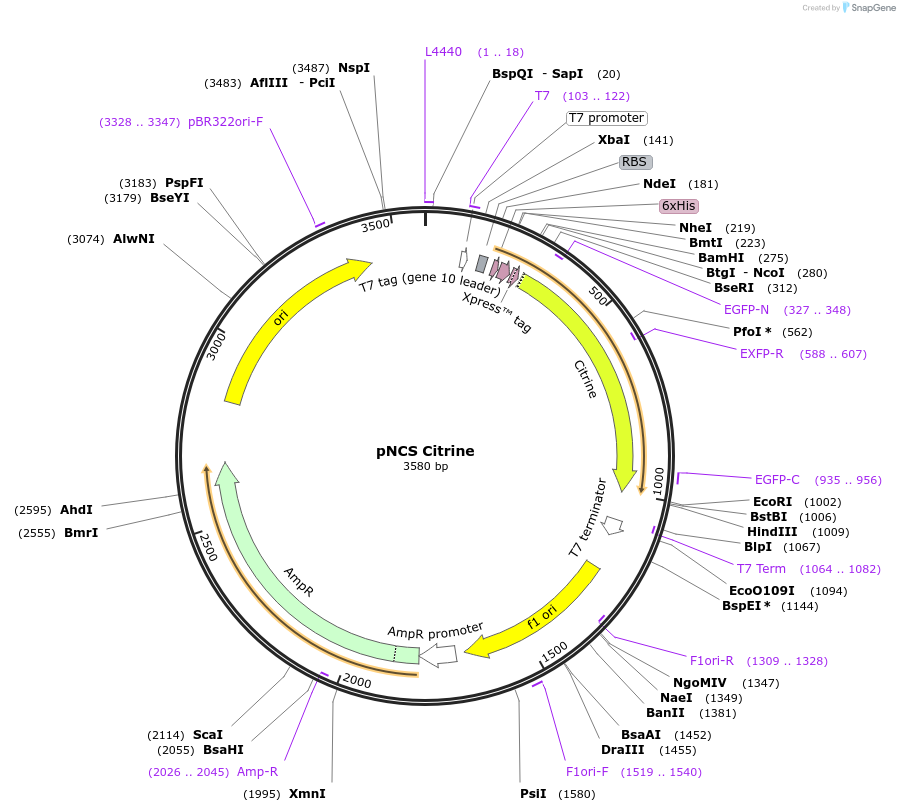
pNCS Citrine
(Plasmid
#91761)
-
PurposeBacterial expression of a fluorescent protein, Citrine
-
Depositing Labs
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 91761 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonemodified pRSETB
-
Backbone manufacturerLife Technologies
- Backbone size w/o insert (bp) 2900
-
Vector typeBacterial Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Growth instructionsE. coli expressing T7 polymerase necessary for fluorescent protein production. Expresses fluorescent protein constitutively. Do not add IPTG.
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameCitrine
-
SpeciesAequorea victoria (jellyfish)
-
Insert Size (bp)735
- Promoter T7
-
Tag
/ Fusion Protein
- Hexa-Histidine tag, Xpress epitope for detection with the anti-Xpress antibody, and enterokinase cleavage site. (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer T7 Promoter Forward
- 3′ sequencing primer T7 Terminator
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
These plasmids are used in University Molecular Biology Laboratories and by High School students to purify fluorescent proteins and create fluorescent protein plate artwork.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pNCS Citrine was a gift from Erik Rodriguez & Roger Tsien (Addgene plasmid # 91761 ; http://n2t.net/addgene:91761 ; RRID:Addgene_91761) -
For your References section:
Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. Griesbeck O, Baird GS, Campbell RE, Zacharias DA, Tsien RY. J Biol Chem. 2001 Aug 3;276(31):29188-94. Epub 2001 May 31. 10.1074/jbc.M102815200 PubMed 11387331