We narrowed to 25,146 results for: NTS
-
Plasmid#52170PurposeEncodes module 7 with amino acids 12S-13Y-16R for recognition of nt 2CDepositorInsertPumilio homology domain amino acids 241-283 (PUM1 Human)
ExpressionBacterialMutationChanged 13N to 13Y and 16E to 16R in module 7Available SinceMay 5, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-1NQ
Plasmid#52141PurposeEncodes module 1 with amino acids 12N-16Q for recognition of nt 8UDepositorInsertPumilio homology domain amino acids 25-60 (PUM1 Human)
ExpressionBacterialMutationChanged 12S to 12N in Module 1Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-2CQ
Plasmid#52143PurposeEncodes module 2 with amino acids 12C-16Q for recognition of nt 7ADepositorInsertPumilio homology domain amino acids 61-96 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C in Module 2Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-2SR
Plasmid#52146PurposeEncodes module 2 with amino acids 12S-16R for recognition of nt 7CDepositorInsertPumilio homology domain amino acids 61-96 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S and 16Q to 16R in Module 2Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-3NQ
Plasmid#52149PurposeEncodes module 3 with amino acids 12N-16Q for recognition of nt 6UDepositorInsertPumilio homology domain amino acids 97-132 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12N in Module 3Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-3SR
Plasmid#52150PurposeEncodes module 3 with amino acids 12S-16R for recognition of nt 6CDepositorInsertPumilio homology domain amino acids 97-132 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12S and 16Q to 16R in Module 3Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4CQ
Plasmid#52151PurposeEncodes module 4 with amino acids 12C-16Q for recognition of nt 5ADepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4SR
Plasmid#52154PurposeEncodes module 4 with amino acids 12S-16R for recognition of nt 5CDepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S and 16Q to 16R in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4CYQ
Plasmid#52155PurposeEncodes module 4 with amino acids 12C-13Y-16Q for recognition of nt 5ADepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C and 13H to 13Y in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4SYE
Plasmid#52156PurposeEncodes module 4 with amino acids 12S-13Y-16E for recognition of nt 5GDepositorInsertD43951 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S, 13H to 13Y, and 16Q to 16E in…Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4NYQ
Plasmid#52157PurposeEncodes module 4 with amino acids 12N-13Y-16Q for recognition of nt 5UDepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 13H to 13Y in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-5SE
Plasmid#52160PurposeEncodes module 5 with amino acids 12S-16E for recognition of nt 4GDepositorInsertPumilio homology domain amino acids 169-204 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12S and 16Q to 16E in Module 5Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-5NQ
Plasmid#52161PurposeEncodes module 5 with amino acids 12N-16Q for recognition of nt 4UDepositorInsertPumilio homology domain amino acids 169-204 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12N in Module 5Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-6CQ
Plasmid#52163PurposeEncodes module 6 with amino acids 12C-16Q for recognition of nt 3ADepositorInsertPumilio homology domain amino acids 205-240 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C in Module 6Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-6SR
Plasmid#52166PurposeEncodes module 6 with amino acids 12S-16R for recognition of nt 3CDepositorInsertPumilio homology domain amino acids 205-240 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S and 16Q to 16R in Module 6Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
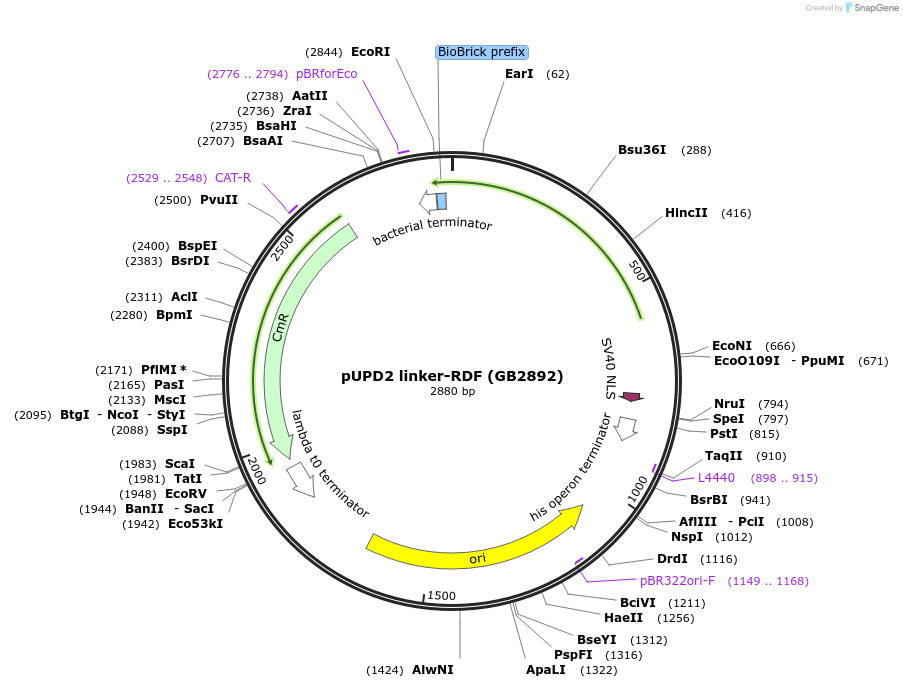
pUPD2 linker-RDF (GB2892)
Plasmid#160604PurposeRDF sequence for C-terminal fusion with PhiC31 recombinase, together with a 18-nt linker sequenceDepositorInsertRDF
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceJan. 14, 2021AvailabilityAcademic Institutions and Nonprofits only -
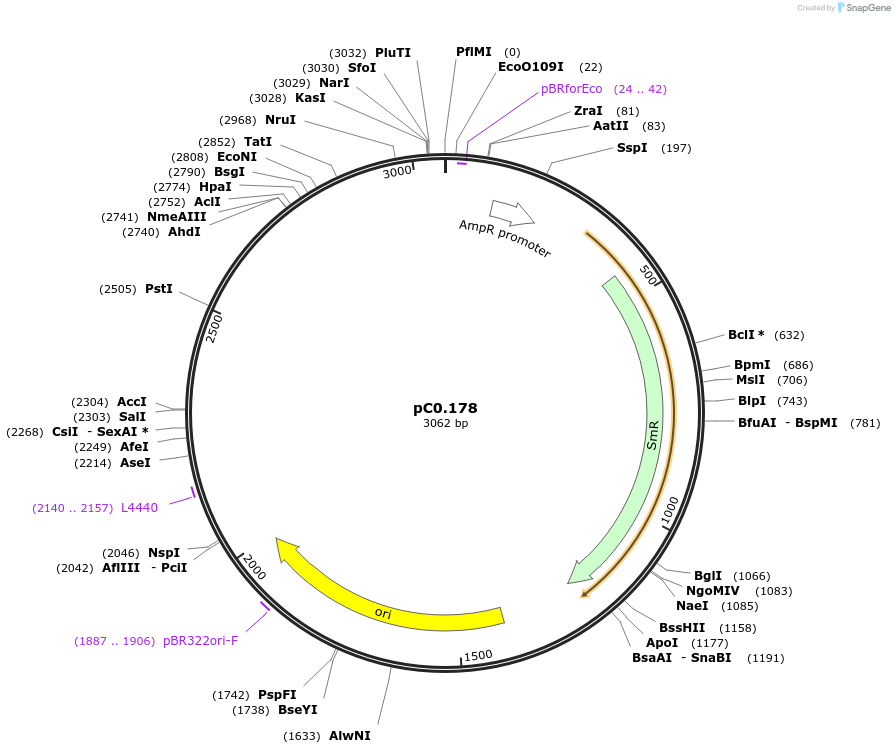
pC0.178
Plasmid#119645PurposeLevel 0 Part. Insertion siteDepositorInsert7942 NS3 Down Flank
UseSynthetic BiologyMutation4 nucleotide insertion between nt 65 and 66 in NS…Available SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
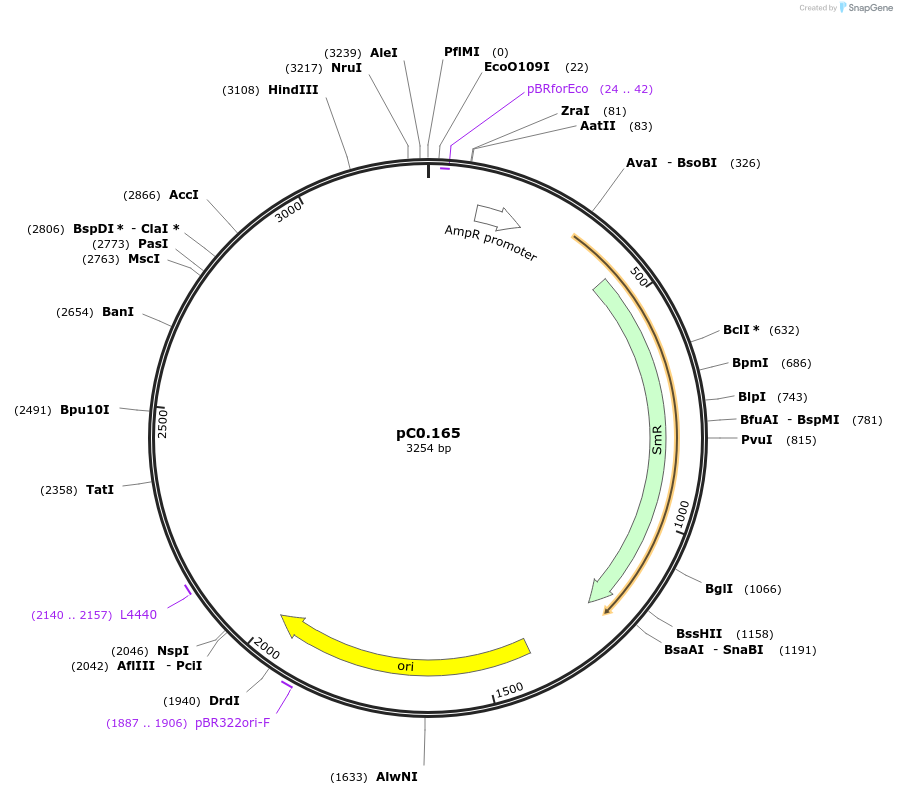
pC0.165
Plasmid#119634PurposeLevel 0 Part. Insertion siteDepositorInsert6803 NS1 Up Flank
UseSynthetic BiologyMutation1 nucleotide insertion between nt 21 and 22 in NS…Available SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
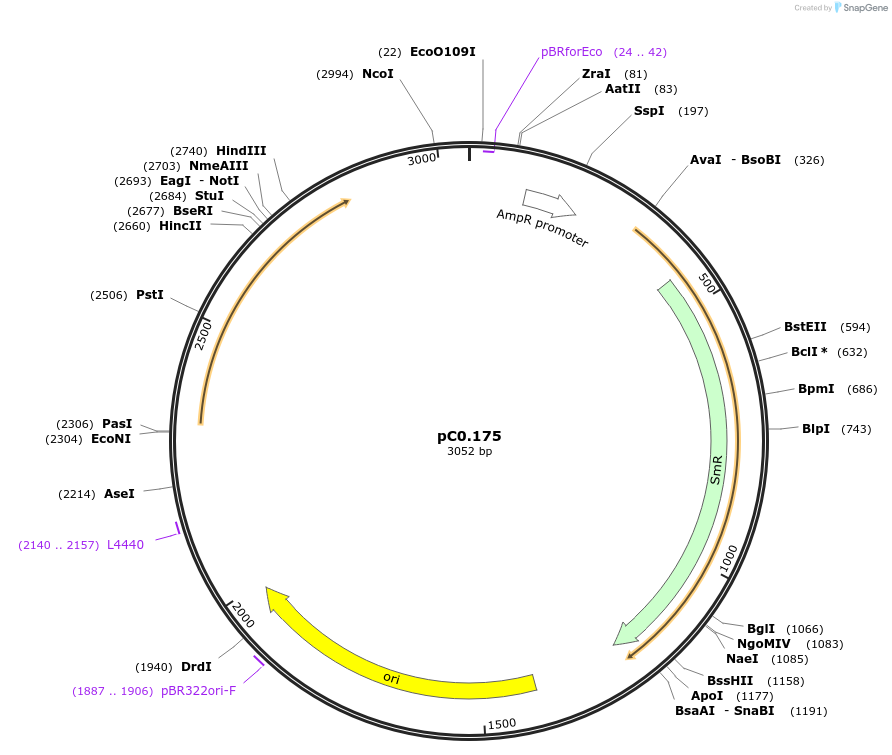
pC0.175
Plasmid#119643PurposeLevel 0 Part. Insertion siteDepositorInsert7942 NS2 Up Flank
UseSynthetic BiologyMutationbp 422 A to T, bp 423 A to T, bp 663 inserted G. …Available SinceAug. 6, 2019AvailabilityAcademic Institutions and Nonprofits only -
pVR2
Plasmid#84719PurposeE. coli RecA promoter followed by stretch with no A's followed by riboswitch for single-round transcriptionDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutation12 nt stretch lacking A's inserted before 5&…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only