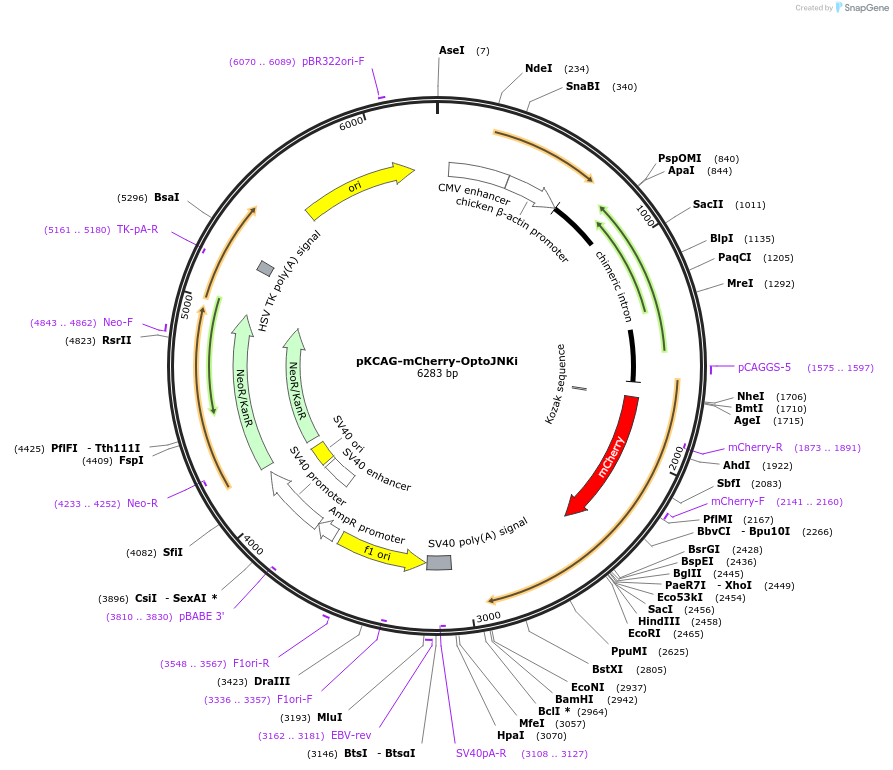
pKCAG-mCherry-OptoJNKi
(Plasmid
#89738)
-
PurposeExpression of light-regulated JNK inhibitor, wild-type photosensor, inhibitor JIP11
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 89738 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepKCAG-mCherry-C1
-
Backbone manufacturerDerived by our lab using pmCherry-C1 and CAG promoter
- Backbone size w/o insert (bp) 5765
- Total vector size (bp) 6283
-
Vector typeMammalian Expression ; Fluorescent protein
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Kanamycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameOptoJNKi
-
Alt nameAsLov2Jalpha (404-546)-JIP11
-
SpeciesSynthetic; Avena Sativa and
-
Insert Size (bp)478
-
GenBank IDKY595943
- Promoter CAG
-
Tag
/ Fusion Protein
- mCherry (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site EcoRI (not destroyed)
- 3′ cloning site BamHI (not destroyed)
- 5′ sequencing primer DsRed_mCherry-fwd
- 3′ sequencing primer fastbac reverse
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byWe designed and assembled the insert from de novo synthesised sequences
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
For further information please see:
Melero-Fernandez de Mera RM*, Li LL*, Popinigis A, Cisek K, Tuittila M, Yadav L, Serva A, Courtney MJ (2017) A simple optogenetic MAPK inhibitor design reveals resonance between transcription-regulating circuitry and temporally-encoded inputs. Nat. Commun. 8, 15017 doi: 10.1038/ncomms15017.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pKCAG-mCherry-OptoJNKi was a gift from Michael Courtney (Addgene plasmid # 89738 ; http://n2t.net/addgene:89738 ; RRID:Addgene_89738) -
For your References section:
A simple optogenetic MAPK inhibitor design reveals resonance between transcription-regulating circuitry and temporally-encoded inputs. Melero-Fernandez de Mera RM, Li LL, Popinigis A, Cisek K, Tuittila M, Yadav L, Serva A, Courtney MJ. Nat Commun. 2017 May 12;8:15017. doi: 10.1038/ncomms15017. 10.1038/ncomms15017 PubMed 28497795