We narrowed to 79,713 results for: ANT;
-
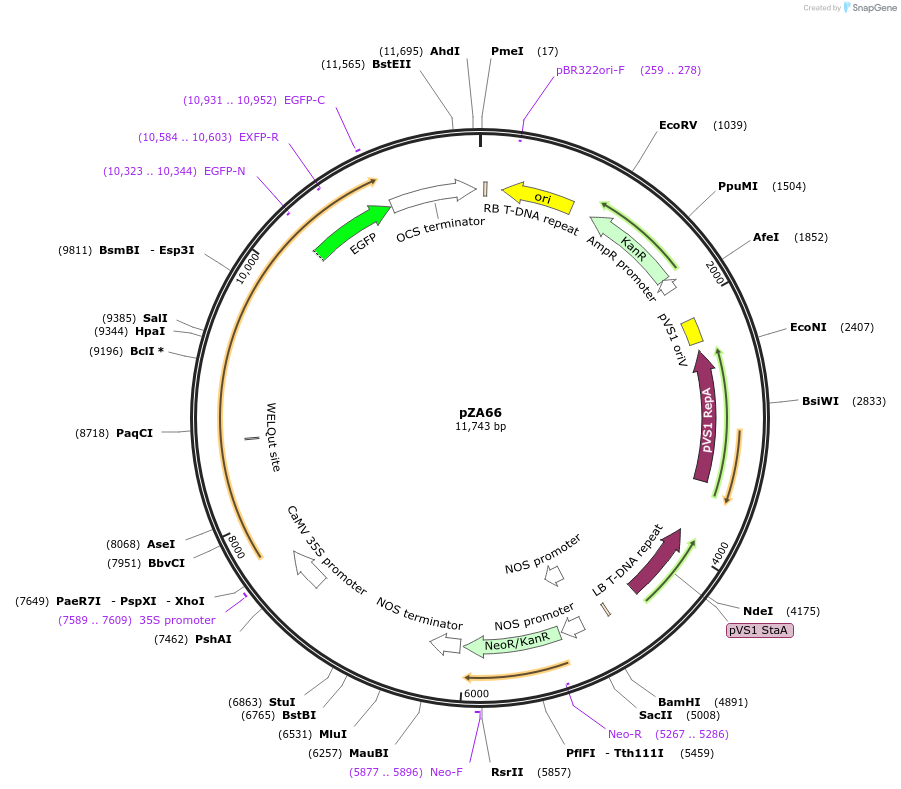
Plasmid#174779PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-D481V-eGFP, Kmr plant selection marker
TagseGFPExpressionPlantMutationD481VAvailable SinceOct. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
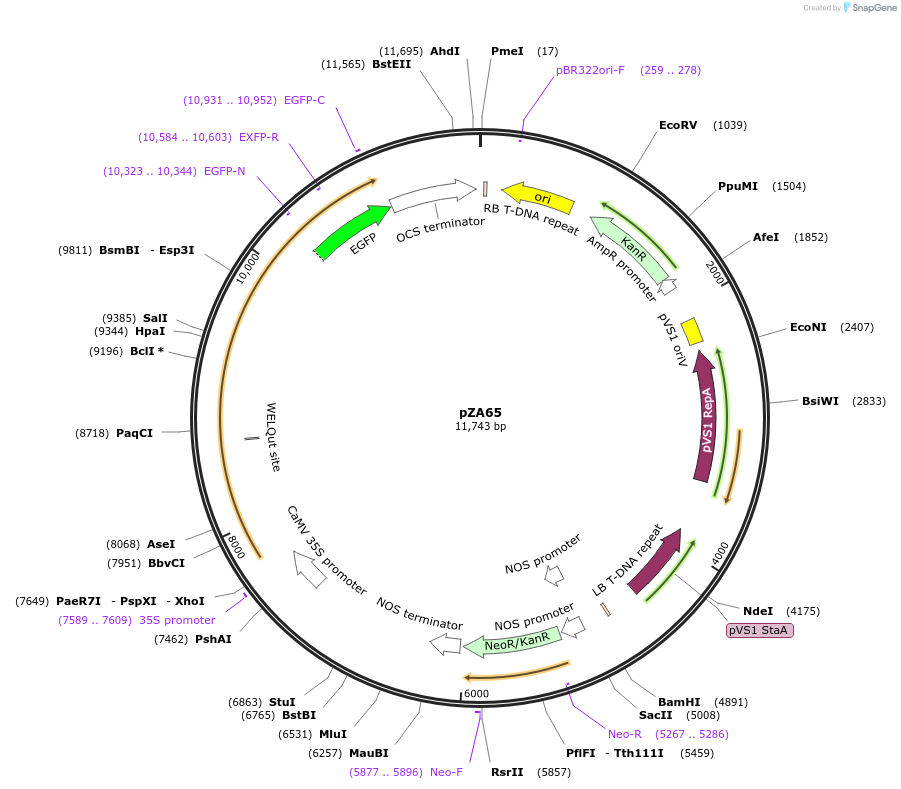
pZA65
Plasmid#174778PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-eGFP, Kmr plant selection marker
TagseGFPExpressionPlantMutationWTAvailable SinceOct. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
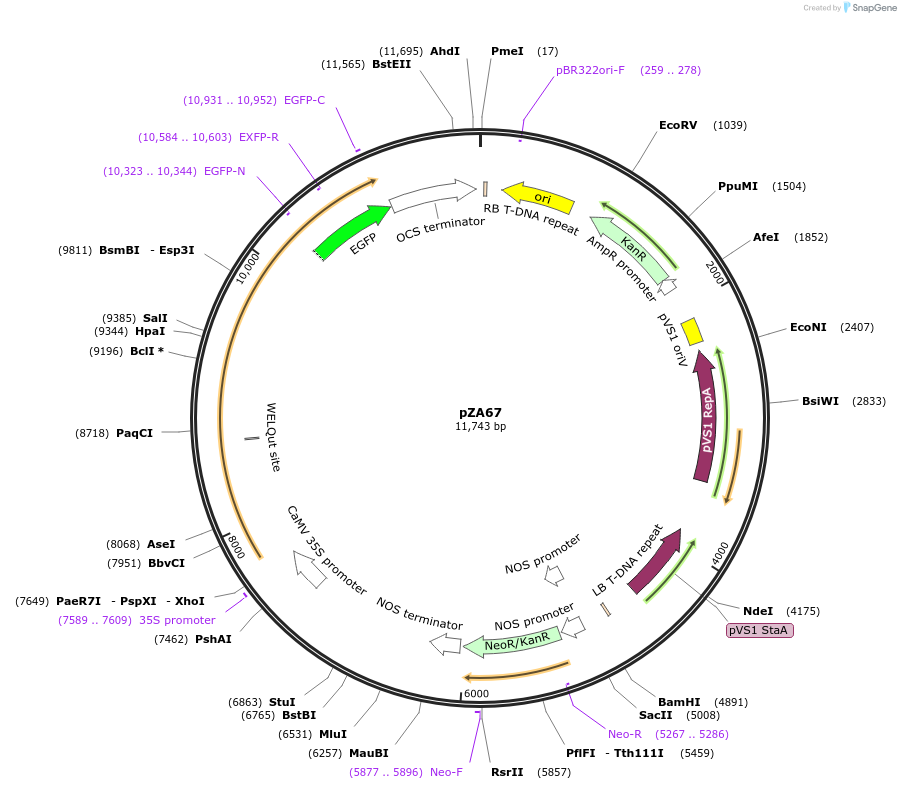
pZA67
Plasmid#174780PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-L17E-eGFP, Kmr plant selection marker
TagseGFPExpressionPlantMutationL17EAvailable SinceOct. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
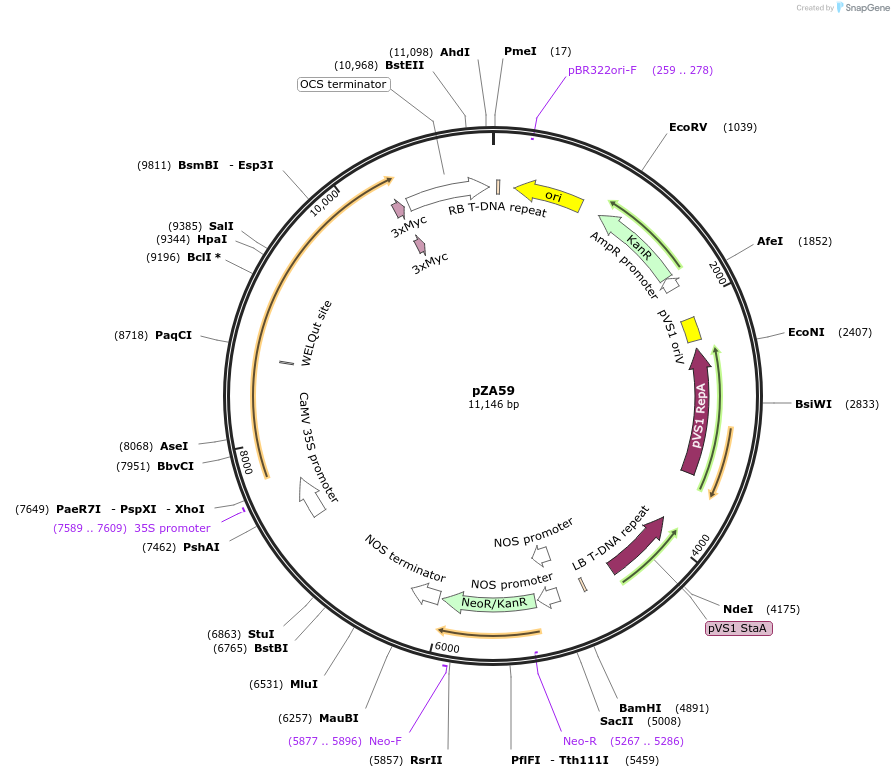
pZA59
Plasmid#174772PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-L17E-4xMyc, Kmr plant selection marker
Tags4xMycExpressionPlantMutationL17EAvailable SinceOct. 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
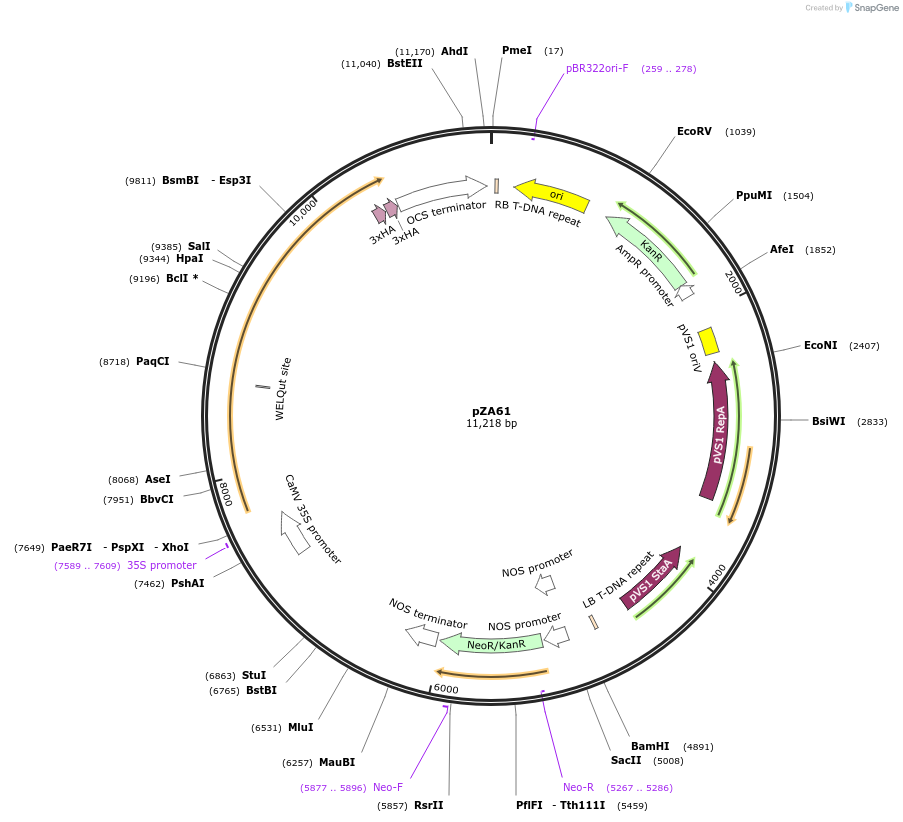
pZA61
Plasmid#174774PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-6xHA, Kmr plant selection marker
Tags6xHAExpressionPlantMutationWTAvailable SinceOct. 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
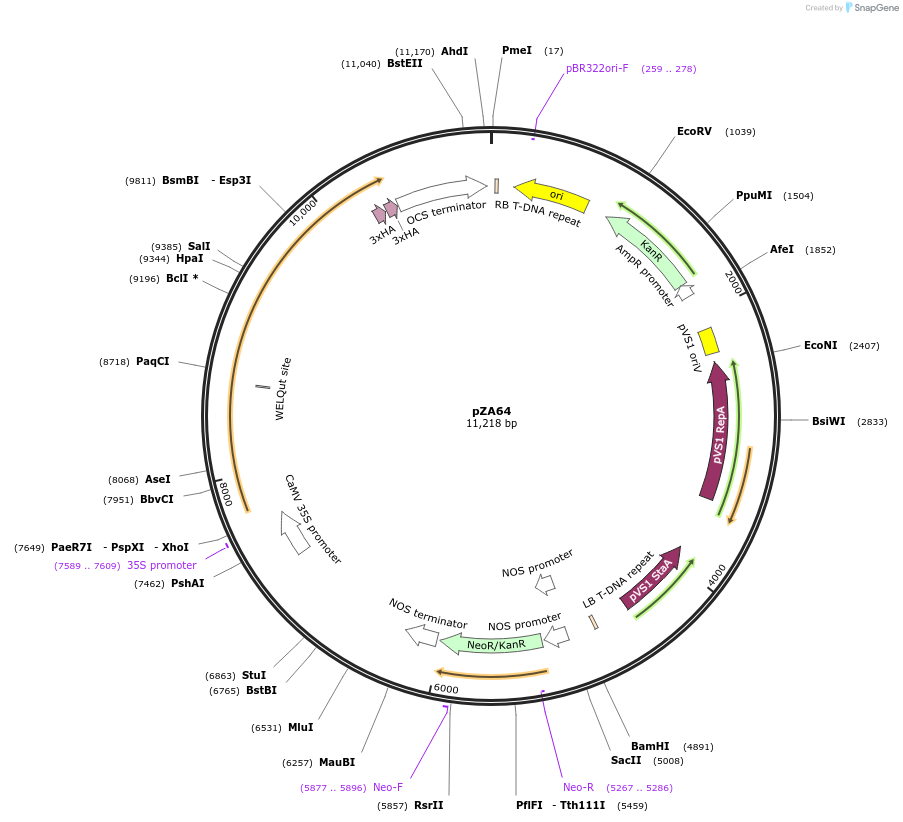
pZA64
Plasmid#174777PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-L17E/D481V-6xHA, Kmr plant selection marker
Tags6xHAExpressionPlantMutationL17E/D481VAvailable SinceOct. 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
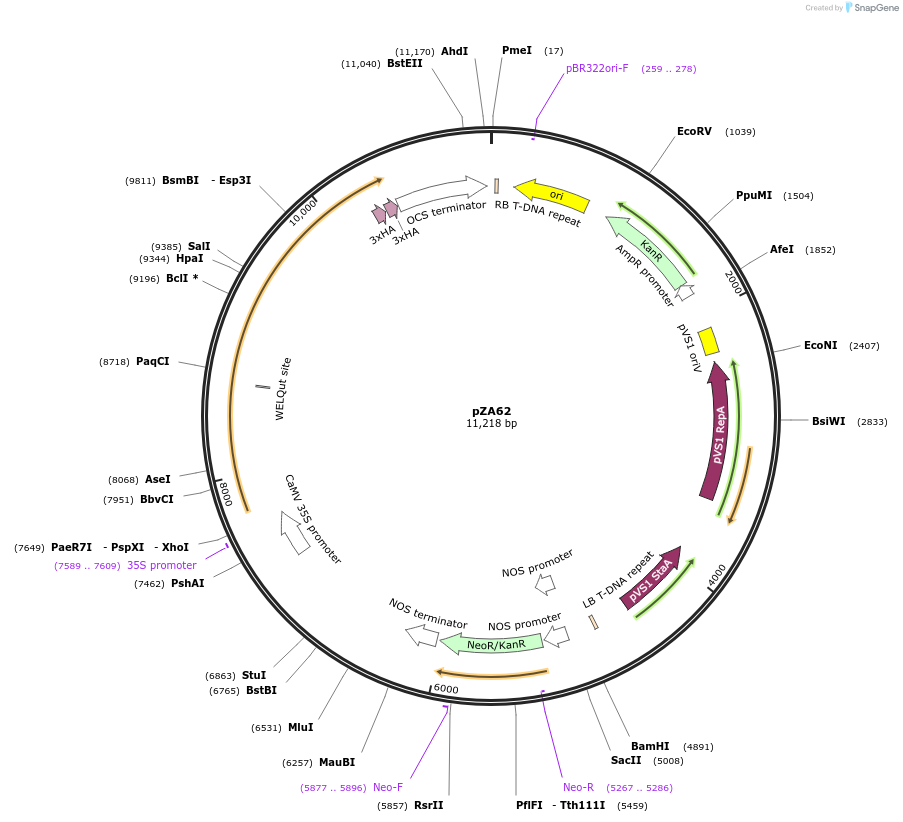
pZA62
Plasmid#174775PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-D481V-6xHA, Kmr plant selection marker
Tags6xHAExpressionPlantMutationD481VAvailable SinceOct. 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
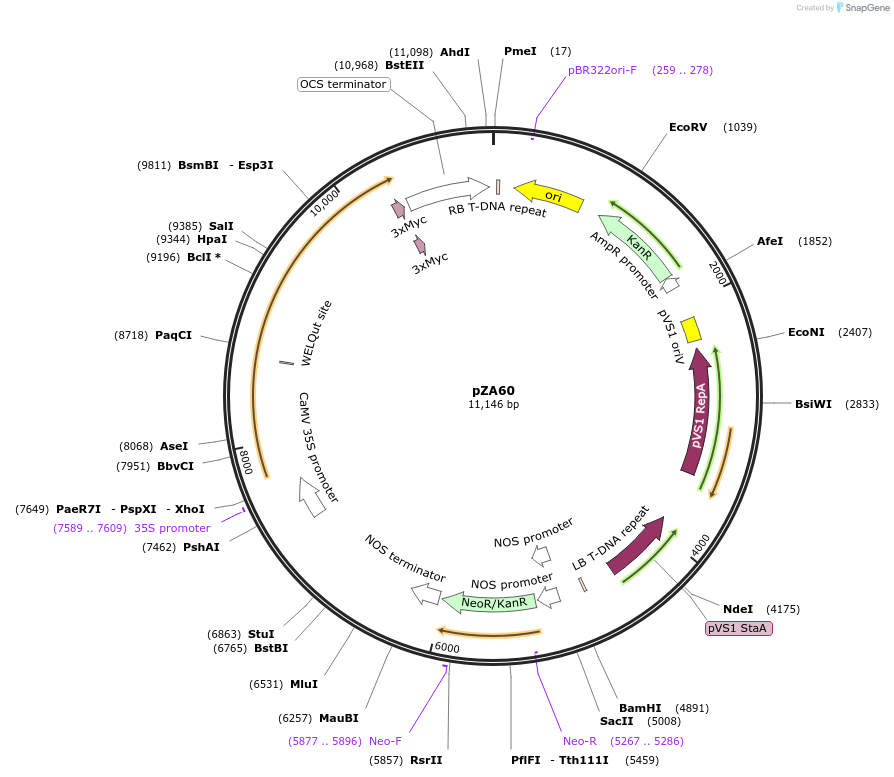
pZA60
Plasmid#174773PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-L17E/D481V-4xMyc, Kmr plant selection marker
Tags4xMycExpressionPlantMutationL17E/D481VAvailable SinceOct. 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
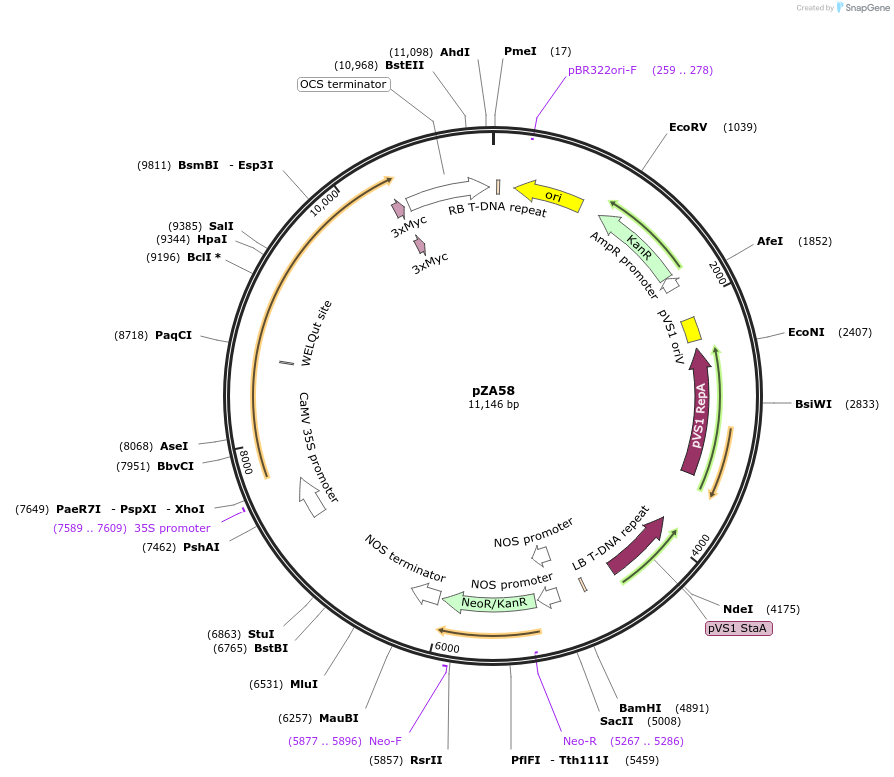
pZA58
Plasmid#174771PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-D481V-4xMyc, Kmr plant selection marker
Tags4xMycExpressionPlantMutationD481VAvailable SinceOct. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
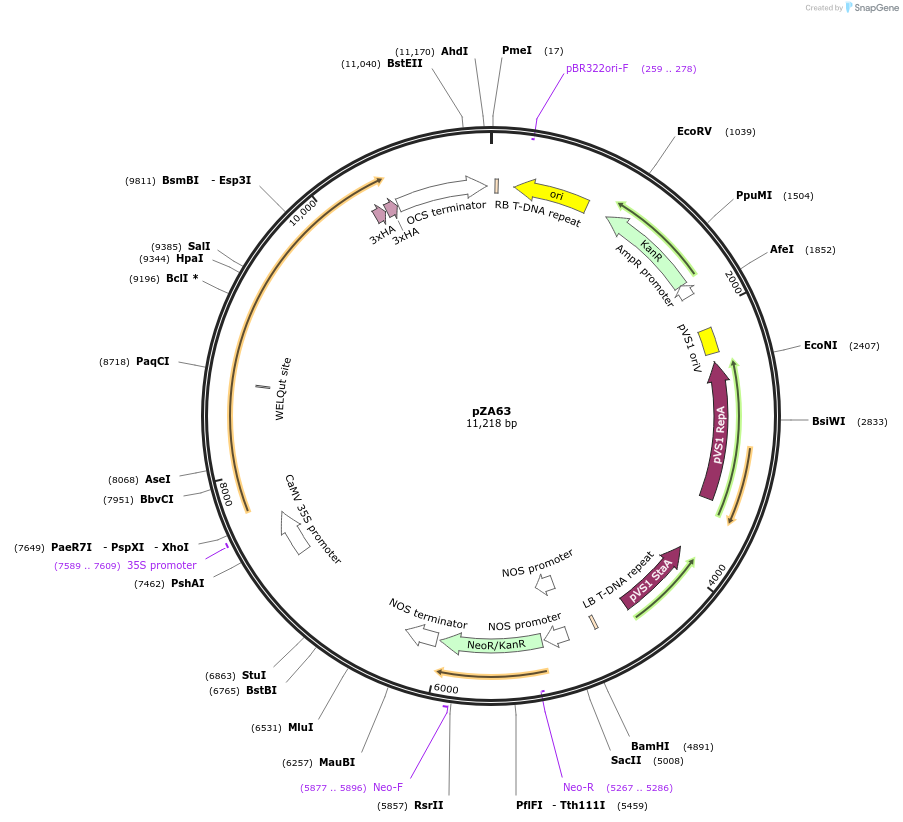
pZA63
Plasmid#174776PurposeIn planta gene expressionDepositorInsertGG_NbZAR1syn-L17E-6xHA, Kmr plant selection marker
Tags6xHAExpressionPlantMutationL17EAvailable SinceOct. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
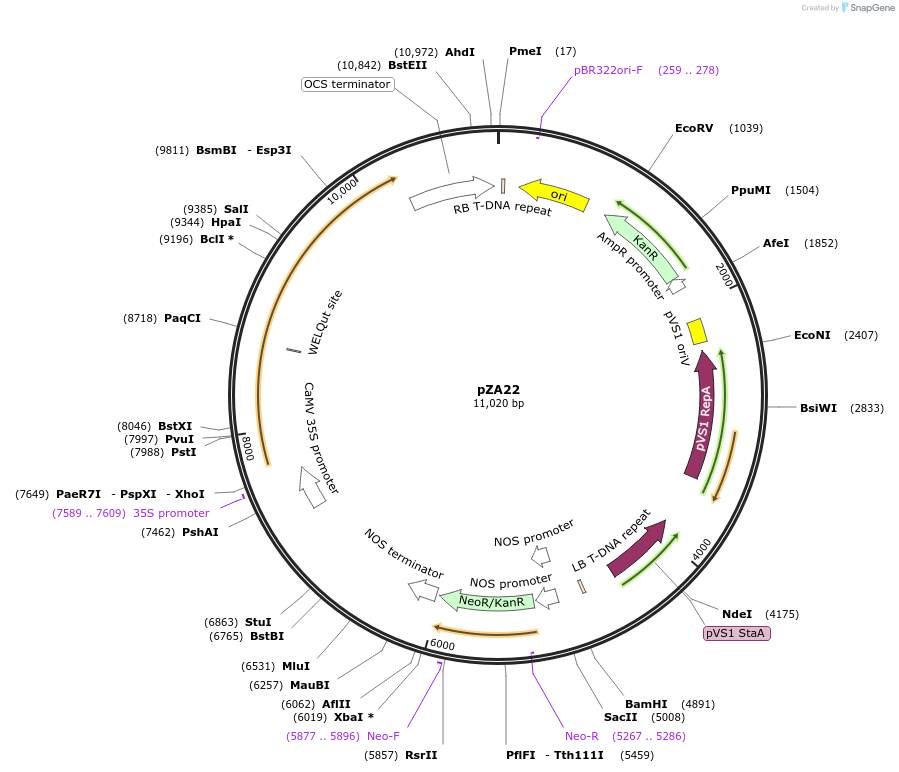
pZA22
Plasmid#158500PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantMutationD481VAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
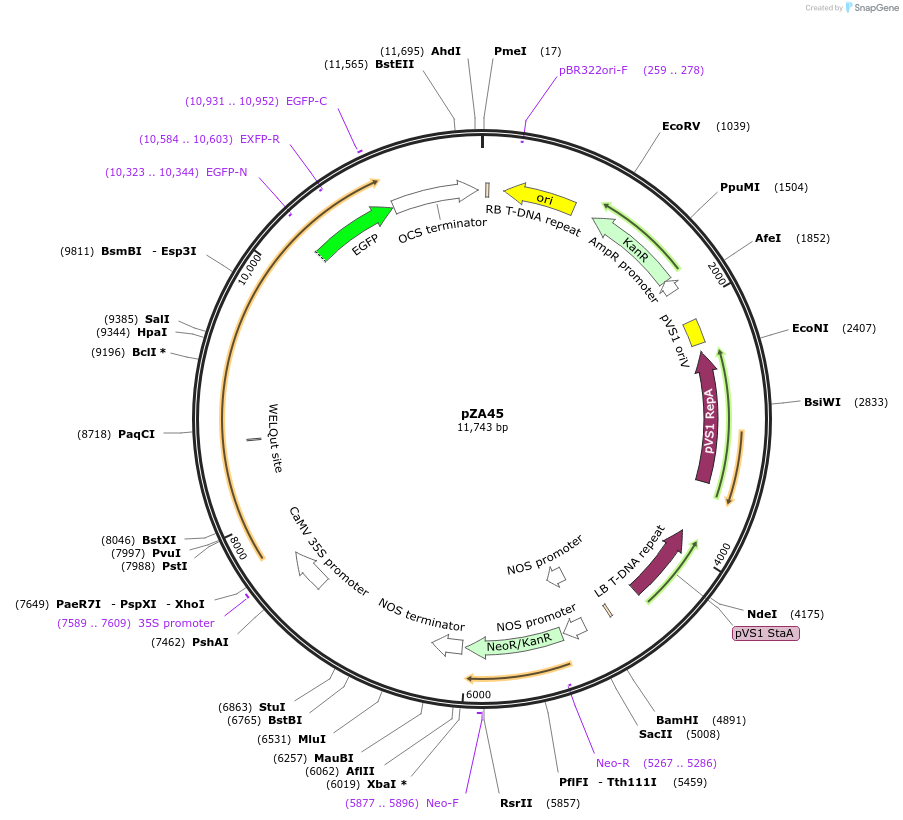
pZA45
Plasmid#158523PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-eGFP. Km plant selection marker.DepositorInsertZAR1-eGFP
TagseGFPExpressionPlantAvailable SinceSept. 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
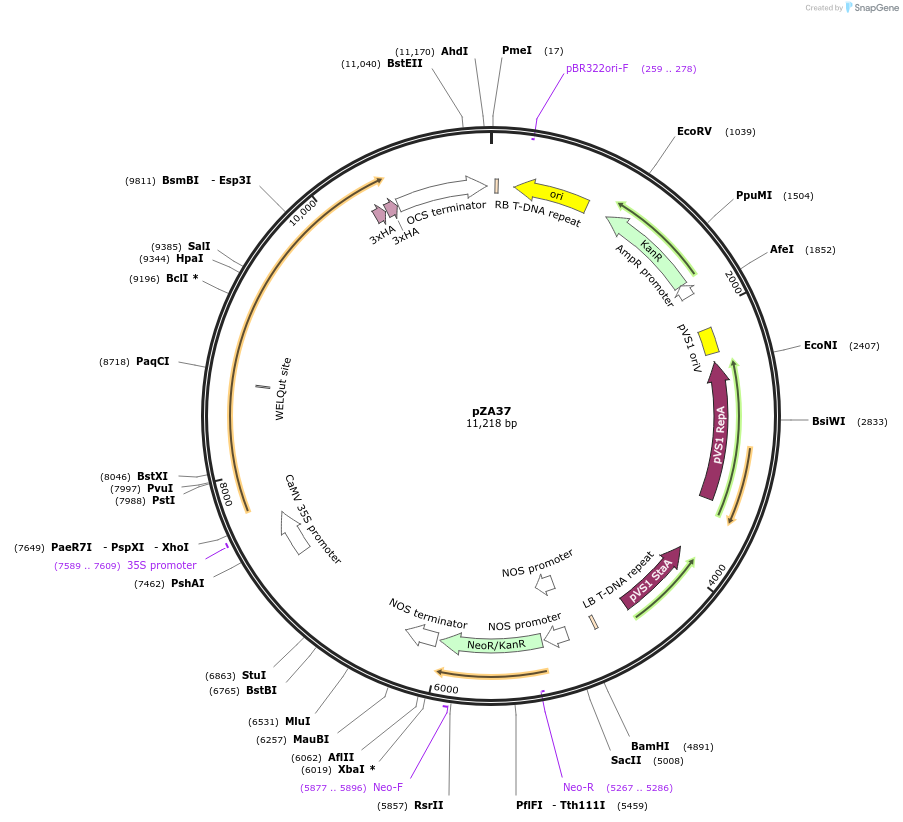
pZA37
Plasmid#158515PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-6xHA. Km plant selection marker.DepositorInsertZAR1-6xHA
Tags6xHAExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
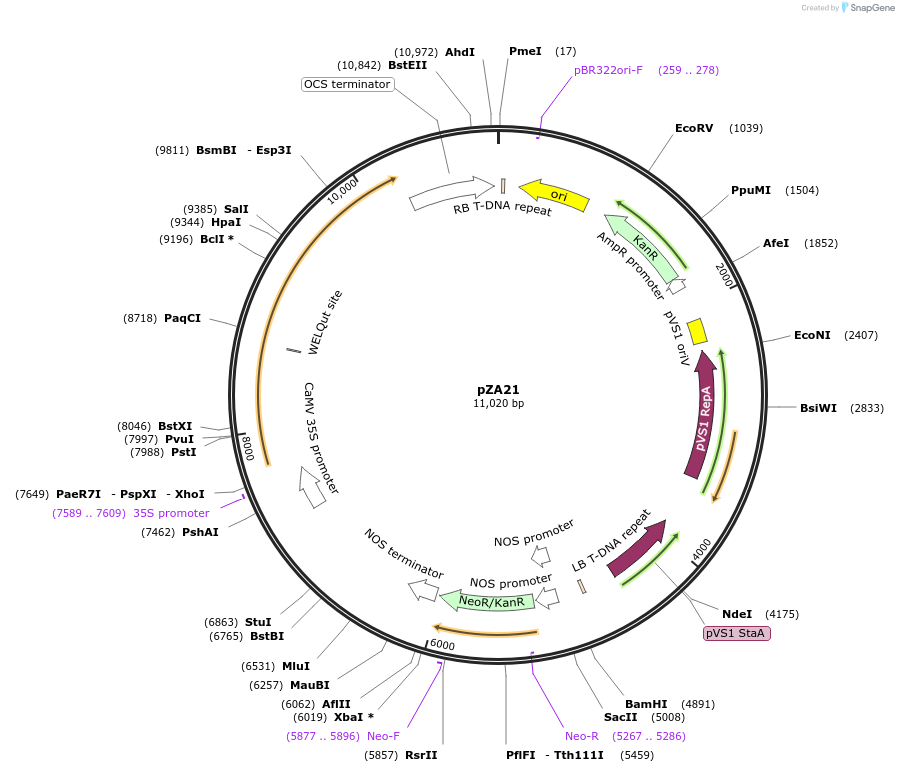
pZA21
Plasmid#158499PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
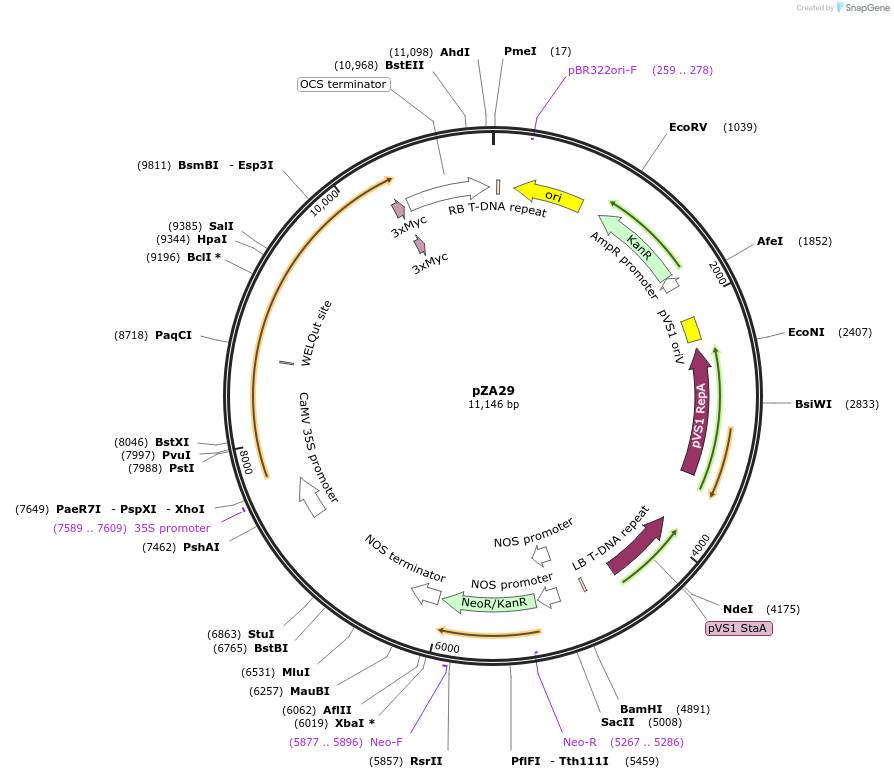
pZA29
Plasmid#158507PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-4xMyc. Km plant selection marker.DepositorInsertZAR1-4xMyc
Tags4xMycExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
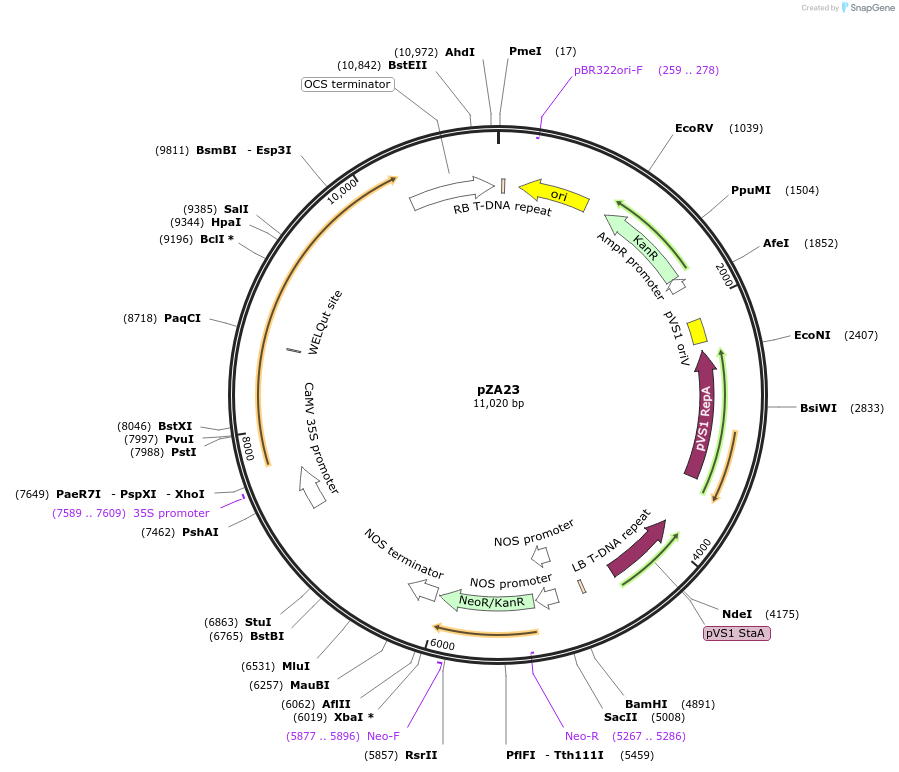
pZA23
Plasmid#158501PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 L17E. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantMutationL17EAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
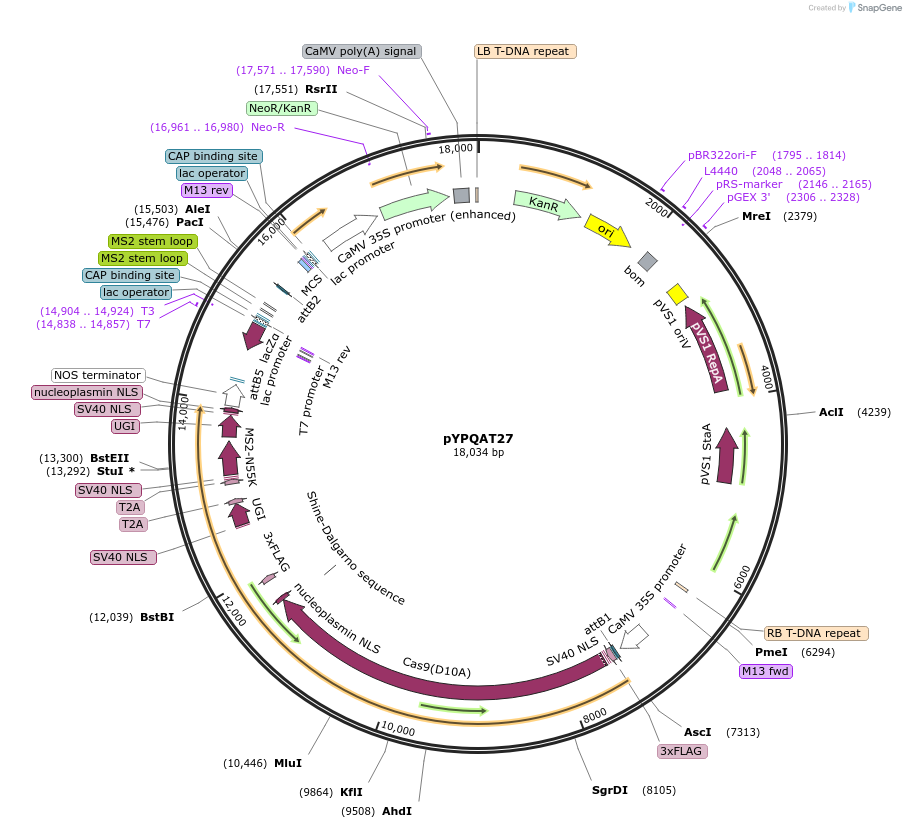
pYPQAT27
Plasmid#223399PurposeT-DNA vector for PmCDA1-CBE_V04 based C-to-T base editing for dicot plants; NGG PAM; 5' editing window; PmCDA1-CBE_V04 was driven by 2x35s and the sgRNA was driven by AtU3; Kanamycin for plant select.DepositorInsert2x35s-SpCas9D10A-PmCDA-UGI-T2A-MS2-UGI-AtU3-gRNA scaffold 2.0
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
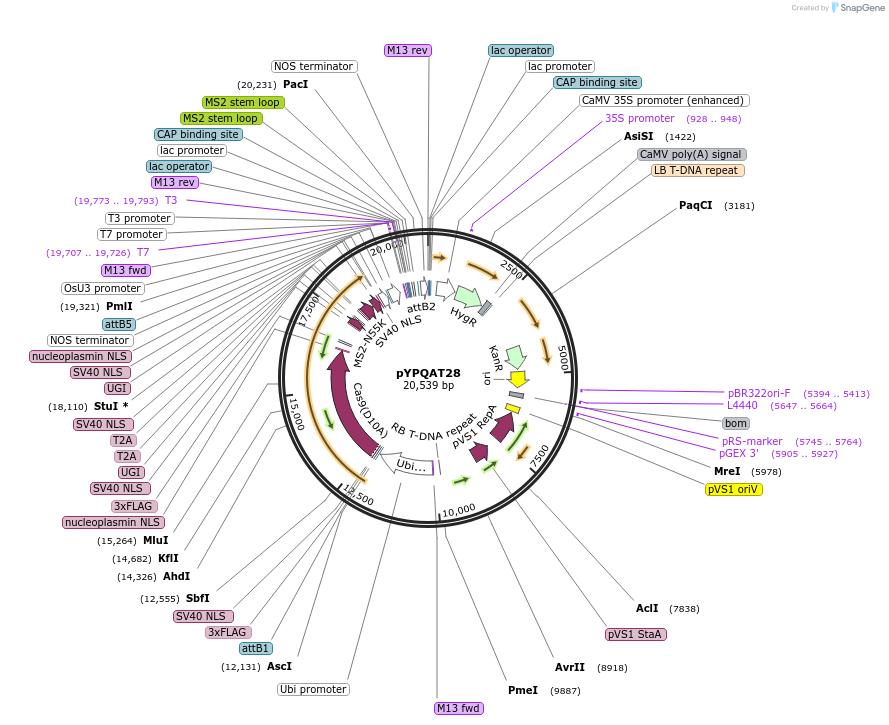
pYPQAT28
Plasmid#223400PurposeT-DNA vector for PmCDA1-CBE_V04 based C-to-T base editing for monocot plants; NGG PAM; 5' editing window; PmCDA1-CBE_V04 was driven by ZmUbi1 and sgRNA was driven by OsU3; Hygromycin for plant select.DepositorInsertZmUbi-SpCas9D10A-PmCDA-UGI-T2A-MS2-UGI-OsU3-gRNA scaffold 2.0
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
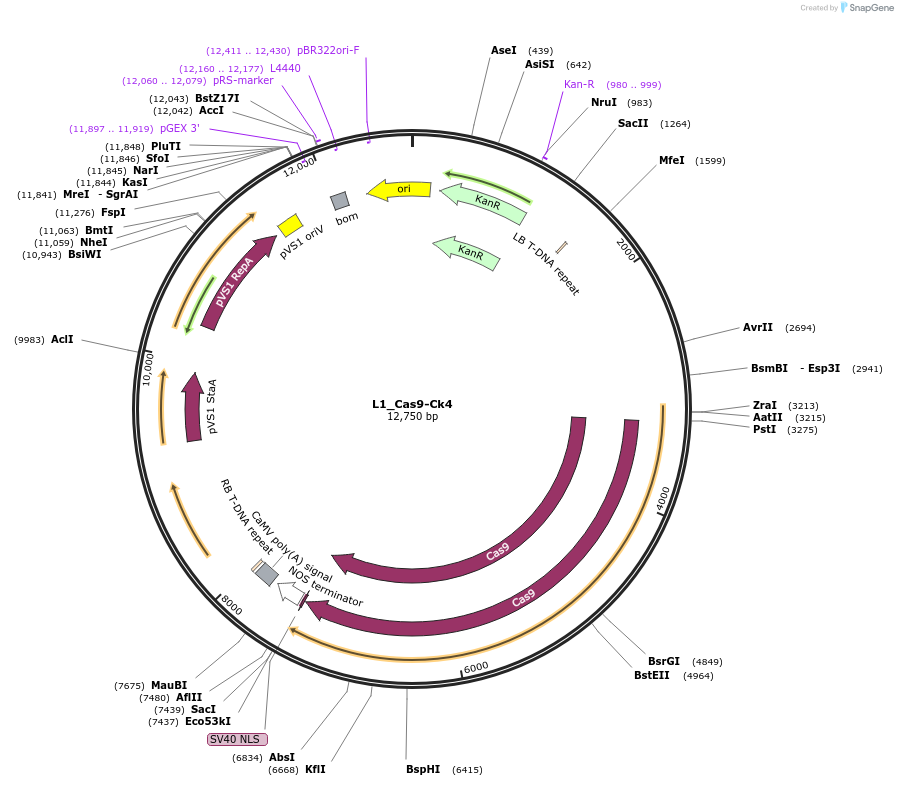
L1_Cas9-Ck4
Plasmid#136135PurposeL1 in position 4, for AtcoCas9 (Puchta) expression in L2 CRISPR vectorsDepositorInsertp5-MpEF1a:Cas9_3tNos-35S
UseSynthetic BiologyExpressionPlantMutationBsaI/ SapI domesticatedAvailable SinceNov. 30, 2021AvailabilityAcademic Institutions and Nonprofits only -
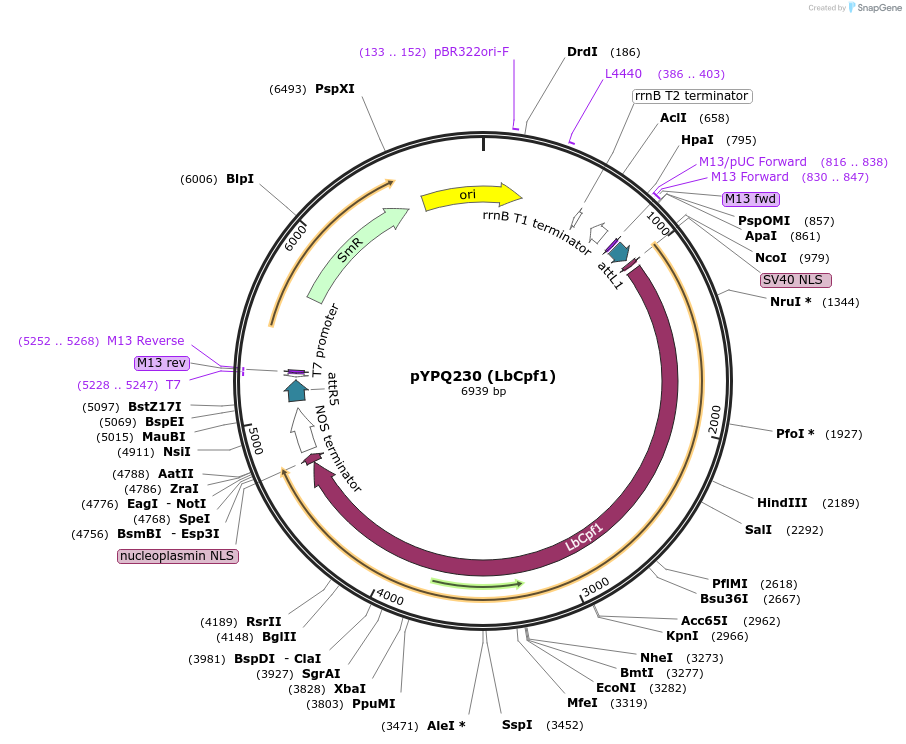
pYPQ230 (LbCpf1)
Plasmid#86210PurposeLbCpf1 Gateway entry plasmidDepositorInsertLbCpf1
UseCRISPR; Gateway compatible lbcpf1 entry cloneTags5' and 3' NLSExpressionPlantAvailable SinceApril 25, 2017AvailabilityAcademic Institutions and Nonprofits only