How to Design a Primer
Primer Design for PCR
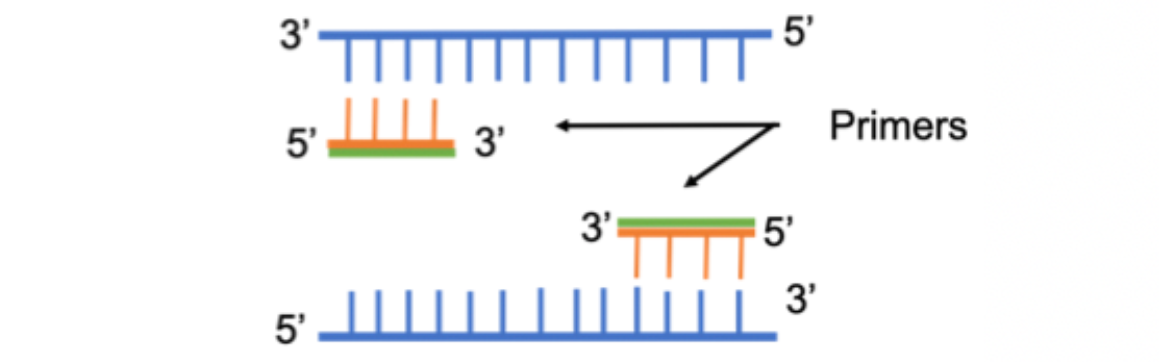
Oligonucleotide primers are necessary when running a PCR reaction. One needs to design primers that are complementary to the template region of DNA. They are synthesized chemically by joining nucleotides together. One must selectively block and unblock repeatedly the reactive groups on a nucleotide when adding a nucleotide one at a time. The main property of primers is that they must correspond to sequences on the template molecule (must be complementary to template strand). However, primers do not need to correspond to the template strand completely; it is essential, however, that the 3’ end of the primer corresponds completely to the template DNA strand so elongation can proceed. Usually a guanine or cytosine is used at the 3’ end, and the 5’ end of the primer usually has stretches of several nucleotides. Also, both of the 3’ ends of the hybridized primers must point toward one another.
The size of the primer is very important as well. Short primers are mainly used for amplifying a small, simple fragment of DNA. On the other hand, a long primer is used to amplify a eukaryotic genomic DNA sample. However, a primer should not be too long (> 30-mer primers) or too short. Short primers produce inaccurate, nonspecific DNA amplification product, and long primers result in a slower hybridizing rate. On average, the DNA fragment that needs to be amplified should be within 1-10 kB in size.
The structure of the primer should be relatively simple and contain no internal secondary structure to avoid internal folding. One also needs to avoid primer-primer annealing which creates primer dimers and disrupts the amplification process. When designing, if unsure about what nucleotide to put at a certain position within the primer, one can include more than one nucleotide at that position termed a mixed site. One can also use a nucleotide-based molecular insert (inosine) instead of a regular nucleotide for broader pairing capabilities.
Taking into consideration the information above, primers should generally have the following properties:
- Length of 18-24 bases
- 40-60% G/C content
- Start and end with 1-2 G/C pairs
- Melting temperature (Tm) of 50-60°C
- Primer pairs should have a Tm within 5°C of each other
-
Primer pairs should not have complementary regions
Note: If you will be including a restriction site at the 5’ end of your primer, note that a 3-6 base pair "clamp" should be added upstream in order for the enzyme to cleave efficiently (e.g. GCGGCG-restriction site-your sequence).
Protocol Video
Watch the protocol video below to learn how to design primers for PCR.


