We narrowed to 83,971 results for: TRI
-
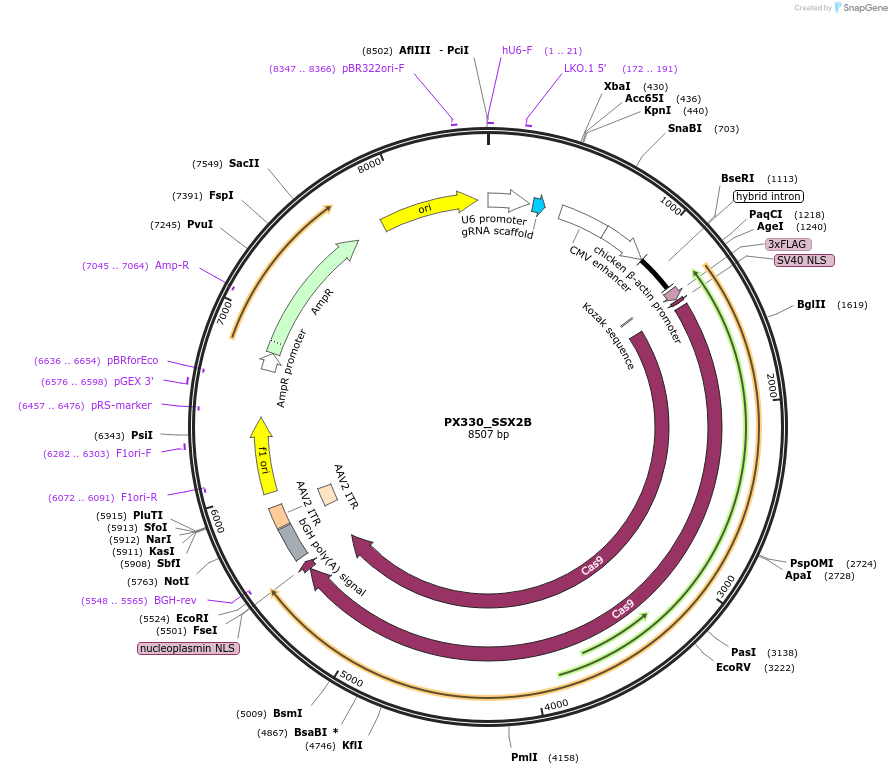
Plasmid#128252PurposeEncodes gRNA for 3' target of human SSX2BDepositorInsertSSX2B gRNA
UseCRISPRAvailable SinceNov. 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
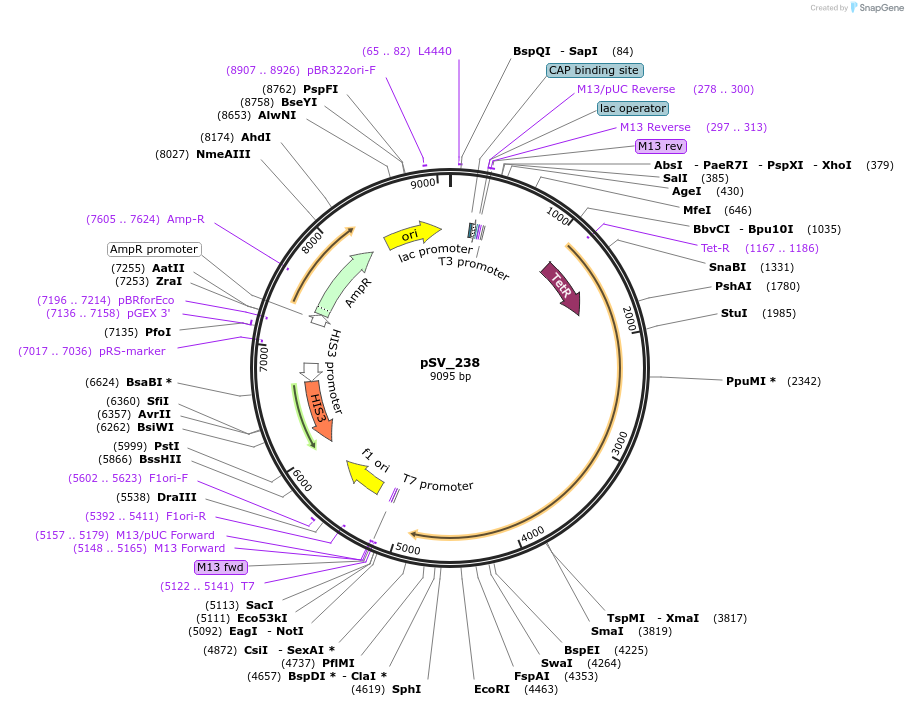
pSV_238
Plasmid#122941PurposeExpression of the repressor to control the insertional promoter.DepositorInsertrTetR.SUM1
ExpressionYeastPromoterPret2Available SinceApril 11, 2019AvailabilityAcademic Institutions and Nonprofits only -
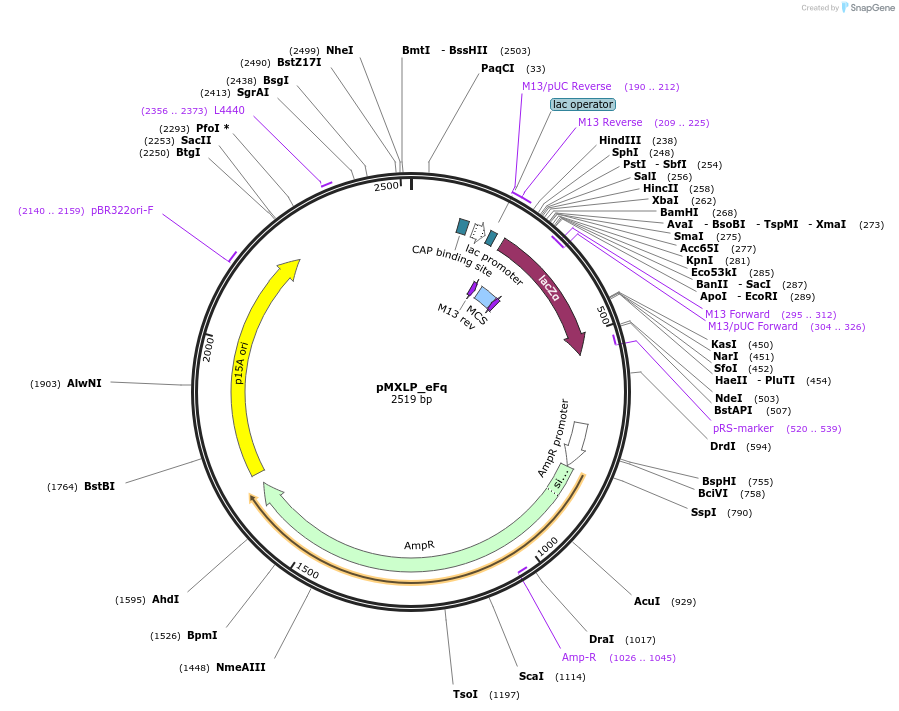
pMXLP_eFq
Plasmid#114248PurposePOC12390, MetClo assembly vector with p15A replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type e-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
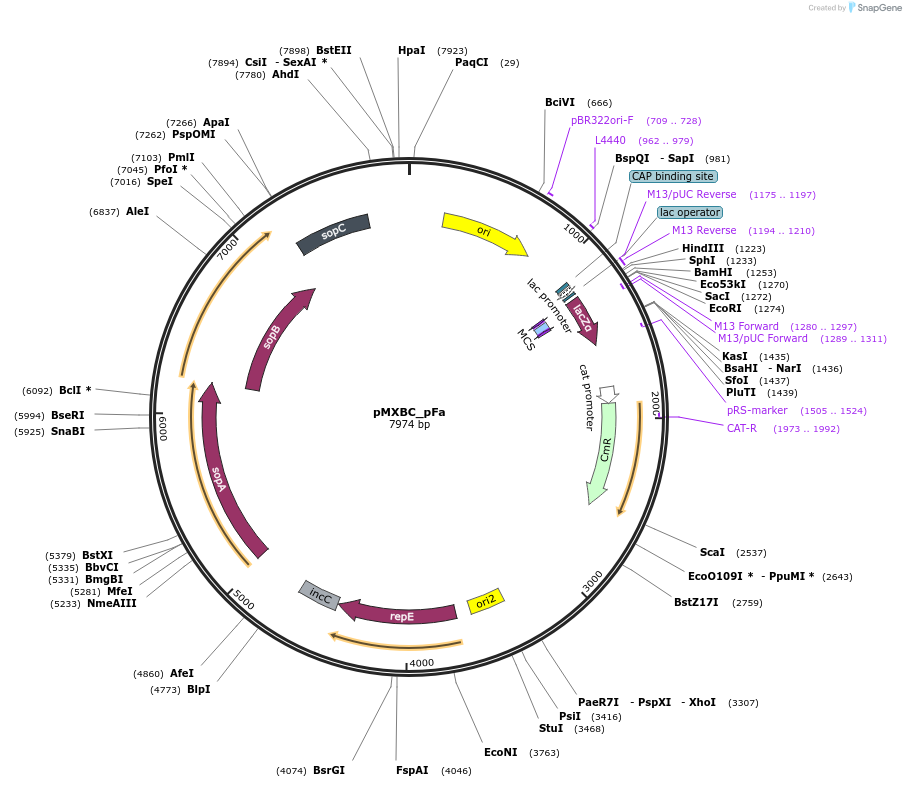
pMXBC_pFa
Plasmid#114249PurposePOC12391, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type p-aDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
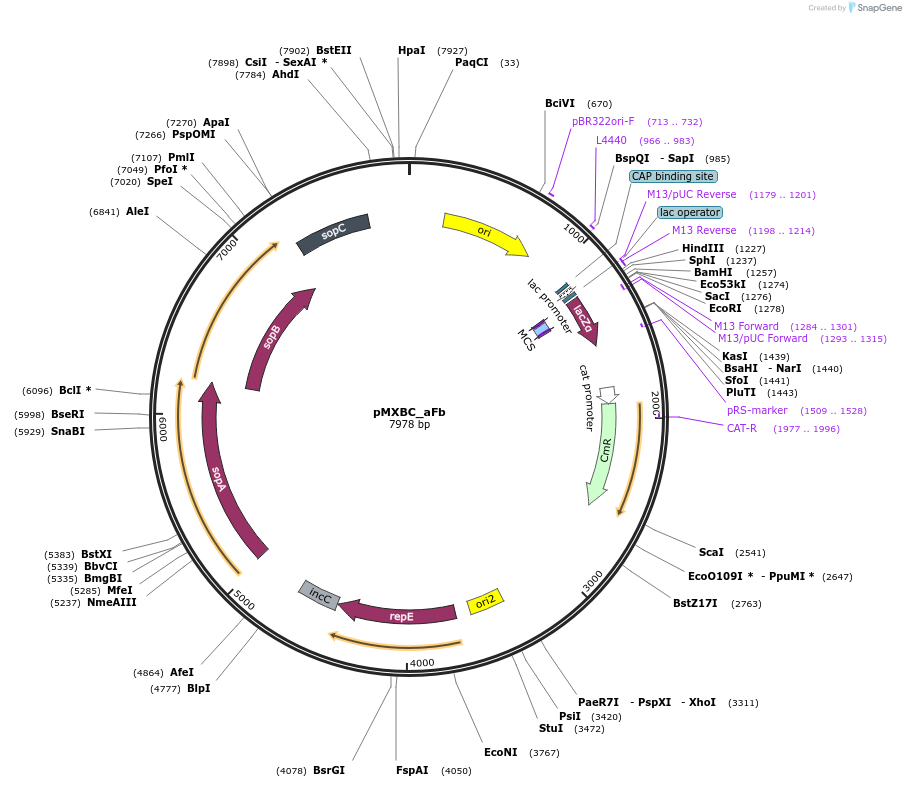
pMXBC_aFb
Plasmid#114250PurposePOC12392, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type a-bDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
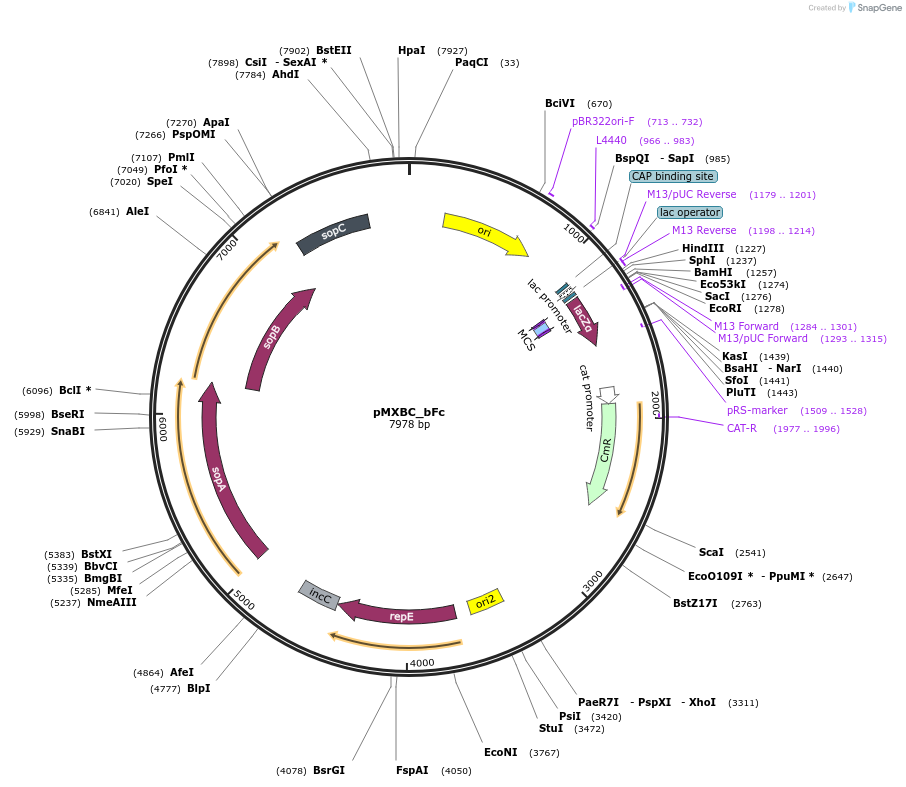
pMXBC_bFc
Plasmid#114251PurposePOC12393, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type b-cDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
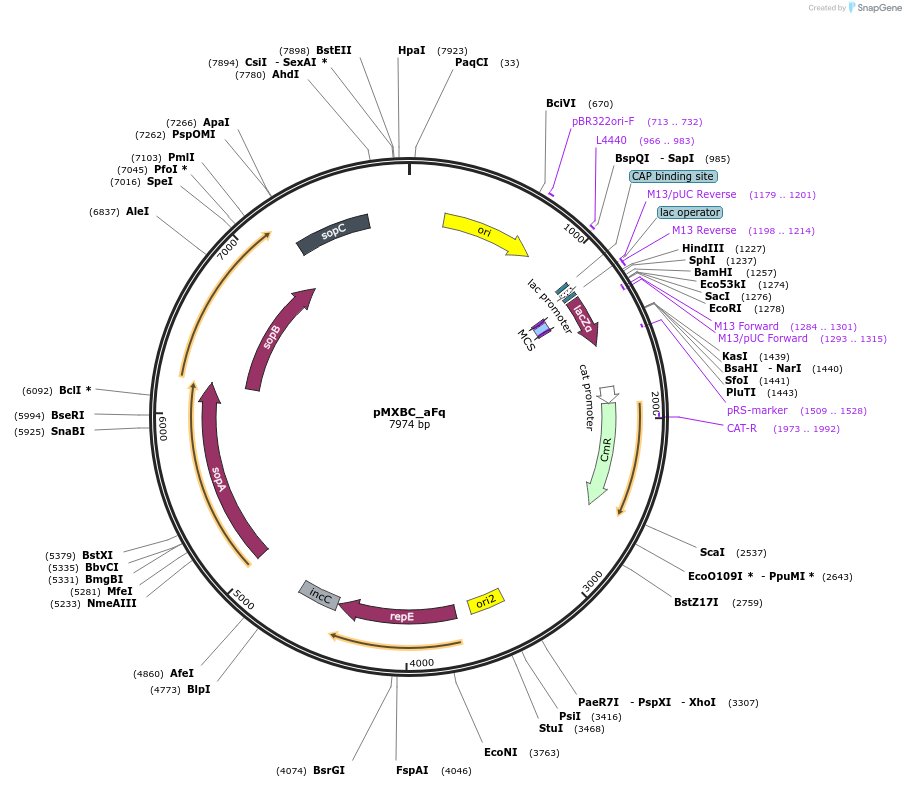
pMXBC_aFq
Plasmid#114252PurposePOC12394, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type a-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
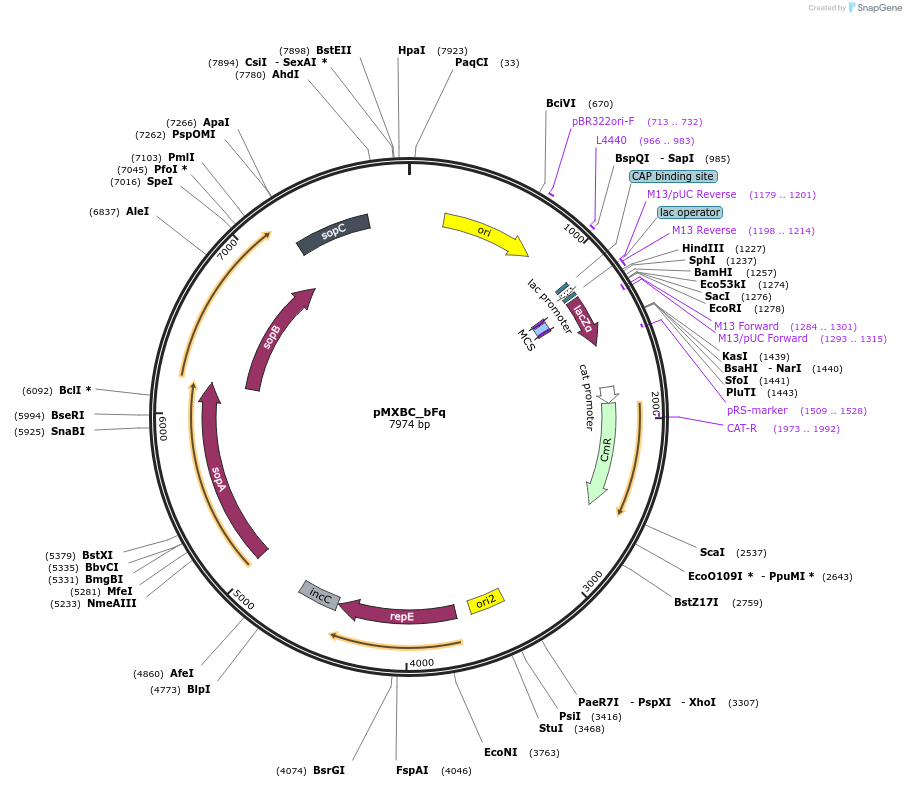
pMXBC_bFq
Plasmid#114253PurposePOC12395, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type b-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
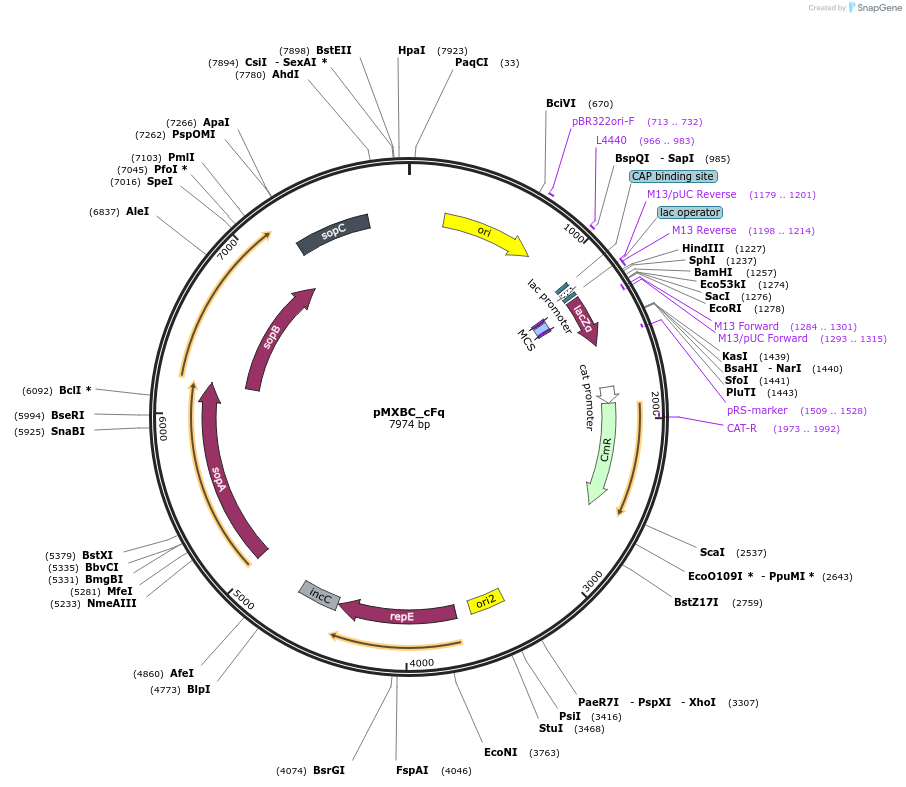
pMXBC_cFq
Plasmid#114254PurposePOC12396, MetClo assembly vector with F replication origin and chloramphenicol resistance for BsaI-based MetClo, adaptor sequence type c-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
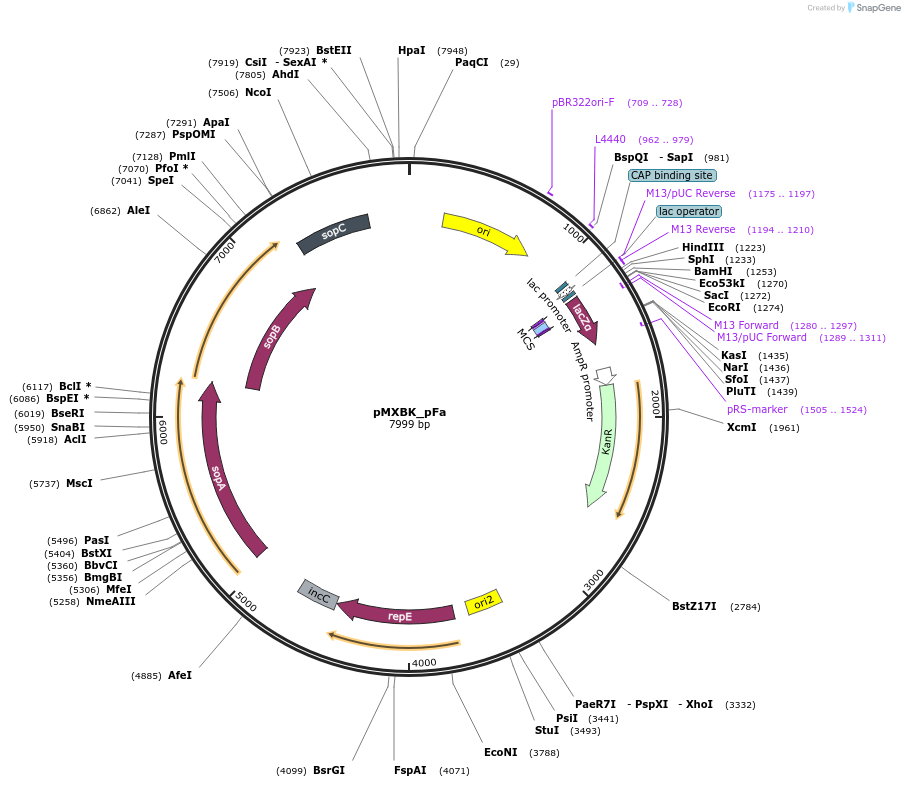
pMXBK_pFa
Plasmid#114255PurposePOC12397, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type p-aDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
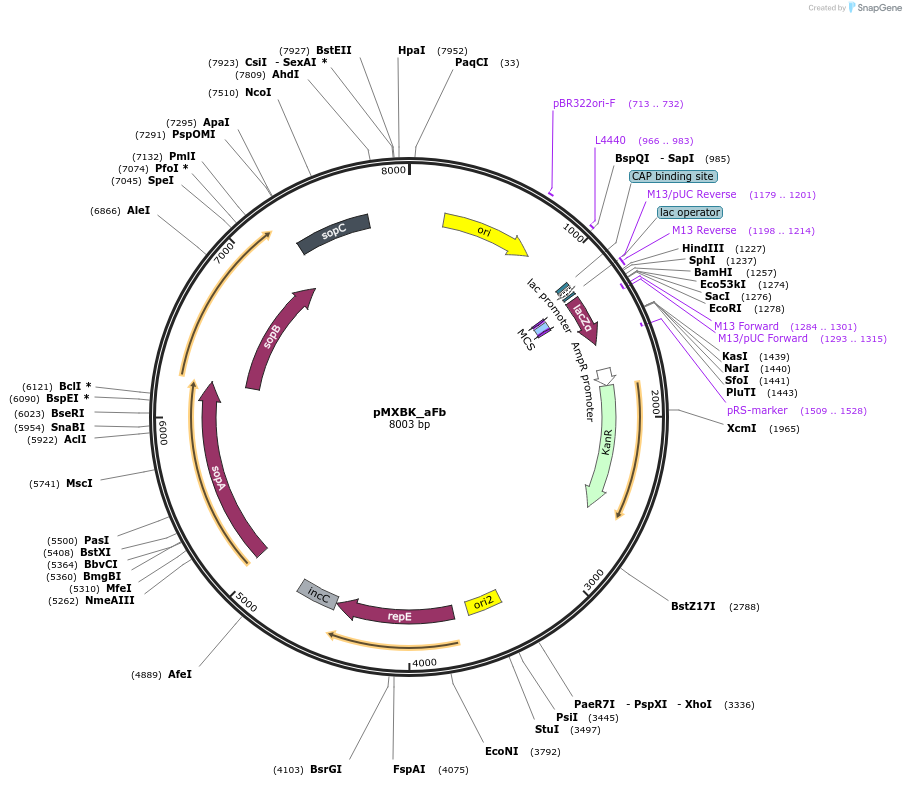
pMXBK_aFb
Plasmid#114256PurposePOC12398, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type a-bDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
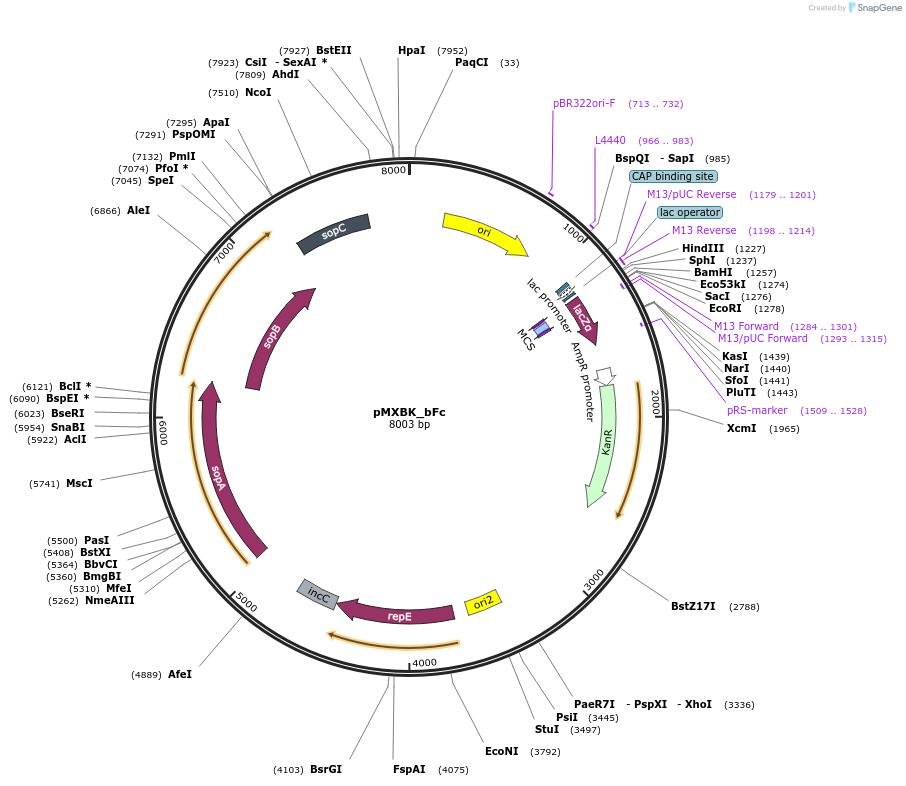
pMXBK_bFc
Plasmid#114257PurposePOC12399, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type b-cDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
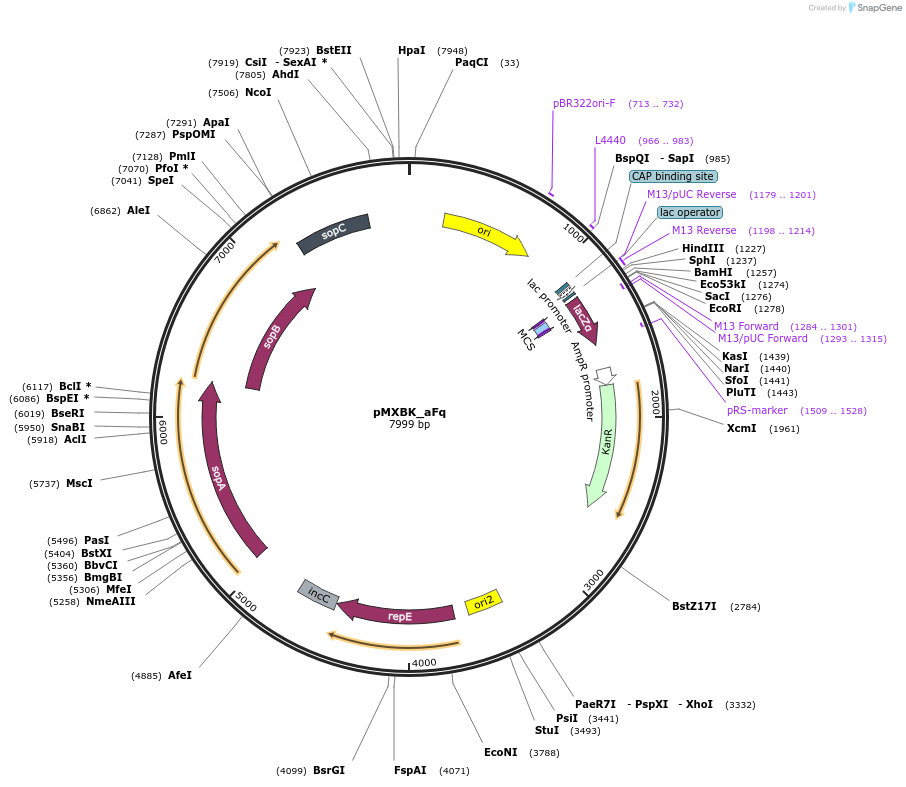
pMXBK_aFq
Plasmid#114258PurposePOC12400, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type a-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
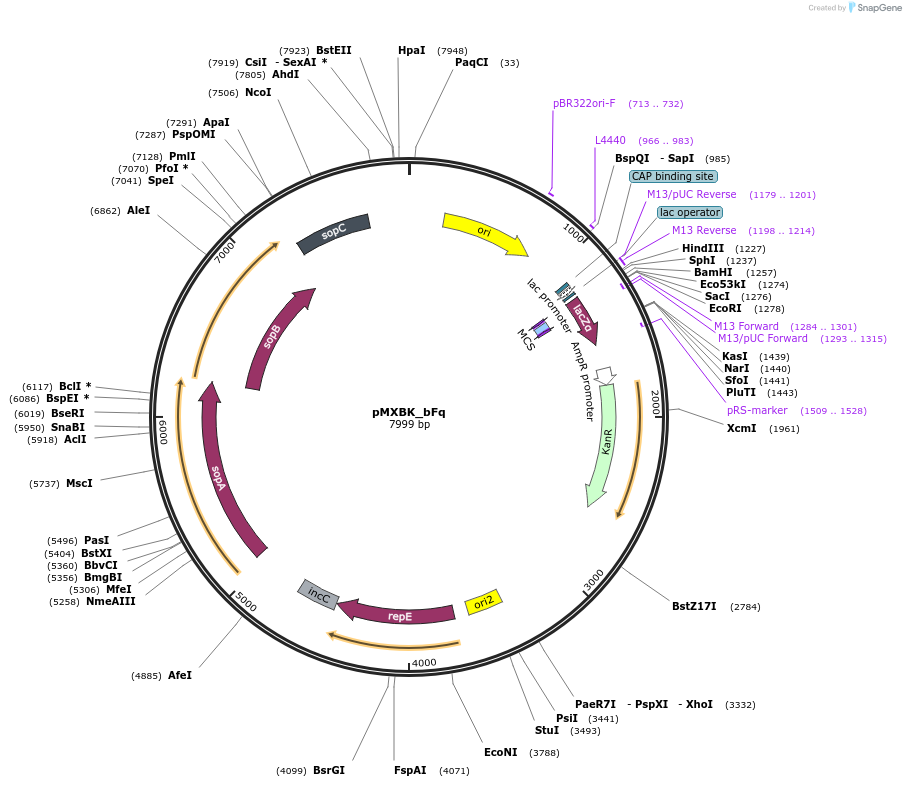
pMXBK_bFq
Plasmid#114259PurposePOC12401, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type b-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
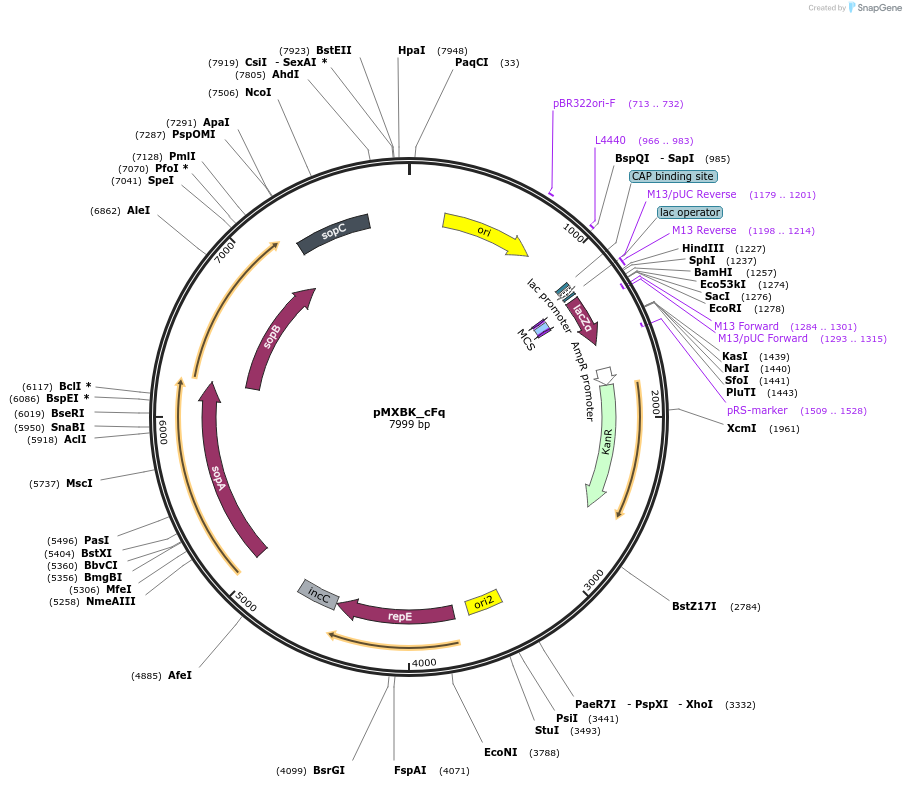
pMXBK_cFq
Plasmid#114260PurposePOC12402, MetClo assembly vector with F replication origin and kanamycin resistance for BsaI-based MetClo, adaptor sequence type c-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
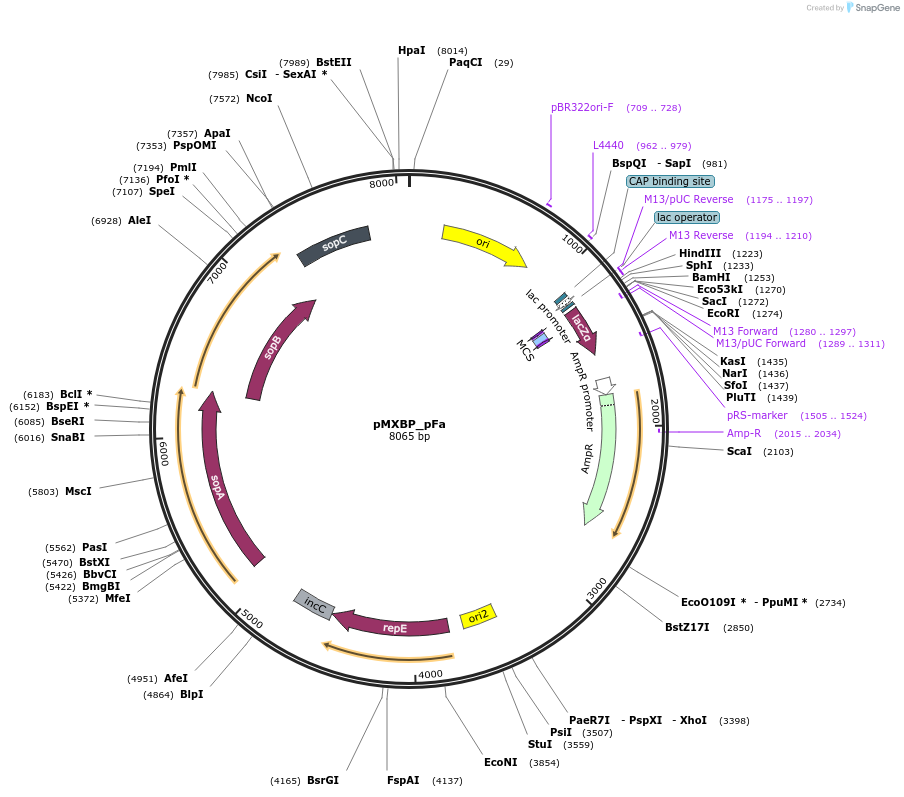
pMXBP_pFa
Plasmid#114261PurposePOC12403, MetClo assembly vector with F replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type p-aDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
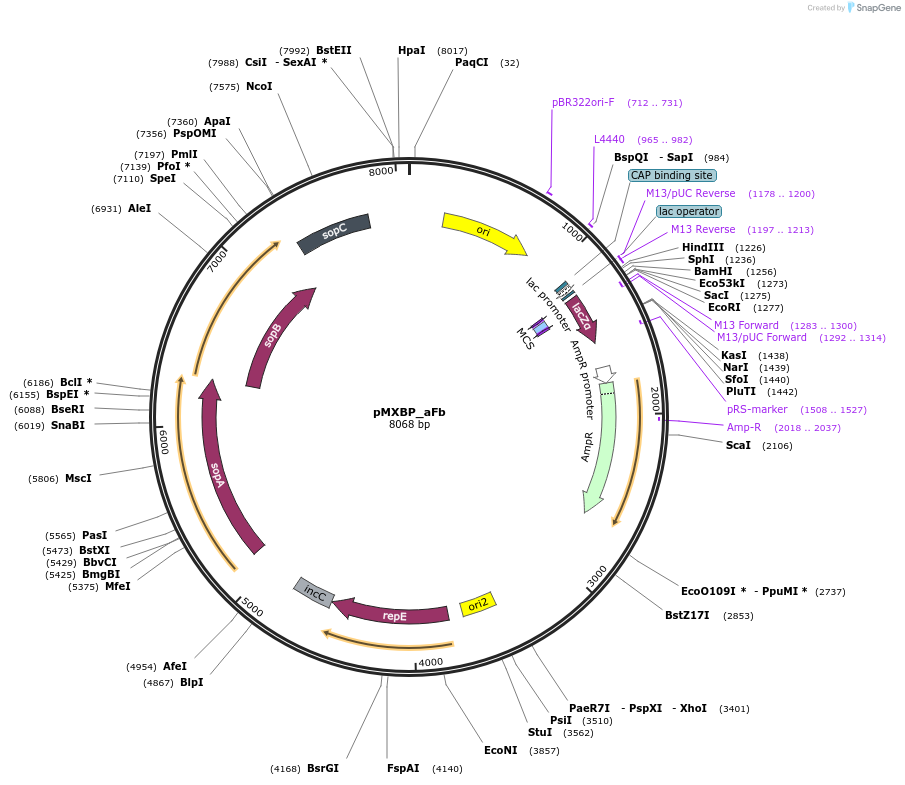
pMXBP_aFb
Plasmid#114262PurposePOC12404, MetClo assembly vector with F replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type a-bDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
pMXBP_bFc
Plasmid#114263PurposePOC12405, MetClo assembly vector with F replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type b-cDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
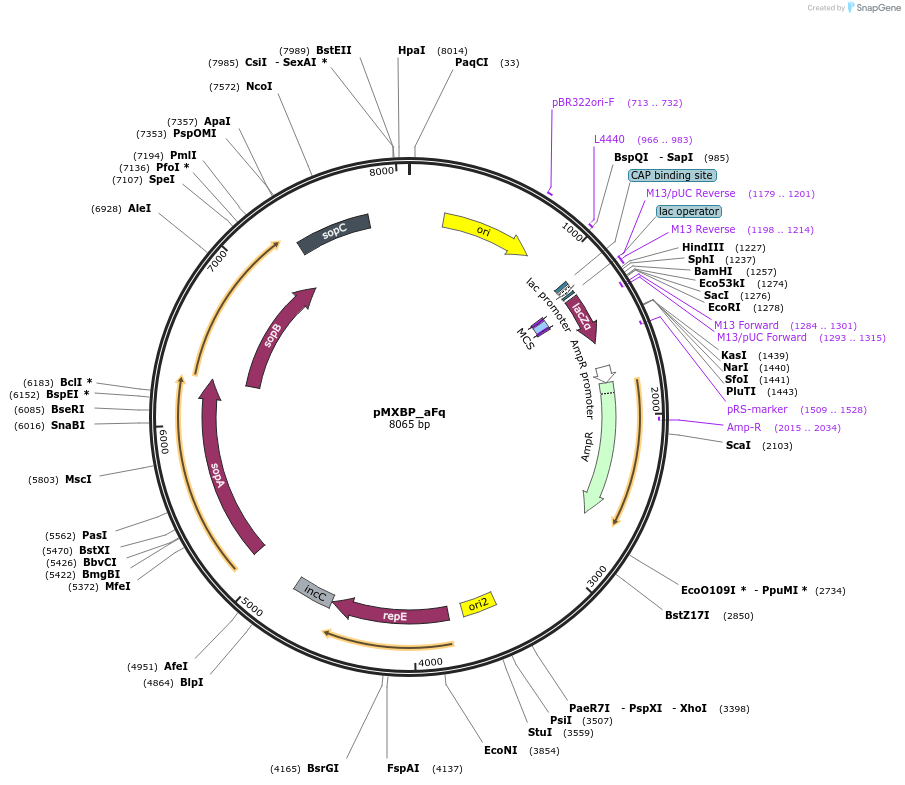
pMXBP_aFq
Plasmid#114264PurposePOC12406, MetClo assembly vector with F replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type a-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
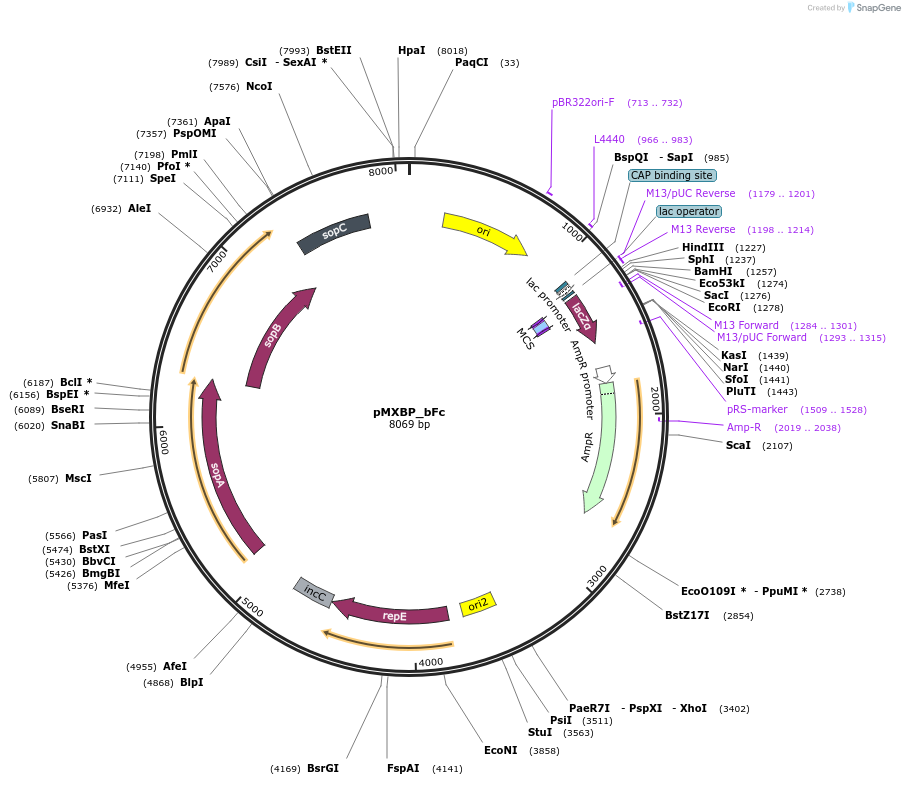
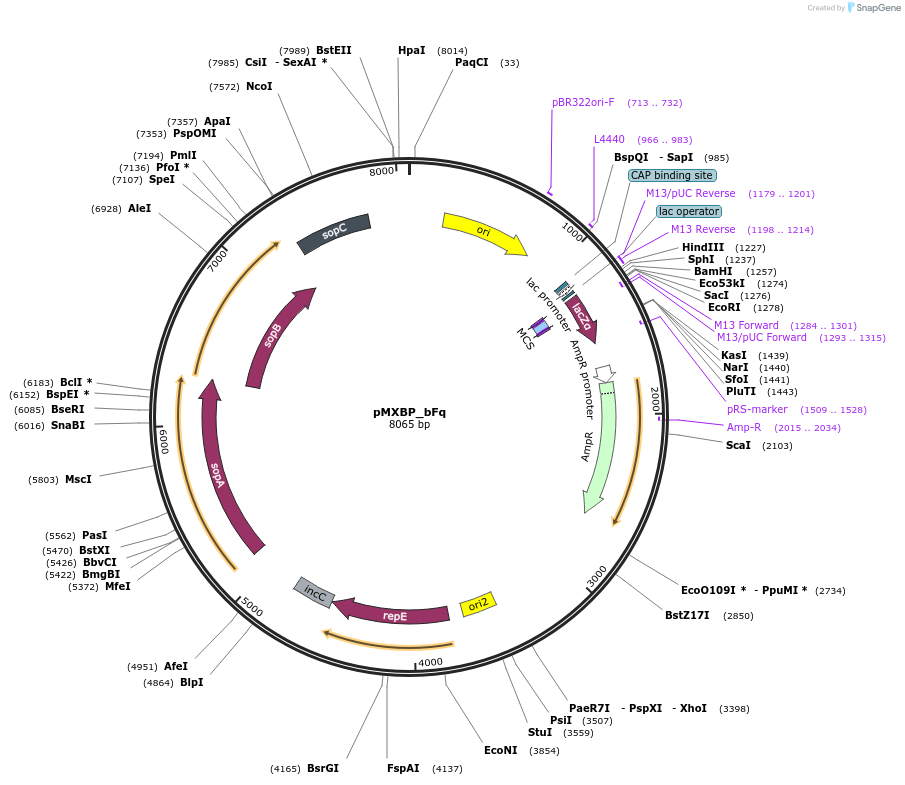
pMXBP_bFq
Plasmid#114265PurposePOC12407, MetClo assembly vector with F replication origin and ampicillin resistance for BsaI-based MetClo, adaptor sequence type b-qDepositorInsertLacZalpha
UseSynthetic BiologyAvailable SinceFeb. 15, 2019AvailabilityAcademic Institutions and Nonprofits only