We narrowed to 167,734 results for: mpi
-
Plasmid#78677PurposeInsulated MoClo promoter for E. coli. 36nt DNA spacer empirically selected. Modified from BBa_I13453. Has near identical expression as 36INS1-pBAD_AB [G:36INS2-pBAD:B]DepositorInsertInsulated pBAD Promoter
UseSynthetic BiologyAvailable SinceAug. 4, 2016AvailabilityAcademic Institutions and Nonprofits only -
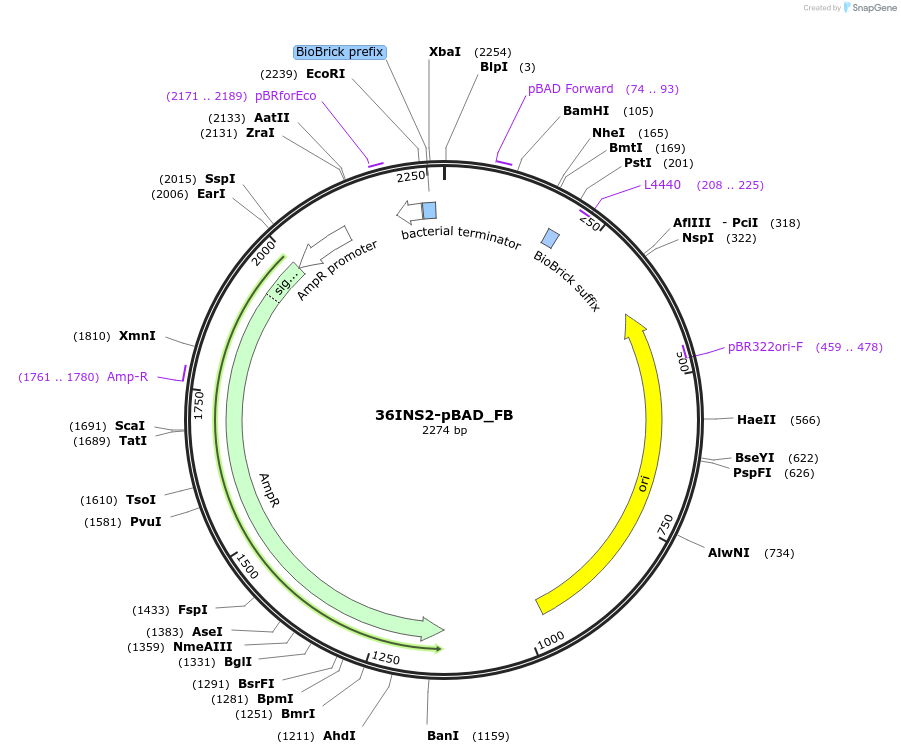
36INS2-pBAD_FB
Plasmid#78676PurposeInsulated MoClo promoter for E. coli. 36nt DNA spacer empirically selected. Modified from BBa_I13453. Has near identical expression as 36INS1-pBAD_AB [F:36INS2-pBAD:B]DepositorInsertInsulated pBAD Promoter
UseSynthetic BiologyAvailable SinceAug. 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
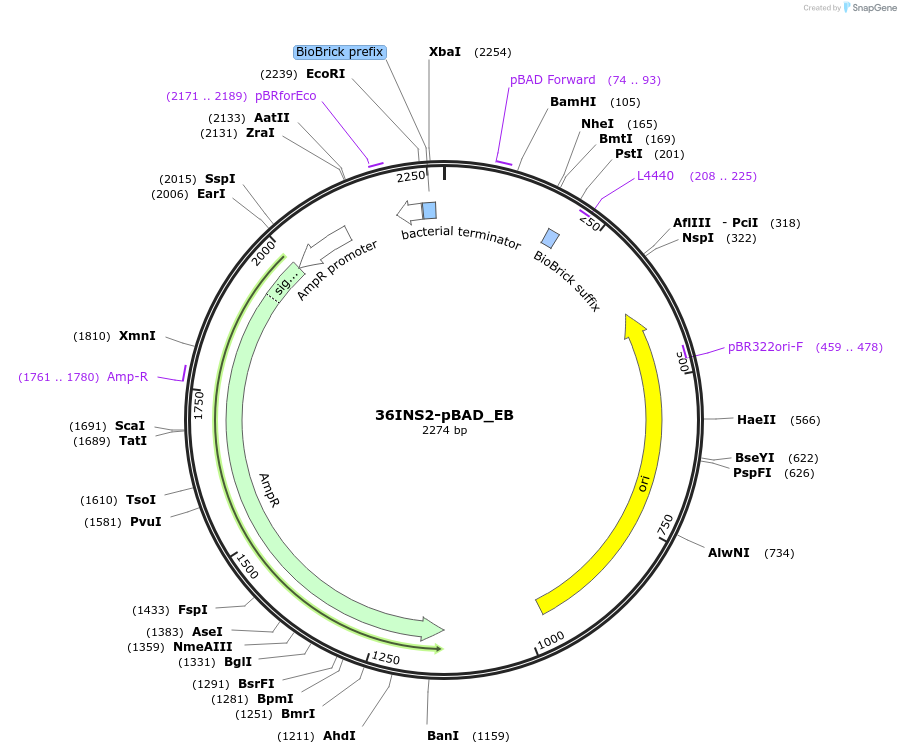
36INS2-pBAD_EB
Plasmid#78675PurposeInsulated MoClo promoter for E. coli. 36nt DNA spacer empirically selected. Modified from BBa_I13453. Has near identical expression as 36INS1-pBAD_AB [E:36INS2-pBAD:B]DepositorInsertInsulated pBAD Promoter
UseSynthetic BiologyAvailable SinceAug. 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
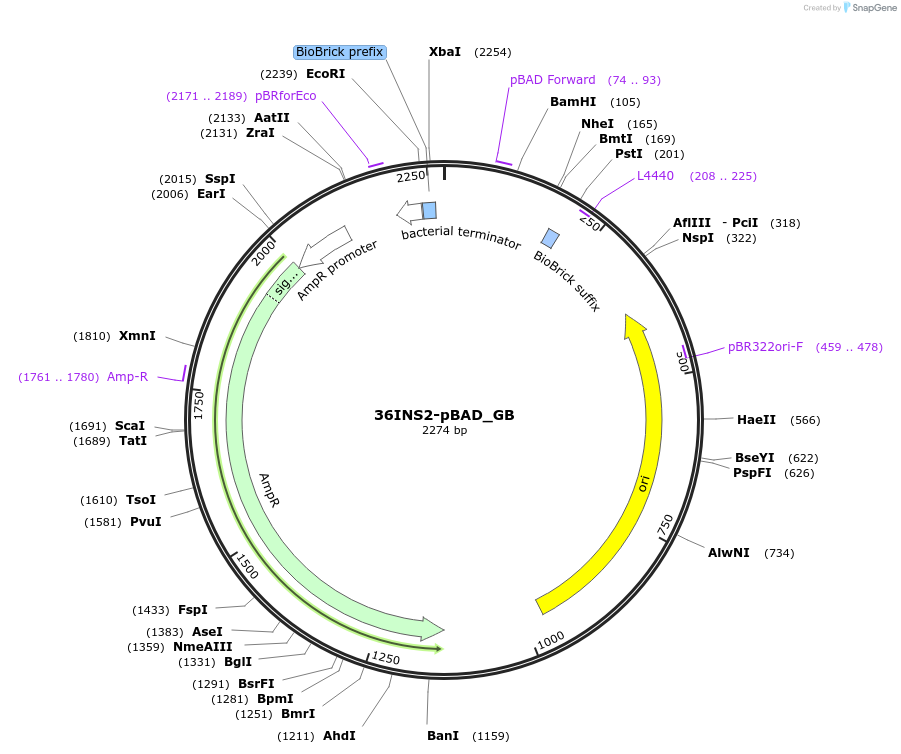
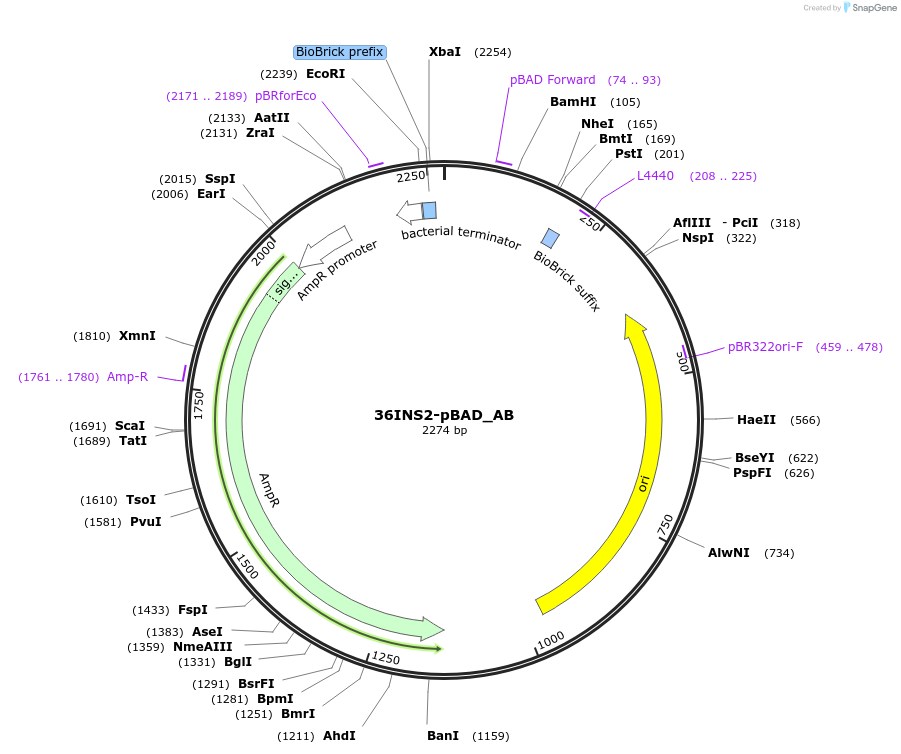
36INS2-pBAD_AB
Plasmid#78674PurposeInsulated MoClo promoter for E. coli. 36nt DNA spacer empirically selected. Modified from BBa_I13453. Has near identical expression as 36INS1-pBAD_AB [A:36INS2-pBAD:B]DepositorInsertInsulated pBAD Promoter
UseSynthetic BiologyAvailable SinceAug. 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
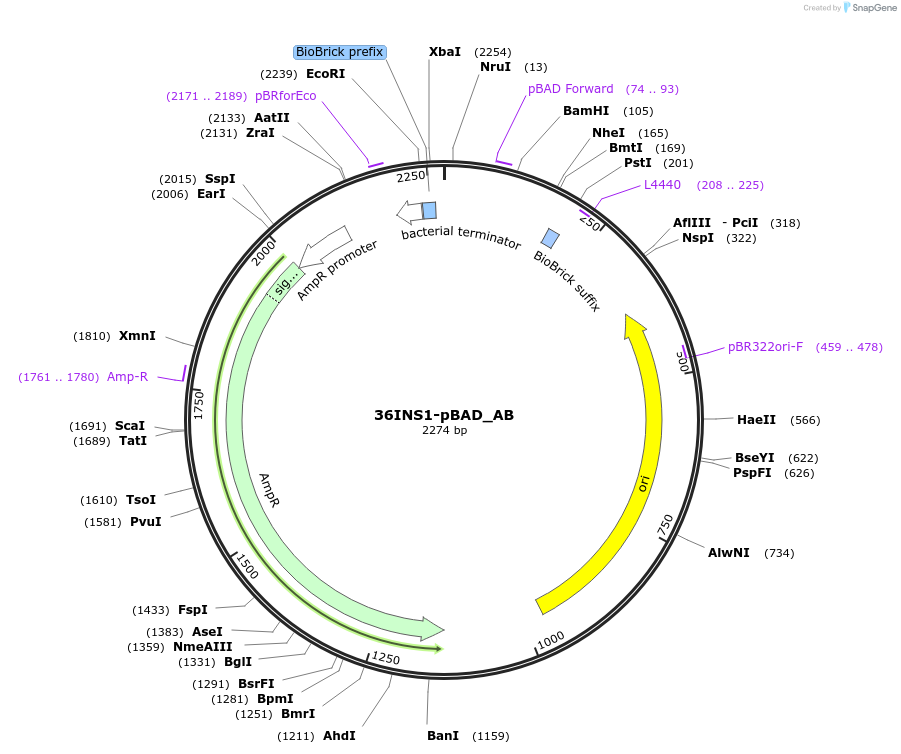
36INS1-pBAD_AB
Plasmid#78670PurposeInsulated MoClo promoter for E. coli. 36nt DNA spacer empirically selected.Modified from BBa_I13453 to adjust spacing in MoClo system. [A:36INS1-pBAD:B]DepositorInsertInsulated pBAD Promoter
UseSynthetic BiologyAvailable SinceAug. 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
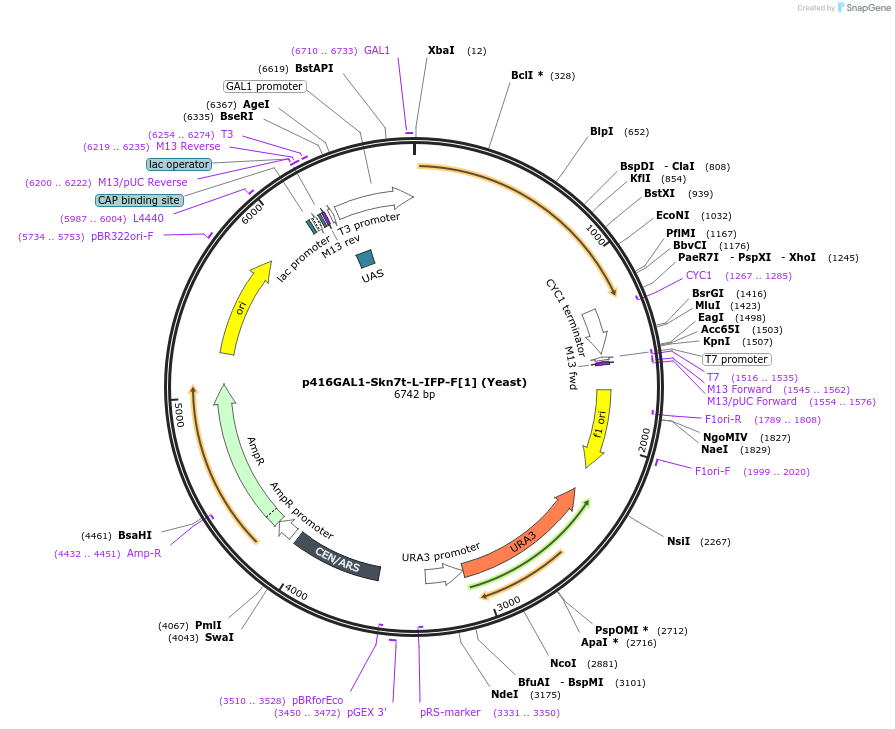
p416GAL1-Skn7t-L-IFP-F[1] (Yeast)
Plasmid#52896PurposeSkn7t replaces the zipper and is fused to the IFP-F[1] through the linker (GGGGS) X 2. Plasmid confers ampicillin resistance to DH5α.DepositorInsertSkn7t-L-IFP-F[1]
TagsL-IFP-F[1]ExpressionYeastPromoterGAL1Available SinceJune 12, 2014AvailabilityAcademic Institutions and Nonprofits only -
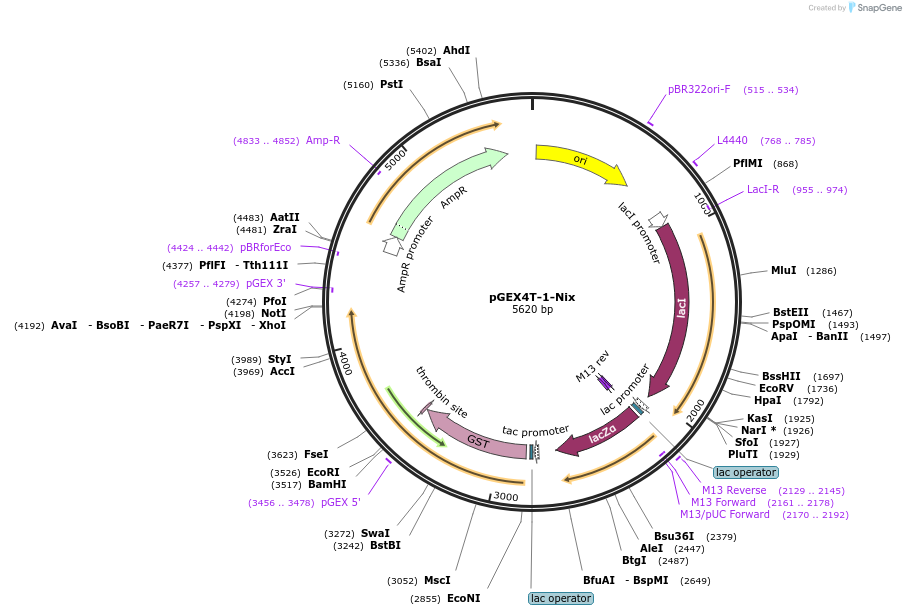
pGEX4T-1-Nix
Plasmid#100759PurposeBacterial expression of GST fused Nix proteinDepositorAvailable SinceSept. 7, 2017AvailabilityAcademic Institutions and Nonprofits only -
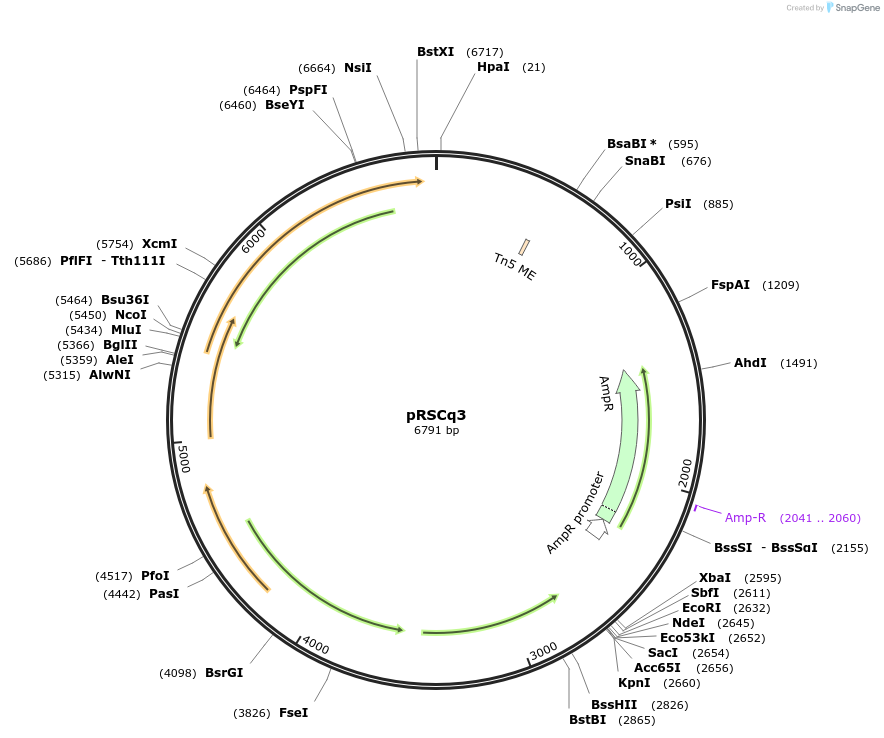
pRSCq3
Plasmid#236106PurposeRalstonia pseudosolanacearum filamentous phage RSCq-based cloning vector carrying ampicillin resistance gene.DepositorTypeEmpty backboneAvailable SinceJan. 7, 2026AvailabilityAcademic Institutions and Nonprofits only -
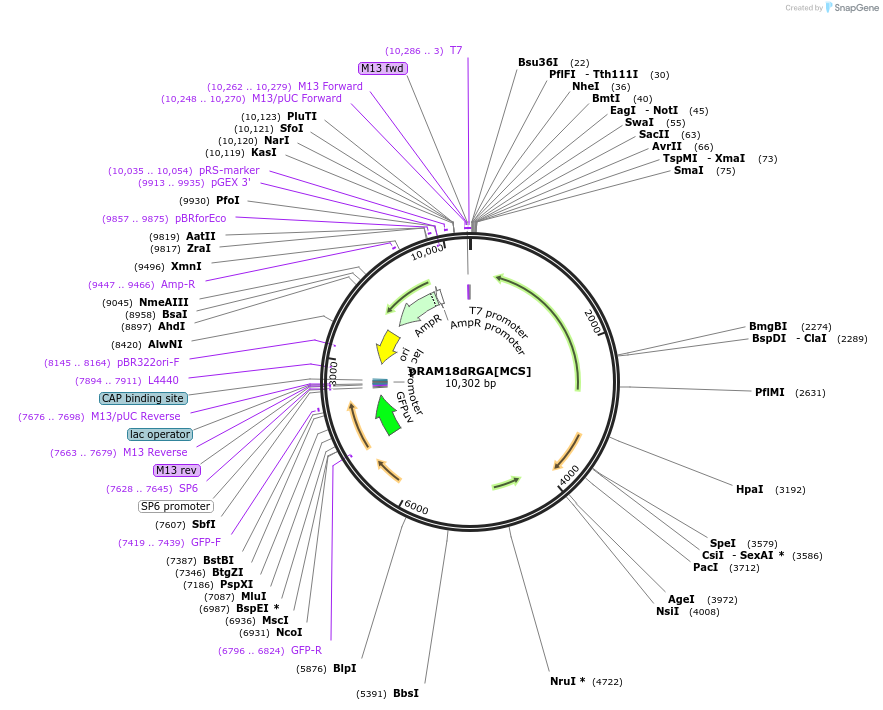
pRAM18dRGA[MCS]
Plasmid#84683PurposeCan be used for the transformation of rickettsiaDepositorTypeEmpty backboneExpressionBacterialAvailable SinceNov. 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
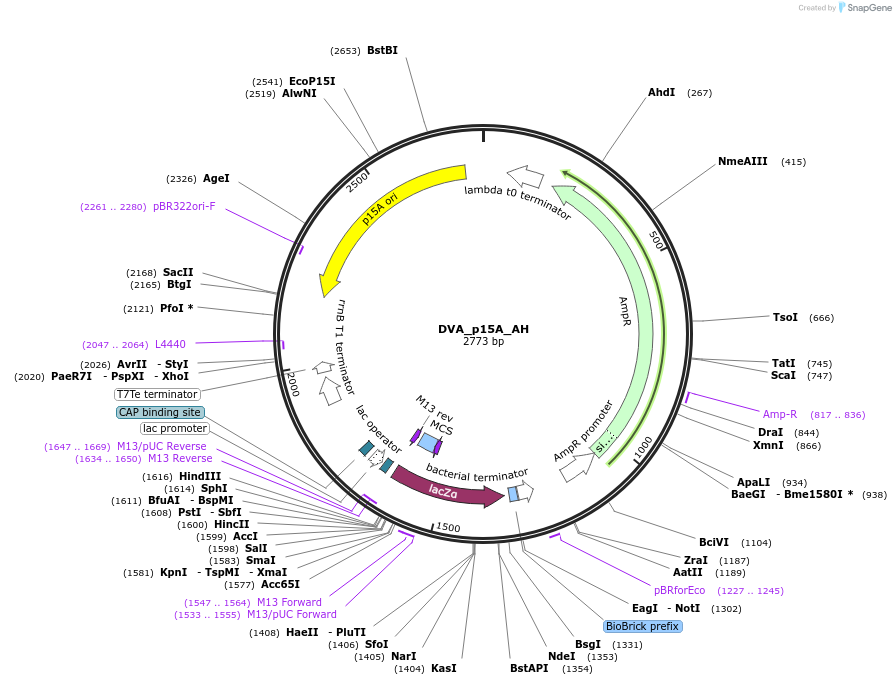
DVA_p15A_AH
Plasmid#217602PurposeDestination vector with ampicillin resistance and p15A origin of replication. Carries A-H CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
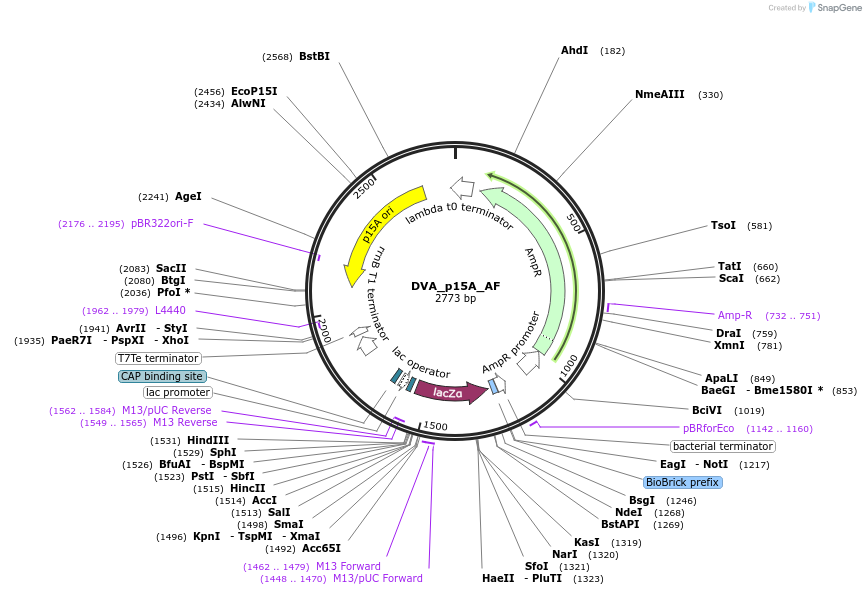
DVA_p15A_AF
Plasmid#217604PurposeDestination vector with ampicillin resistance and p15A origin of replication. Carries A-F CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceAug. 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
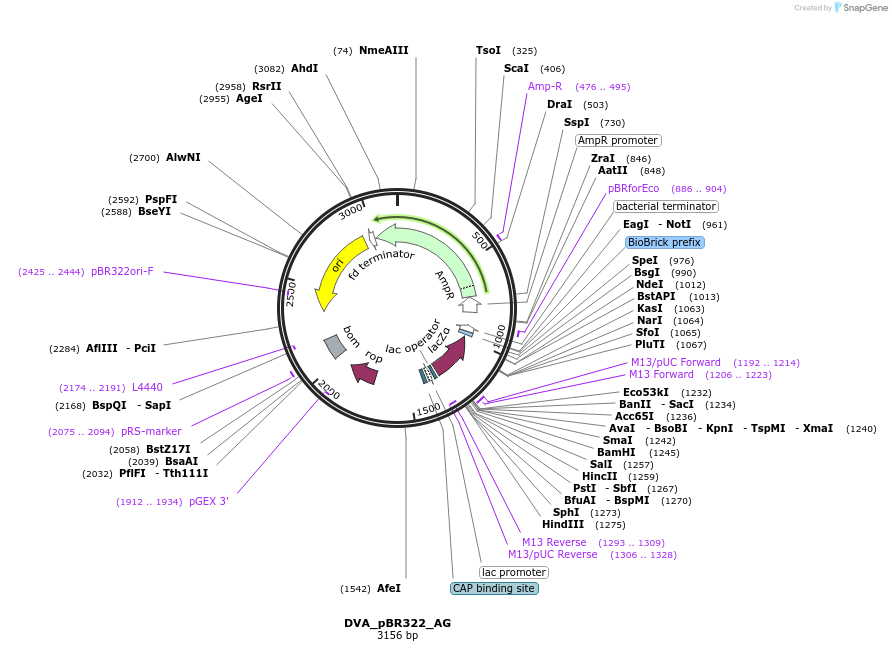
DVA_pBR322_AG
Plasmid#217607PurposeDestination vector with ampicillin resistance and pBR322 origin of replication. Carries A-G CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceAug. 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
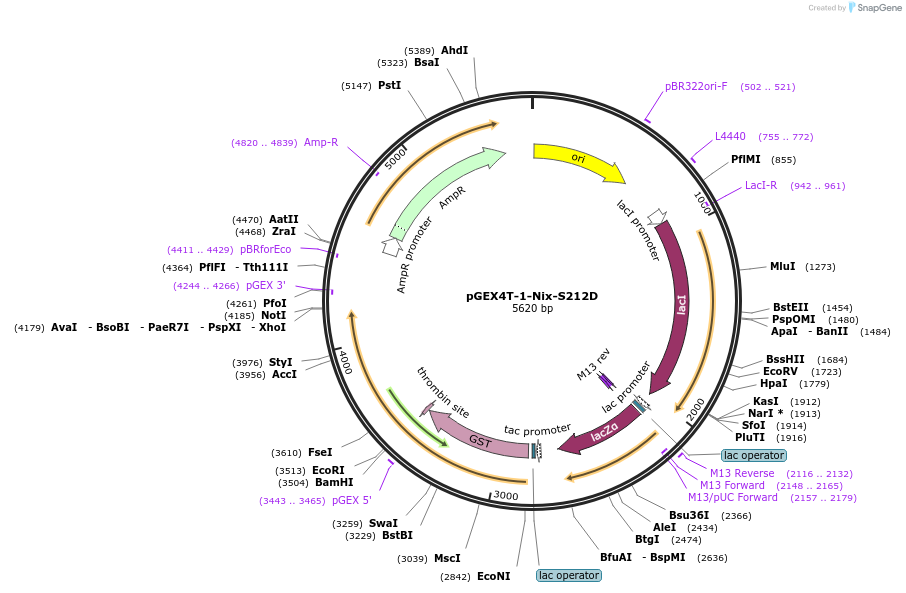
pGEX4T-1-Nix-S212D
Plasmid#100761PurposeBacterial expression of GST fused Nix-S212D proteinDepositorAvailable SinceSept. 7, 2017AvailabilityAcademic Institutions and Nonprofits only -
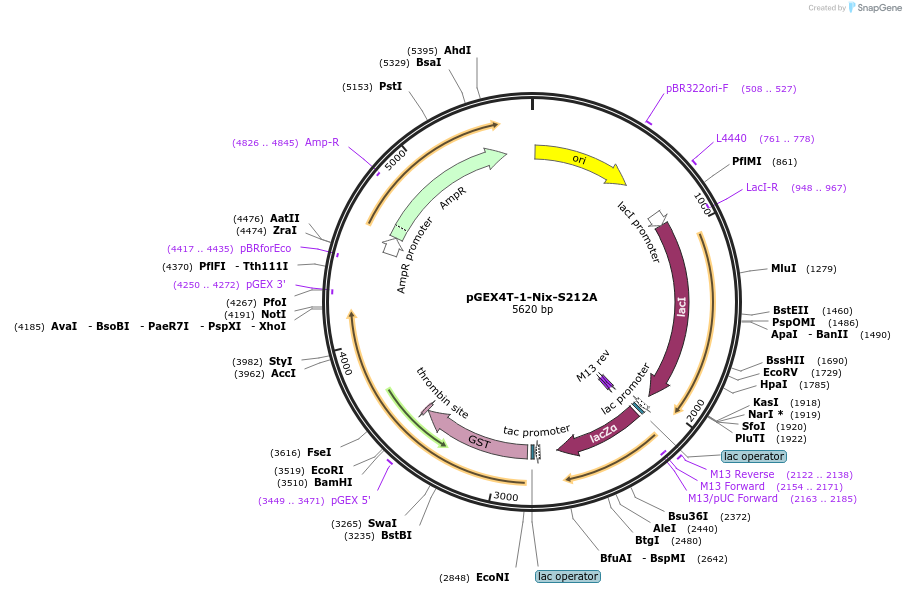
pGEX4T-1-Nix-S212A
Plasmid#100760PurposeBacterial expression of GST fused Nix-S212A proteinDepositorAvailable SinceSept. 7, 2017AvailabilityAcademic Institutions and Nonprofits only -
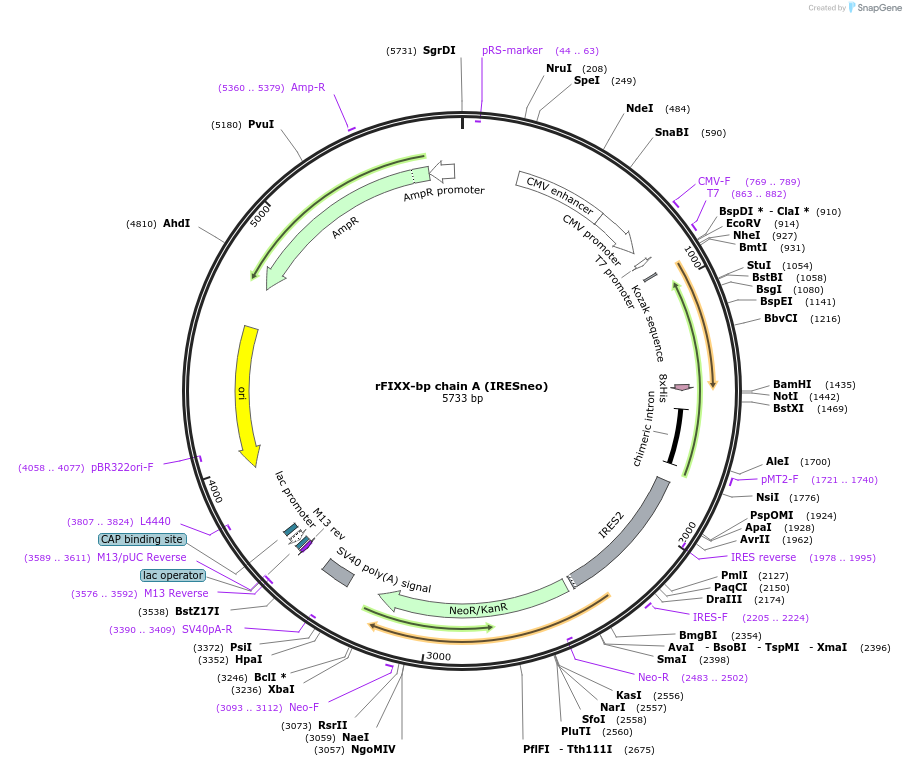
rFIXX-bp chain A (IRESneo)
Plasmid#82450PurposeChain A of the habu snake venom FIX/FX binding protein with a C-terminal polyhistidine-tag. Cloned into pIRESneo3 (Ampicillin) via 5' NheI and 3' BamHI; IRES vector conferring neomycin resistance.DepositorInsertrFIXX-bp chain A
TagsPolyhistidineExpressionBacterial and MammalianPromoterT7 RNA polymerase promoterAvailable SinceNov. 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
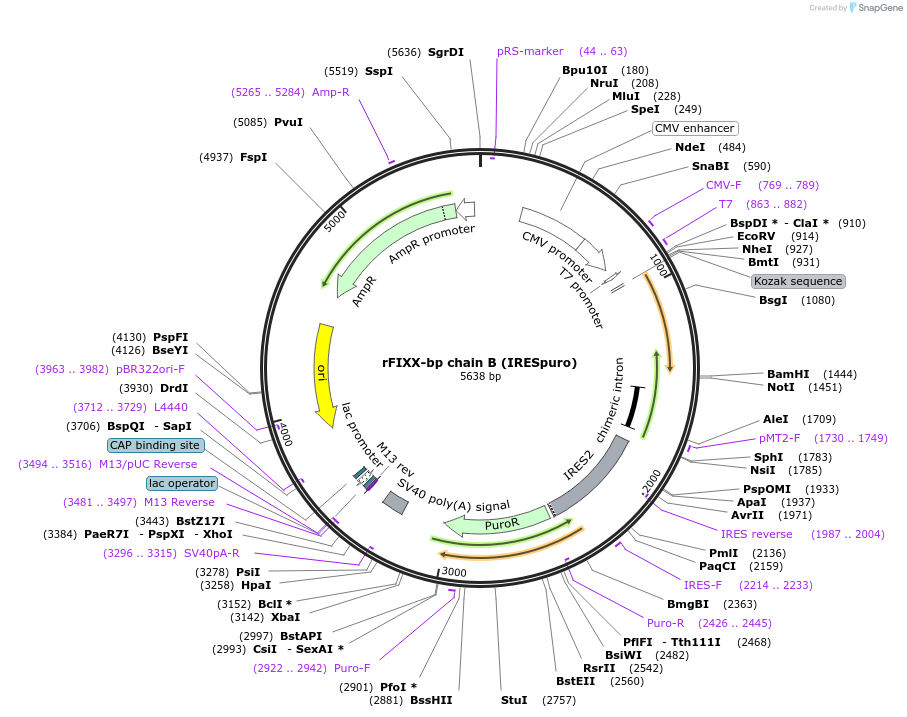
rFIXX-bp chain B (IRESpuro)
Plasmid#82451PurposeChain B of the habu snake venom FIX/FX binding protein with C-terminal biotin-tag. Cloned into pIRESneo3 (Ampicillin) via 5' NheI and 3' BamHI; IRES vector conferring puromycin resistance.DepositorInsertrFIXX-bp chain B
TagsBiotinExpressionBacterial and MammalianPromoterT7 RNA polymerase promoterAvailable SinceNov. 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
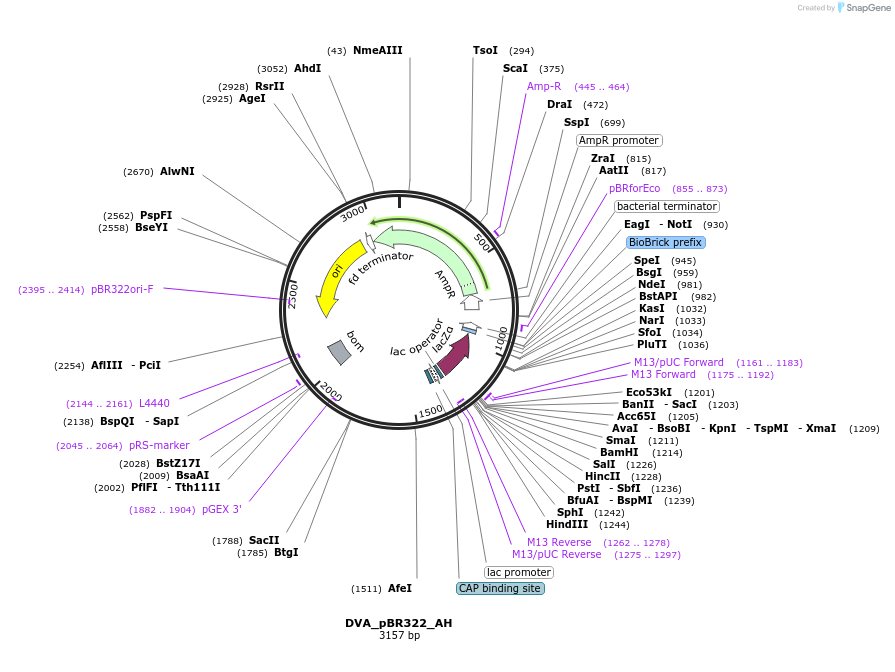
DVA_pBR322_AH
Plasmid#217606PurposeDestination vector with ampicillin resistance and pBR322 origin of replication. Carries A-H CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceSept. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
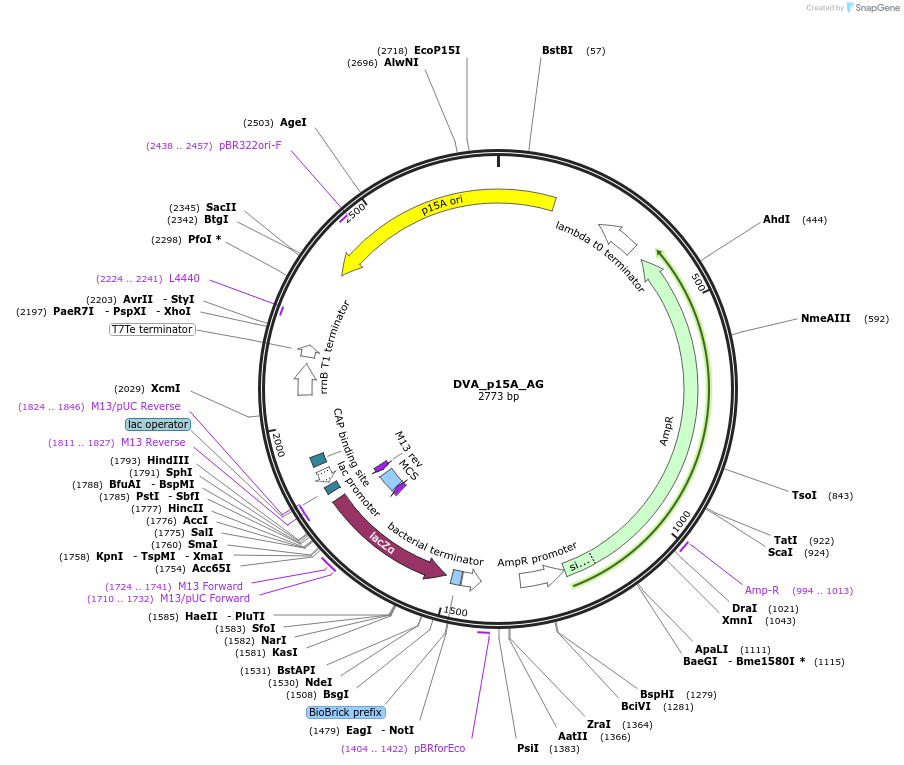
DVA_p15A_AG
Plasmid#217603PurposeDestination vector with ampicillin resistance and p15A origin of replication. Carries A-G CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceAug. 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
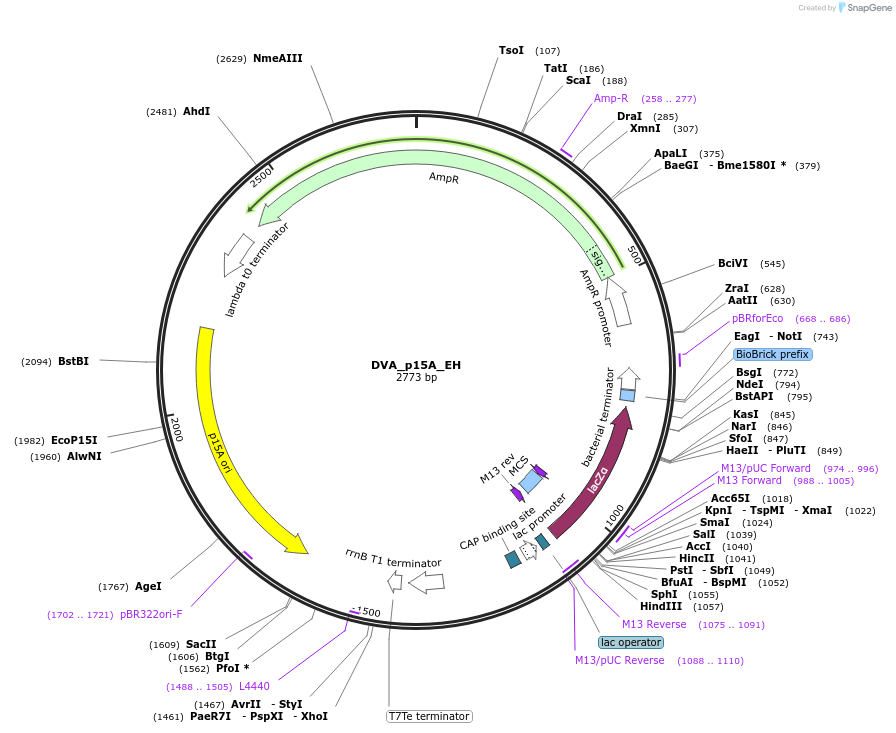
DVA_p15A_EH
Plasmid#217605PurposeDestination vector with ampicillin resistance and p15A origin of replication. Carries E-H CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceAug. 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
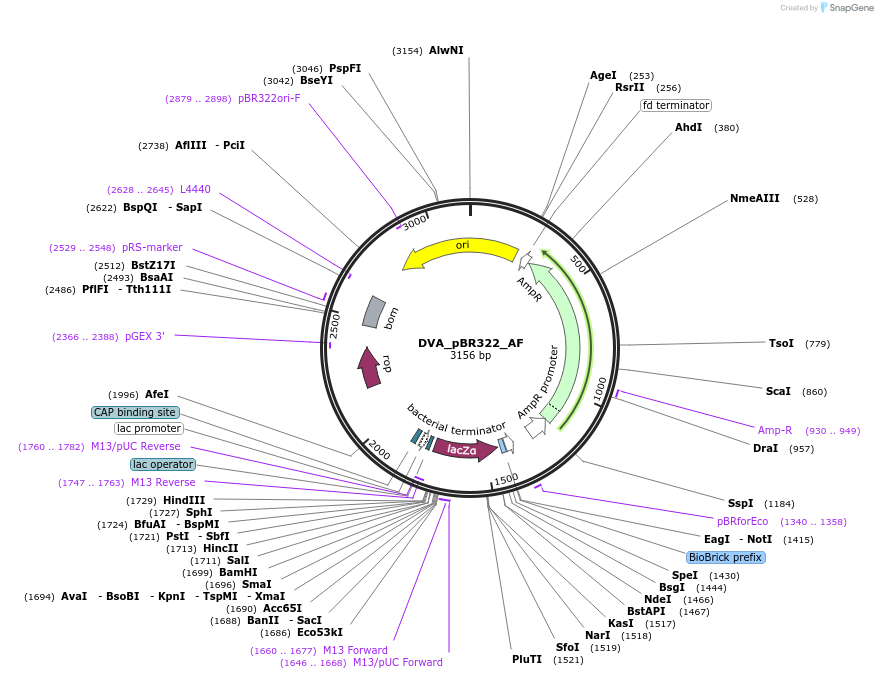
DVA_pBR322_AF
Plasmid#217608PurposeDestination vector with ampicillin resistance and pBR322 origin of replication. Carries A-F CIDAR MoClo overhangs for level 0 or 2 construct cloning.DepositorTypeEmpty backboneUseSynthetic BiologyMutationNoneAvailable SinceAug. 9, 2024AvailabilityAcademic Institutions and Nonprofits only