We narrowed to 3,450 results for: biorxiv
-
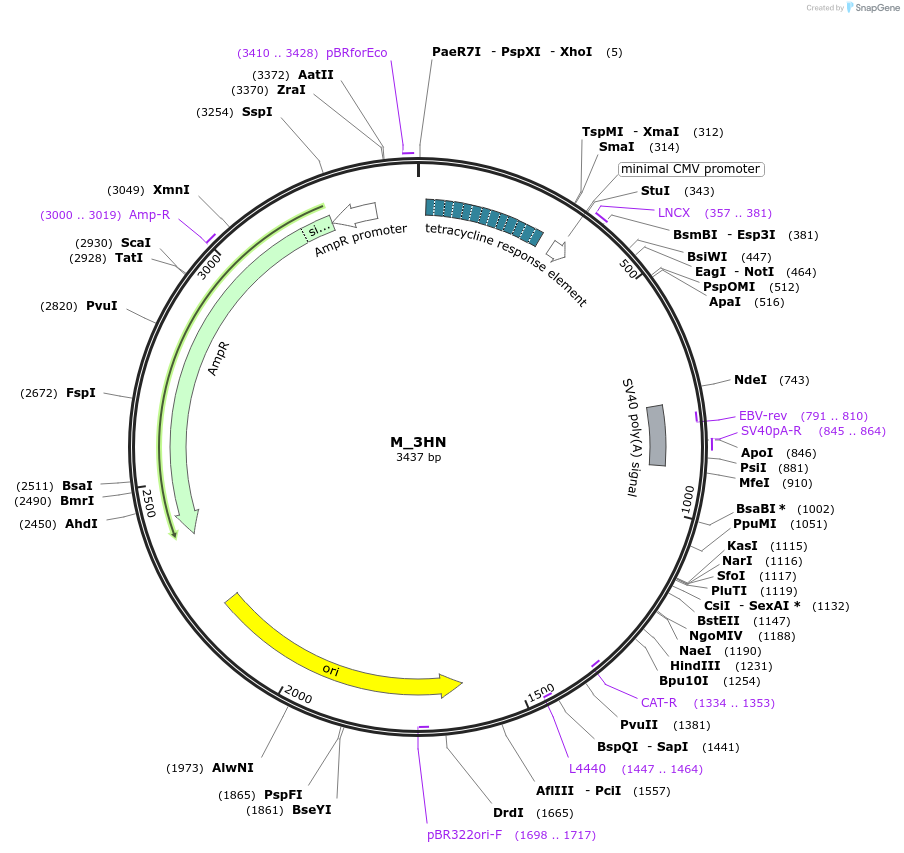
Plasmid#249751PurposeMinimal synthetic construct for assessing speckle localization containing a 3' splice site and an HNRNP motif-rich regionDepositorInsertM_3HN
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 11, 2026AvailabilityAcademic Institutions and Nonprofits only -
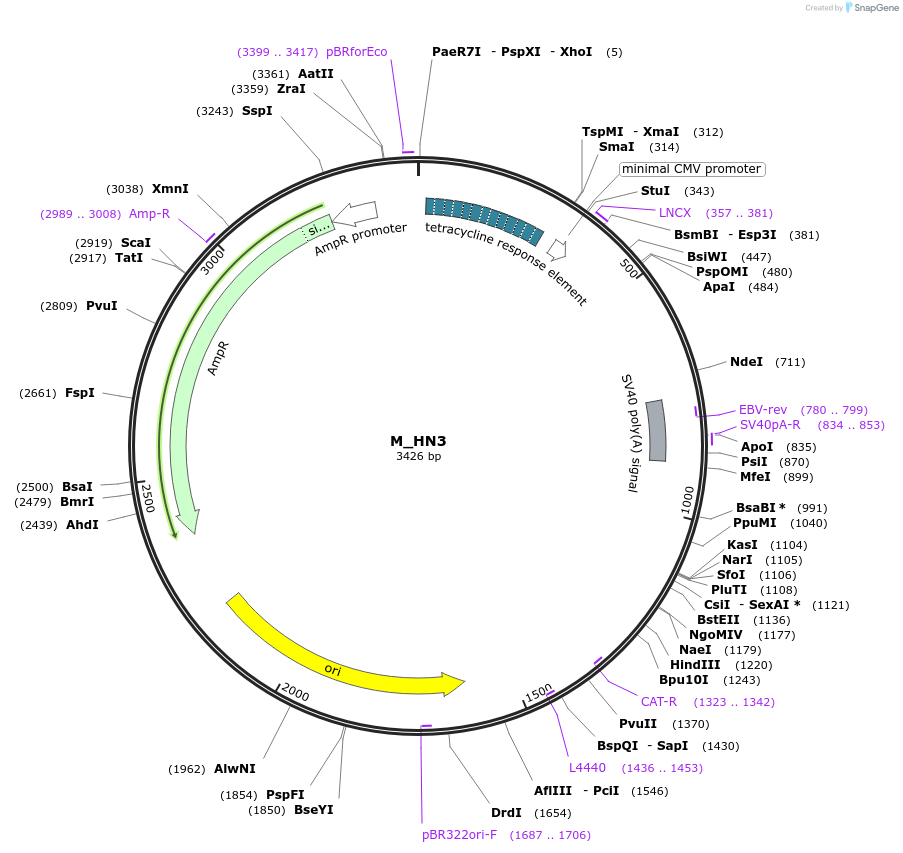
M_HN3
Plasmid#249756PurposeMinimal synthetic construct for assessing speckle localization containing an HNRNP motif-rich region and a 3' splice siteDepositorInsertM_HN3
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 11, 2026AvailabilityAcademic Institutions and Nonprofits only -
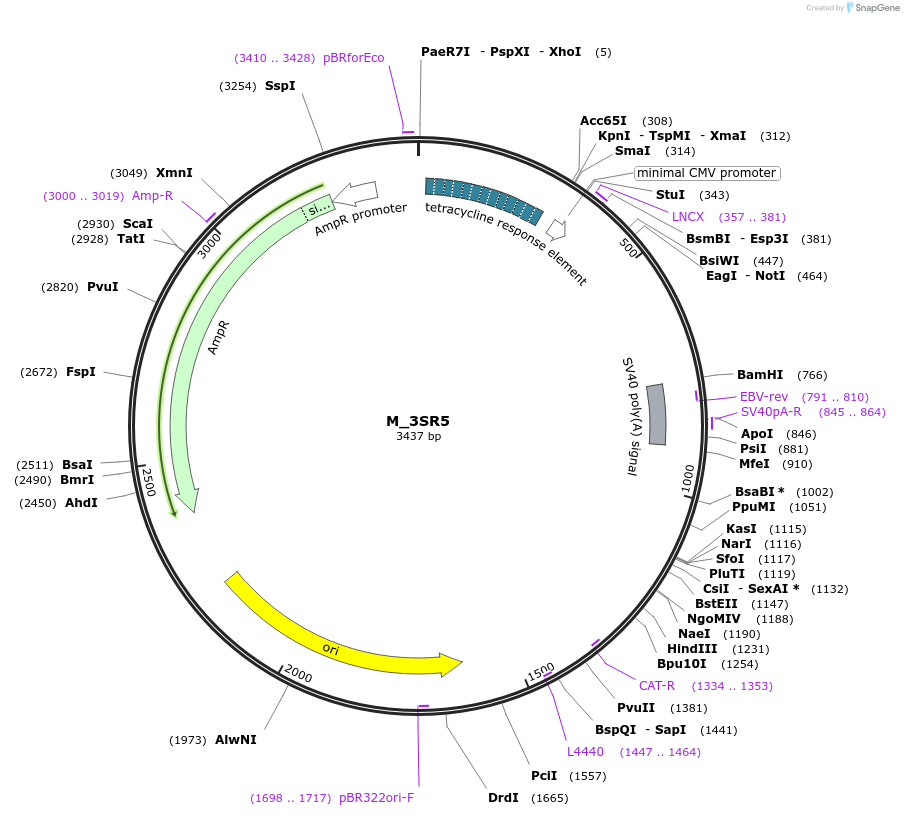
M_3SR5
Plasmid#249745PurposeMinimal synthetic construct for assessing speckle localization containing a 3' splice site, SR motif-rich region, and a 5' splice siteDepositorInsertM_3SR5
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 11, 2026AvailabilityAcademic Institutions and Nonprofits only -
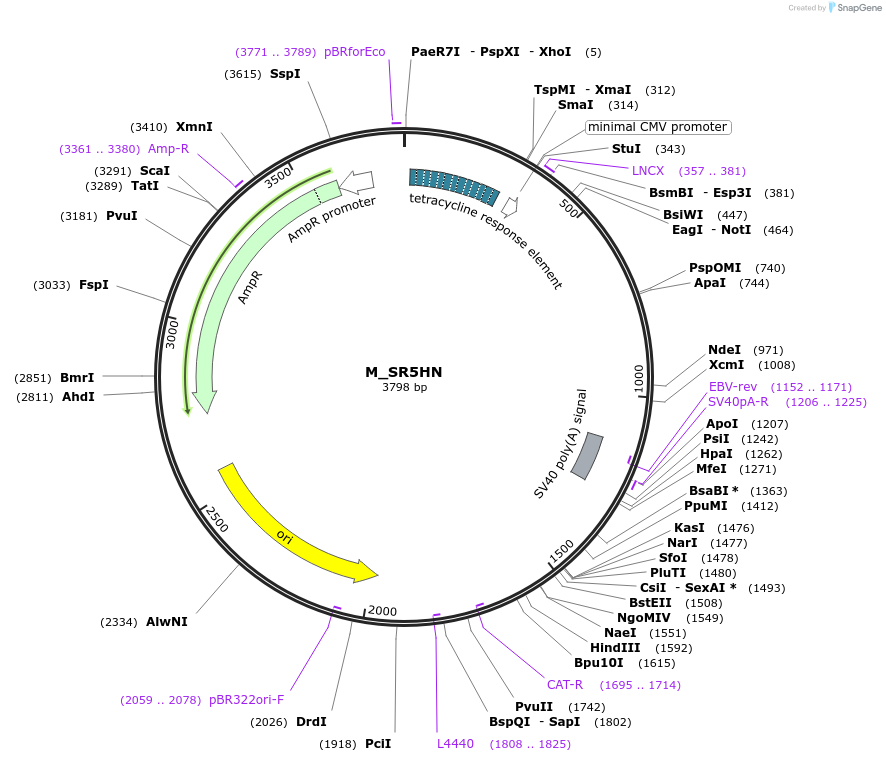
M_SR5HN
Plasmid#249739PurposeMinimal synthetic construct for assessing speckle localization containing an SR motif-rich region, 5' splice site, and an HNRNP motif-rich regionDepositorInsertM_SR5HN
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 11, 2026AvailabilityAcademic Institutions and Nonprofits only -
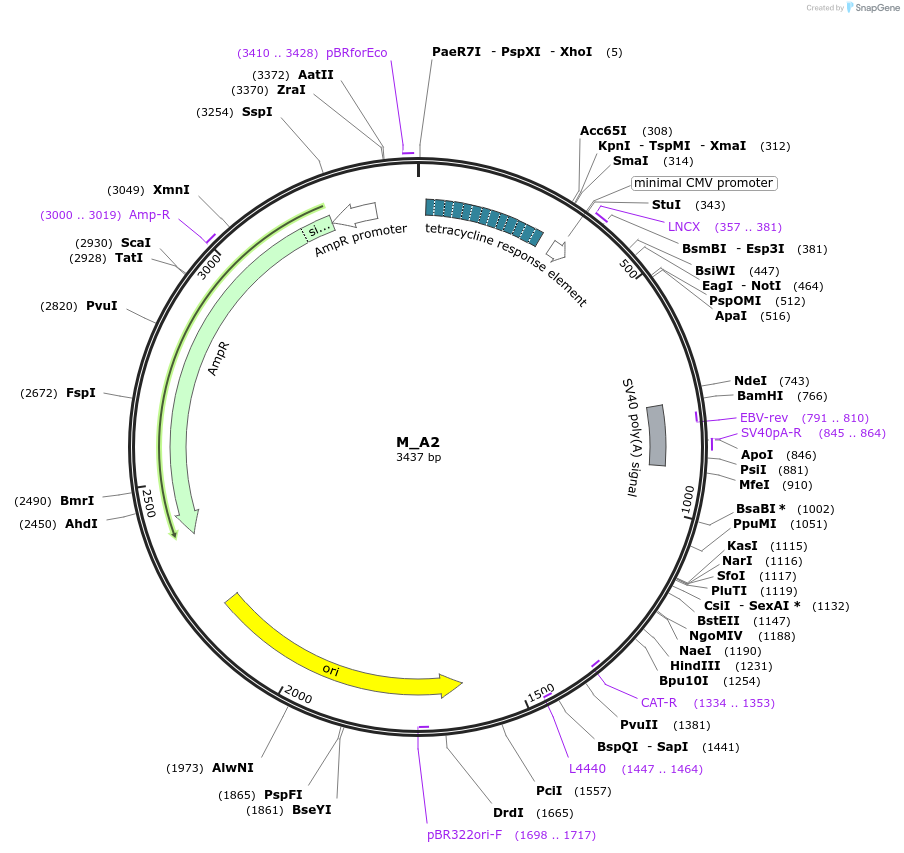
M_A2
Plasmid#249758PurposeMinimal synthetic construct for assessing speckle localization, part of A trajectory series (starting with M_3HN5 and ending in M_3SR5)DepositorInsertM_A2
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 10, 2026AvailabilityAcademic Institutions and Nonprofits only -
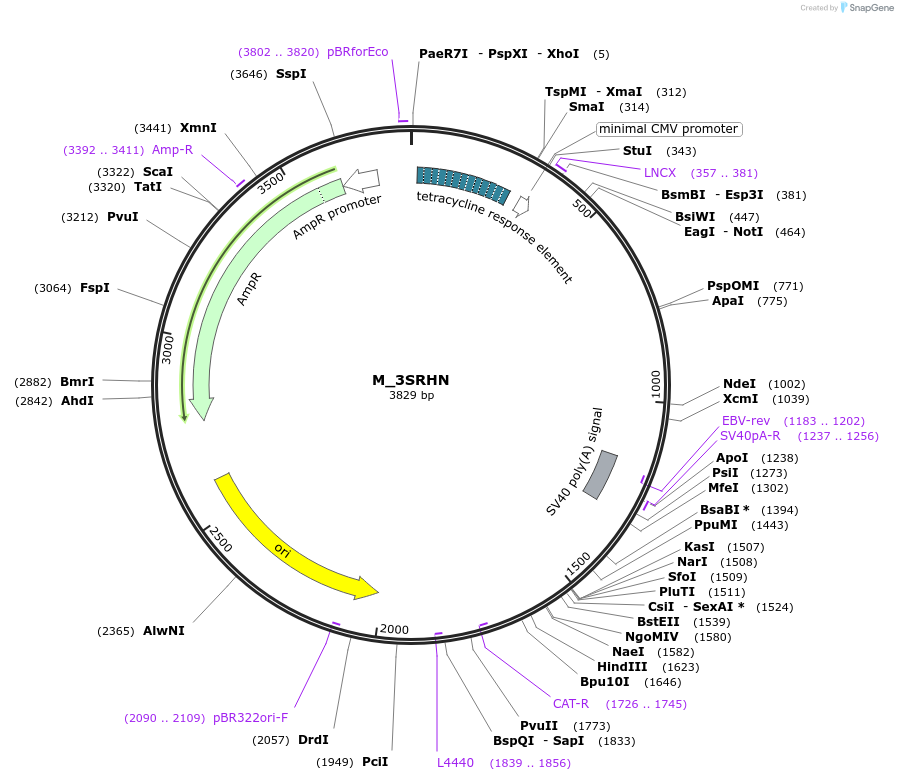
M_3SRHN
Plasmid#249738PurposeMinimal synthetic construct for assessing speckle localization containing a 3' splice site, SR motif-rich region, and an HNRNP motif-rich regionDepositorInsertM_3SRHN
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
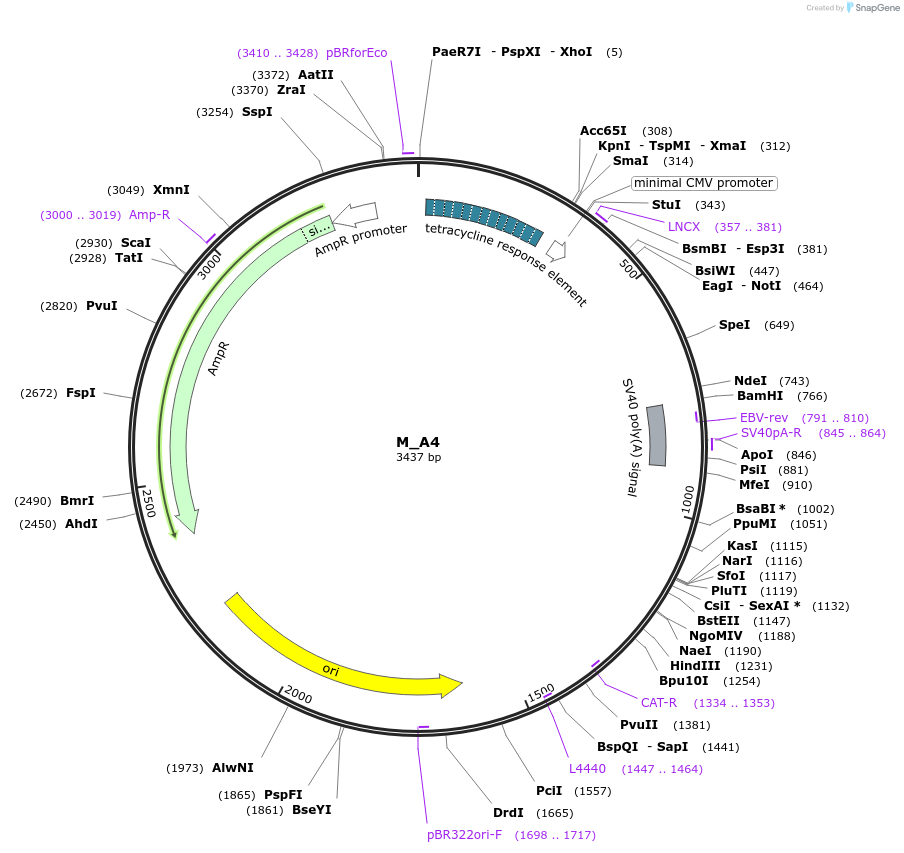
M_A4
Plasmid#249760PurposeMinimal synthetic construct for assessing speckle localization, part of A trajectory series (starting with M_3HN5 and ending in M_3SR5)DepositorInsertM_A4
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
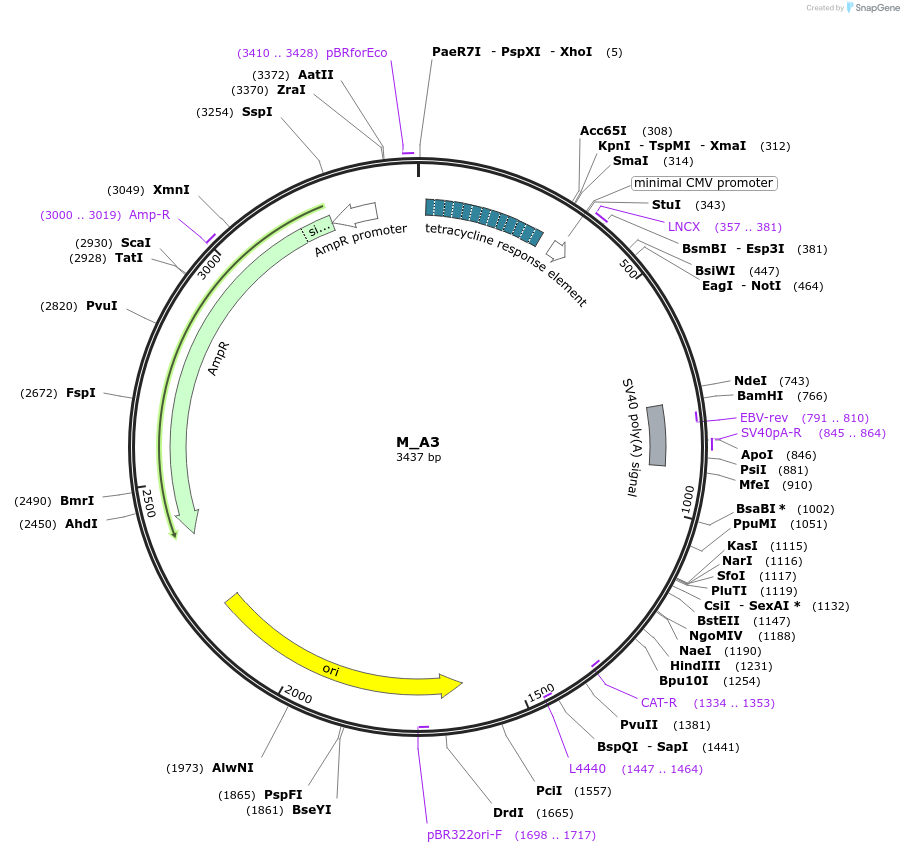
M_A3
Plasmid#249759PurposeMinimal synthetic construct for assessing speckle localization, part of A trajectory series (starting with M_3HN5 and ending in M_3SR5)DepositorInsertM_A3
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
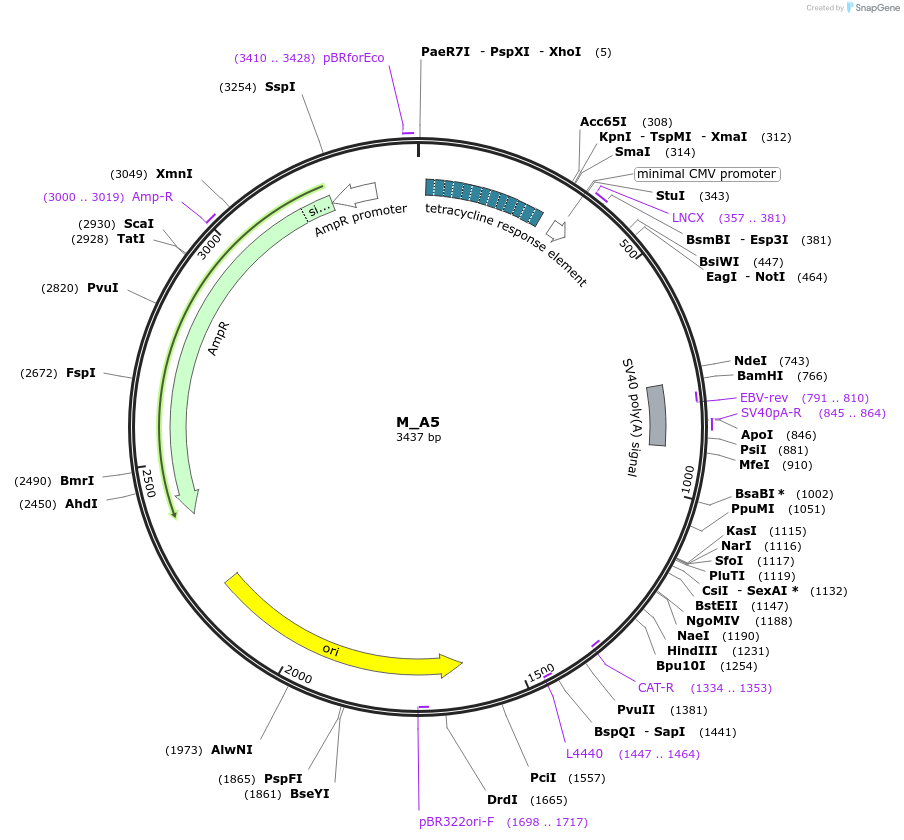
M_A5
Plasmid#249761PurposeMinimal synthetic construct for assessing speckle localization, part of A trajectory series (starting with M_3HN5 and ending in M_3SR5)DepositorInsertM_A5
ExpressionMammalianPromoterTRE+CMVAvailable SinceMarch 6, 2026AvailabilityAcademic Institutions and Nonprofits only -
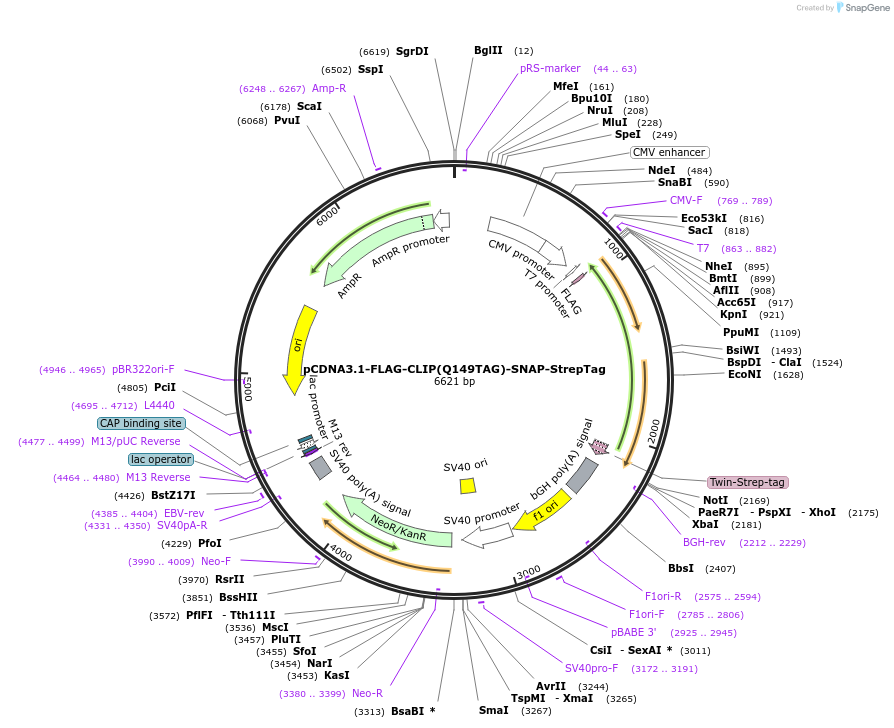
pCDNA3.1-FLAG-CLIP(Q149TAG)-SNAP-StrepTag
Plasmid#248185PurposeFor the mammalian expression and non-cannonical labelling of FLAG-CLIP(Q149TAG)-SNAP-StrepTag fusion protein.DepositorInsertFLAG-CLIP(Q149TAG)-SNAP-StrepTag
Use1257TagsFLAG, CLIP and SNAP, StrepTagExpressionMammalianMutationQ149TAGPromoterCMV & T7Available SinceMarch 6, 2026AvailabilityAcademic Institutions and Nonprofits only -
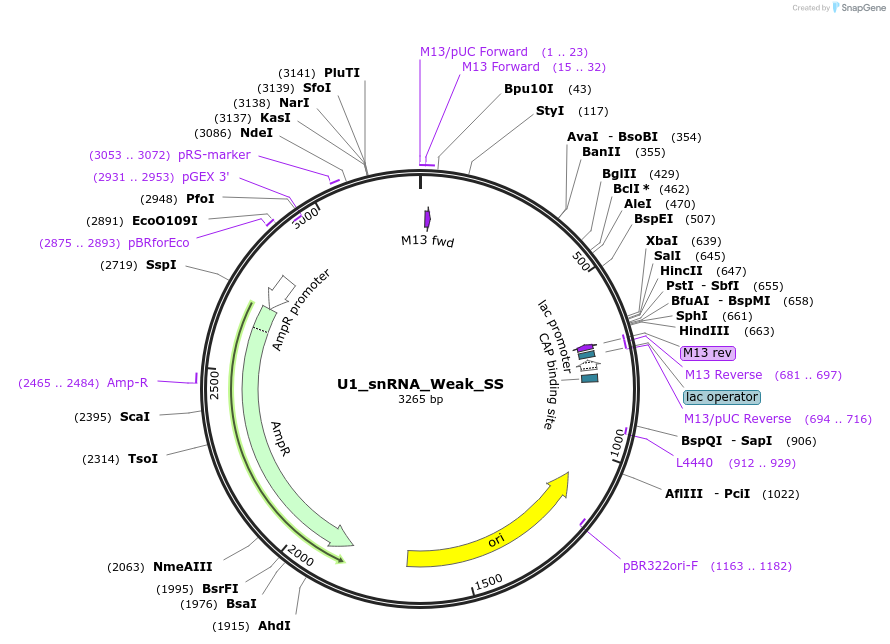
U1_snRNA_Weak_SS
Plasmid#249810PurposeMutated human U1 snRNA complementary to a weak 5' splice site sequence CAGgtacgaDepositorInsertMutated human U1 snRNA
ExpressionMammalianPromoterU1Available SinceMarch 4, 2026AvailabilityAcademic Institutions and Nonprofits only -
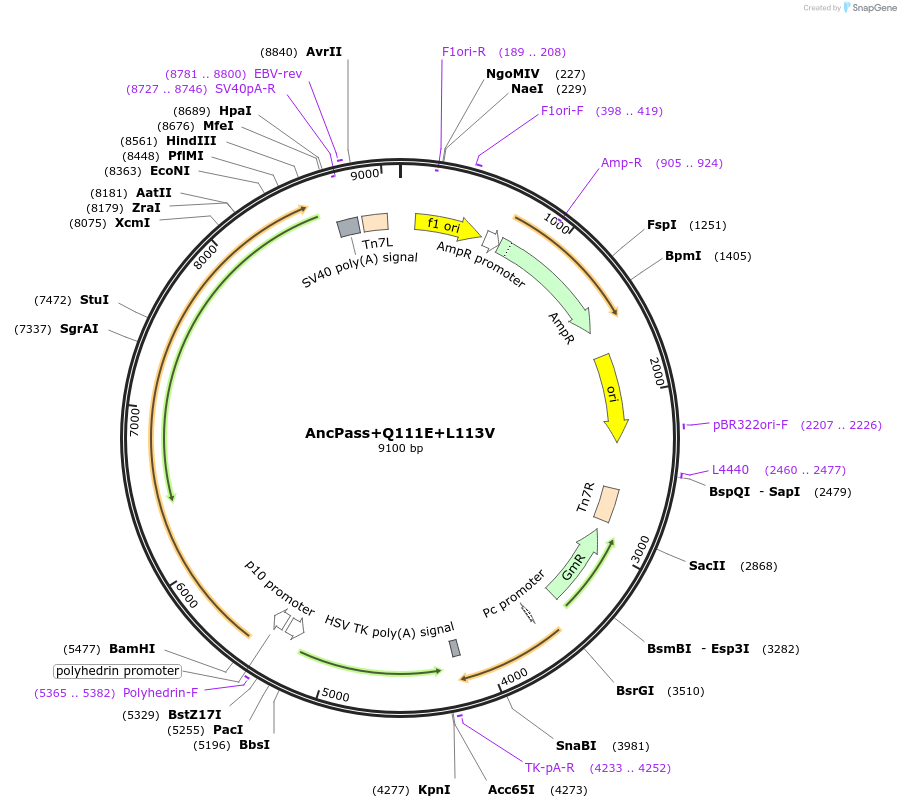
AncPass+Q111E+L113V
Plasmid#196470PurposeExpresses Na,K-ATPase (ATP1A1 and ATP1B1 in pFastBac Dual vector) in insect cellsDepositorInsertsATP1A1
ATP1B1
ExpressionInsectPromoterPH and p10Available SinceFeb. 19, 2026AvailabilityAcademic Institutions and Nonprofits only -
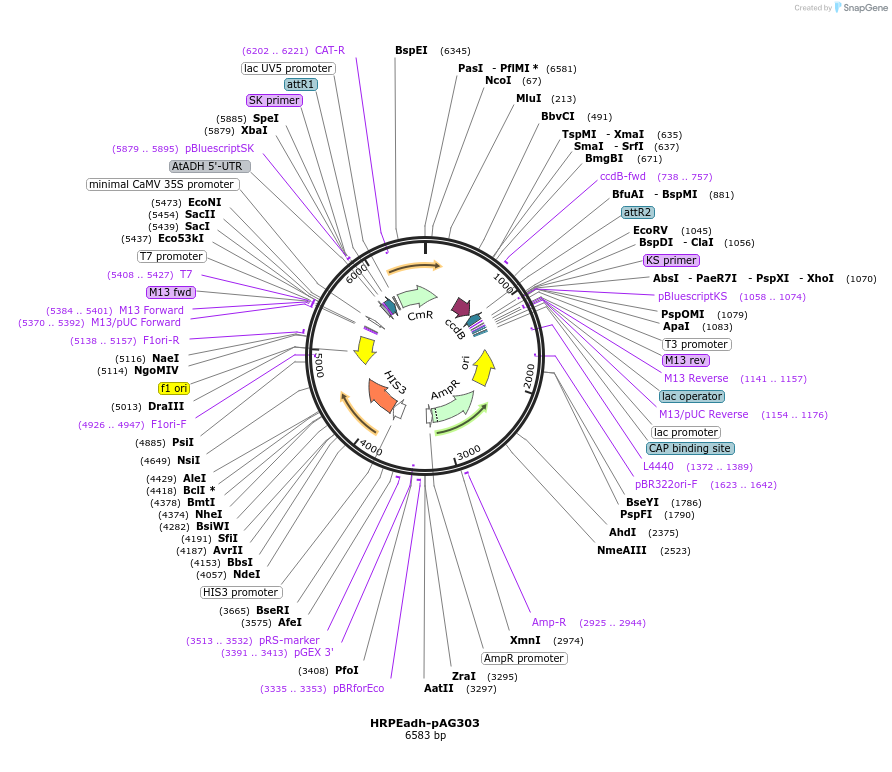
HRPEadh-pAG303
Plasmid#249735PurposeDerived from pAG303GPD-ccdb (#14134); GPD replaced with a synthetic 5xHRPE-ADHutr sequence for orthogonal activation by plant RAP2.12-based TFs.DepositorTypeEmpty backboneExpressionYeastAvailable SinceFeb. 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
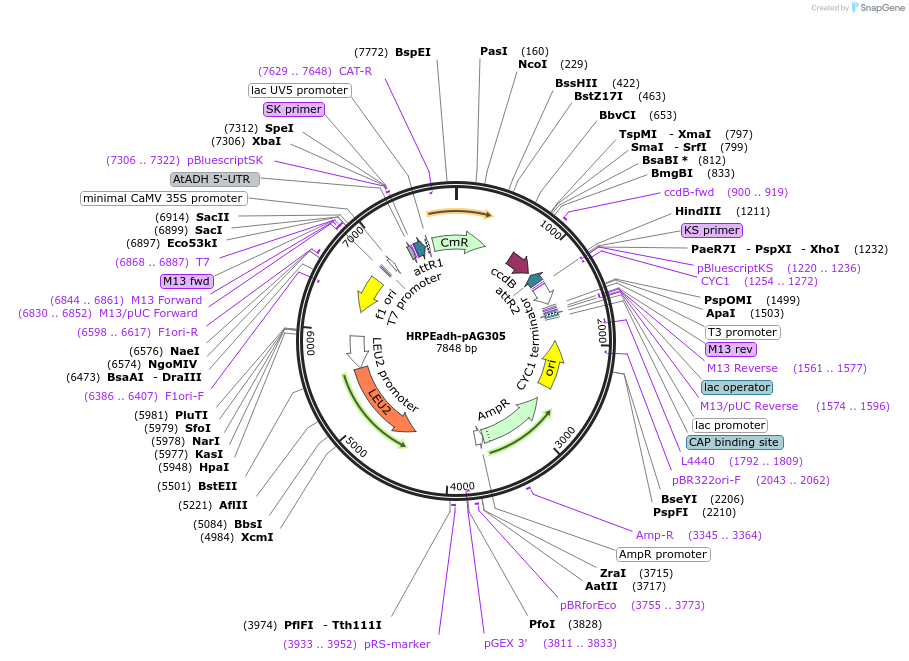
HRPEadh-pAG305
Plasmid#249734PurposeDerived from pAG305GPD-ccdb (#14138); GPD replaced with a synthetic 5xHRPE-ADHutr sequence for orthogonal activation by plant RAP2.12-based TFs.DepositorTypeEmpty backboneExpressionYeastAvailable SinceFeb. 5, 2026AvailabilityAcademic Institutions and Nonprofits only -
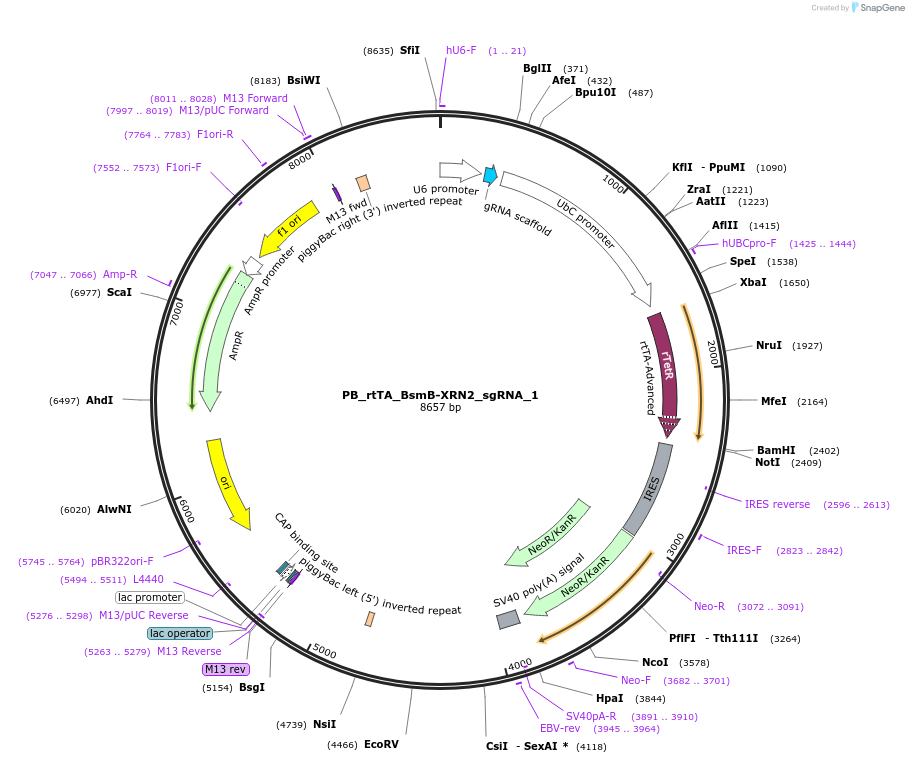
PB_rtTA_BsmB-XRN2_sgRNA_1
Plasmid#242895PurposePiggyBac cargo vector with XRN2 sgRNA 1 for dox-inducible knockdownDepositorAvailable SinceFeb. 5, 2026AvailabilityAcademic Institutions and Nonprofits only -
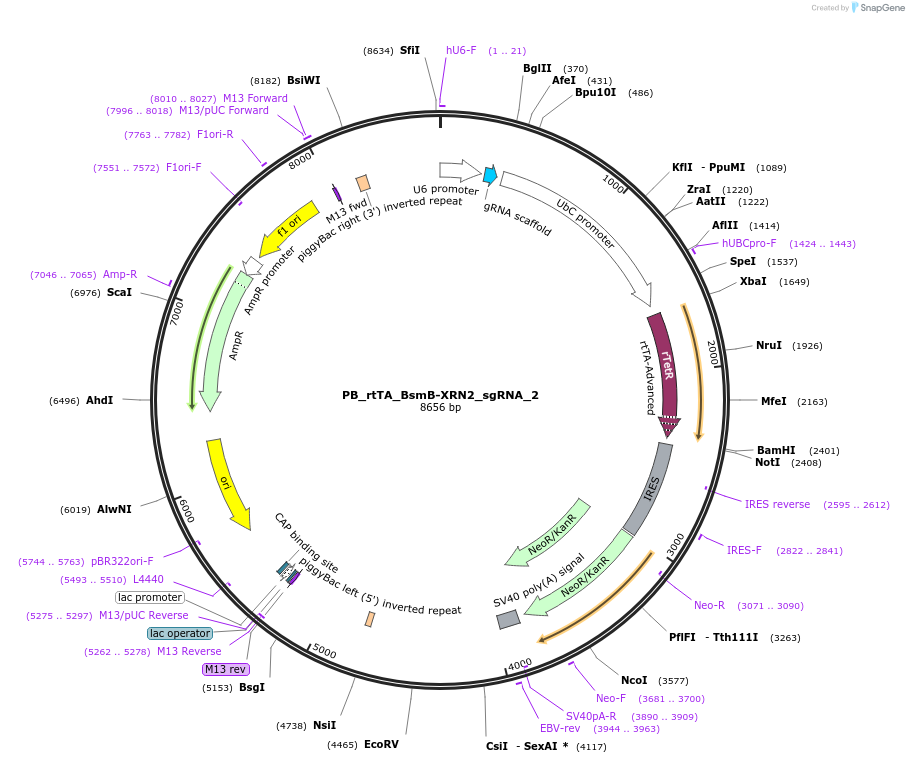
PB_rtTA_BsmB-XRN2_sgRNA_2
Plasmid#242896PurposePiggyBac cargo vector with XRN2 sgRNA 2 for dox-inducible knockdownDepositorAvailable SinceFeb. 5, 2026AvailabilityAcademic Institutions and Nonprofits only -
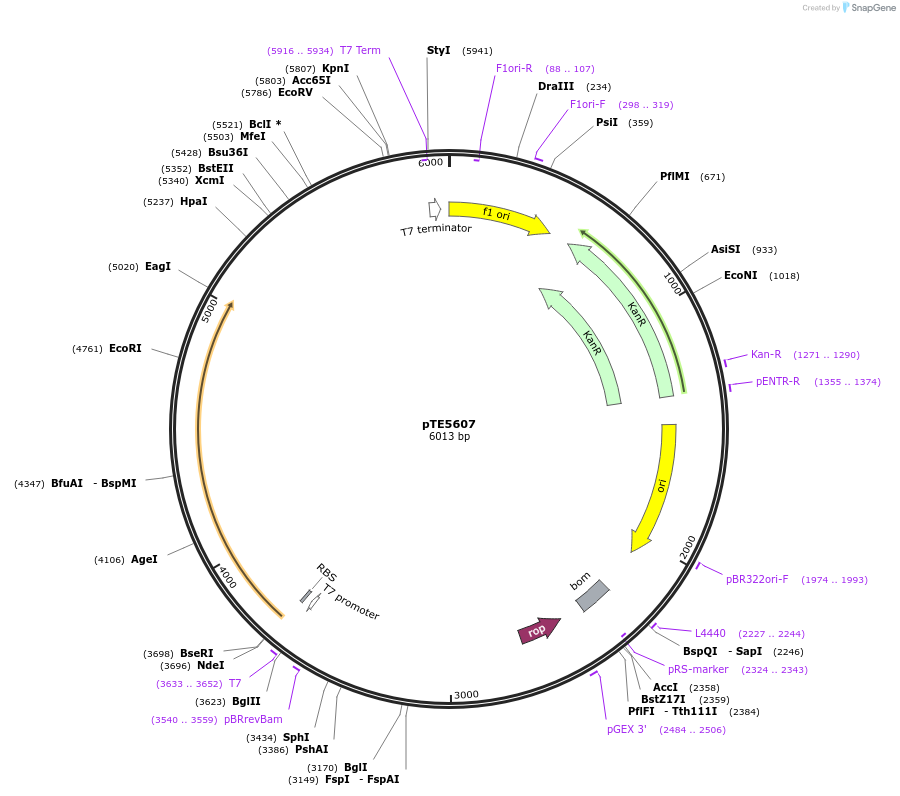
pTE5607
Plasmid#250513PurposepSecYEG; cell-free expression of functional SecYEGDepositorInsertT7-secYEG
ExpressionBacterialPromoterT7Available SinceFeb. 3, 2026AvailabilityAcademic Institutions and Nonprofits only -
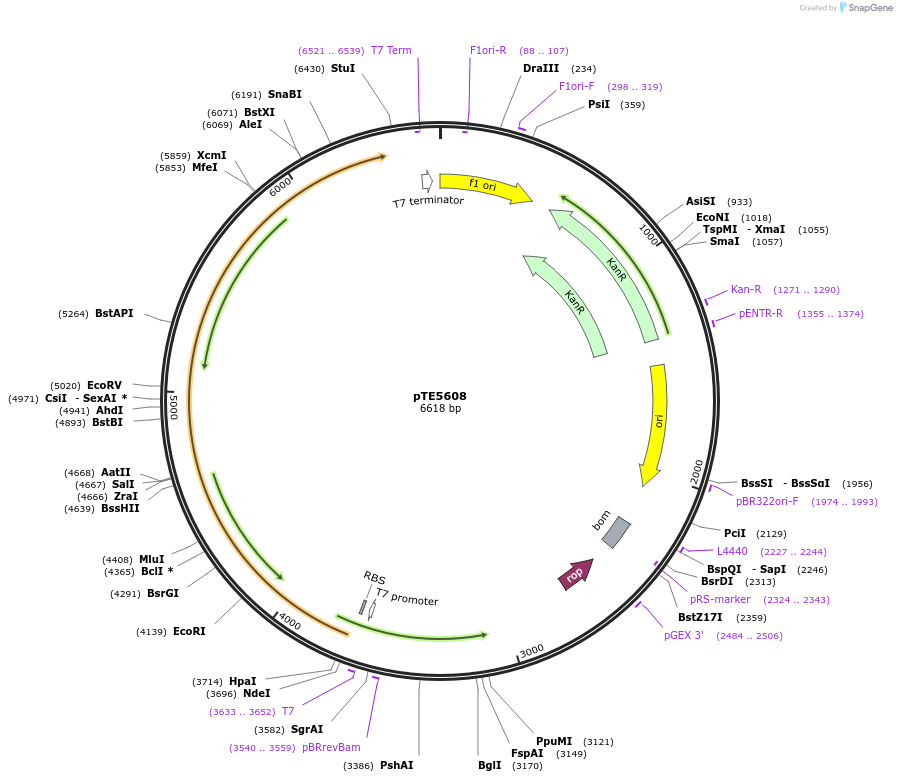
pTE5608
Plasmid#250514PurposeCell-free expression of SecA motor protein for translocation through SecYEGDepositorInsertT7-secA
ExpressionBacterialPromoterT7Available SinceFeb. 3, 2026AvailabilityAcademic Institutions and Nonprofits only -
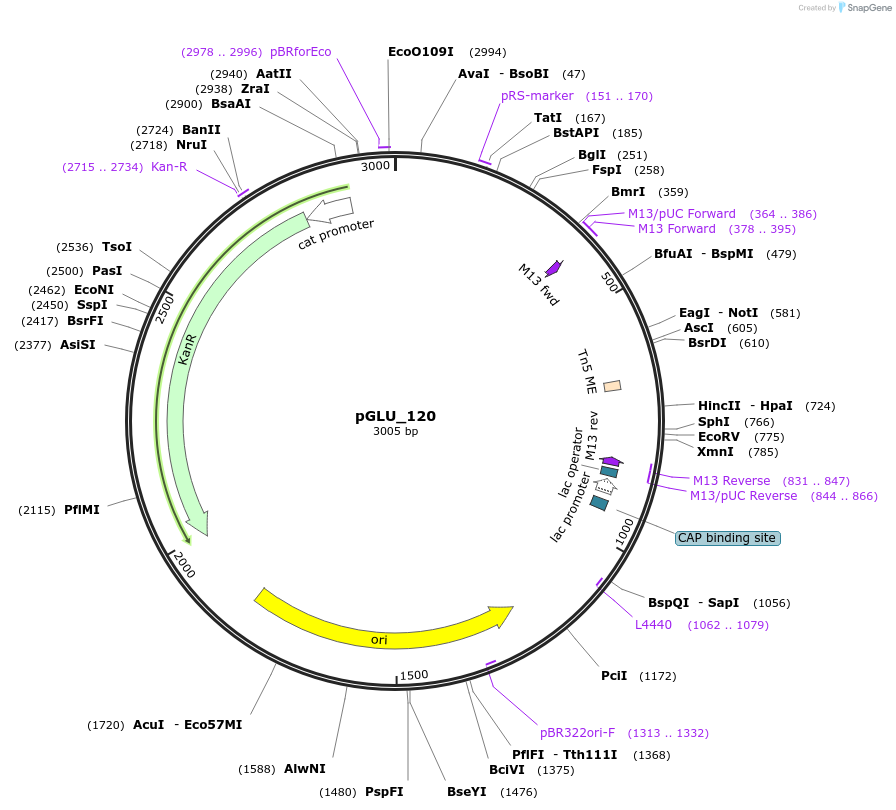
pGLU_120
Plasmid#249825PurposeRS421 library variant 13481 in IDT backboneDepositorInsert13481_[Tn5]
UseSynthetic BiologyMutationN/AAvailable SinceJan. 29, 2026AvailabilityAcademic Institutions and Nonprofits only -
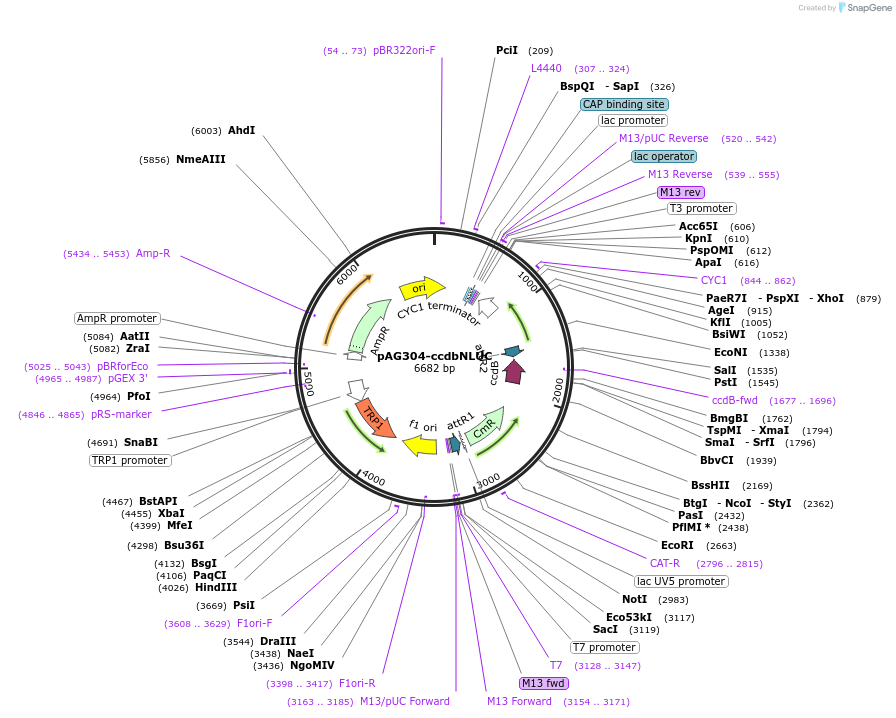
pAG304-ccdbNLUC
Plasmid#249736PurposeDerived from pAG304GPD-ccdb (#14136); GPD removed and NLUC cds inserted downstream gateway cassette. Useful for cloning promoter-NLUC fusions in Saccharomyces cerevisiaeDepositorTypeEmpty backboneExpressionYeastAvailable SinceJan. 28, 2026AvailabilityAcademic Institutions and Nonprofits only