We narrowed to 8,780 results for: construct
-
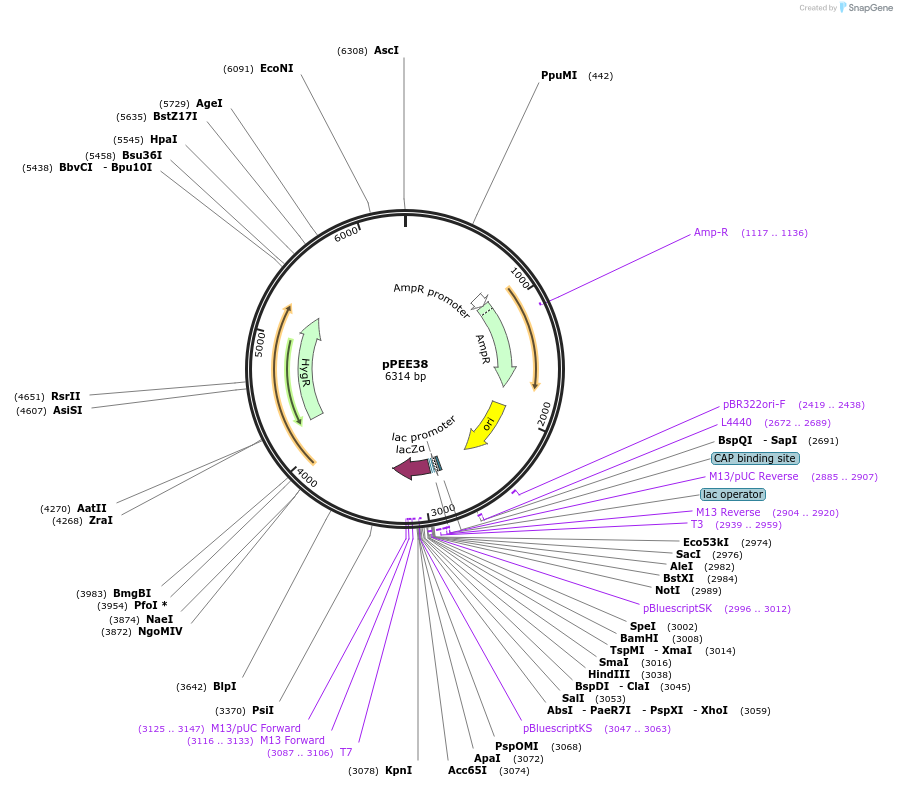
Plasmid#128378PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceJan. 8, 2020AvailabilityAcademic Institutions and Nonprofits only
-
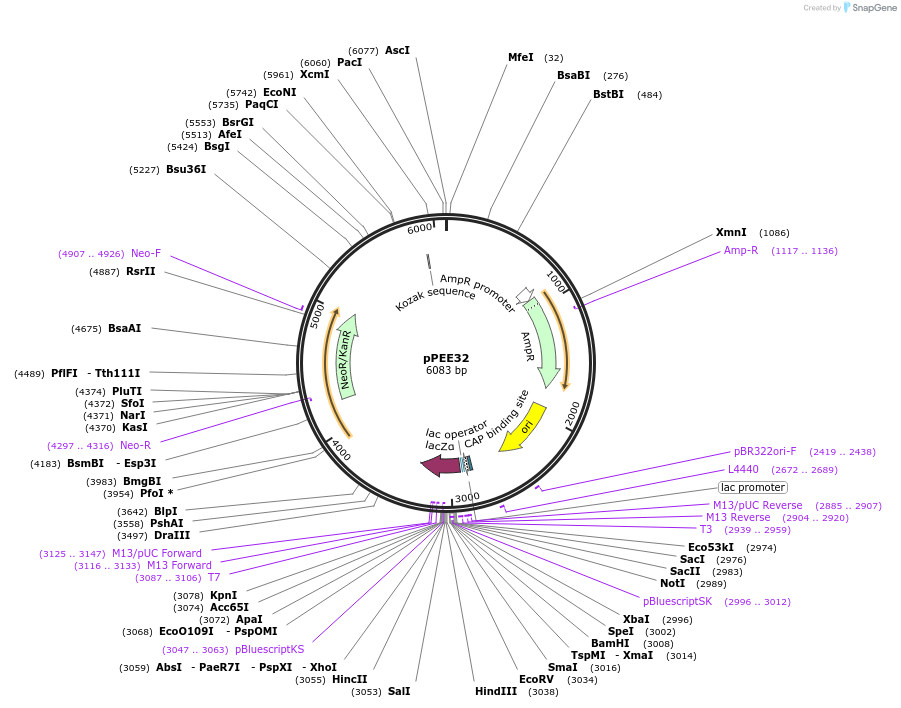
pPEE32
Plasmid#128372PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceJan. 8, 2020AvailabilityAcademic Institutions and Nonprofits only -
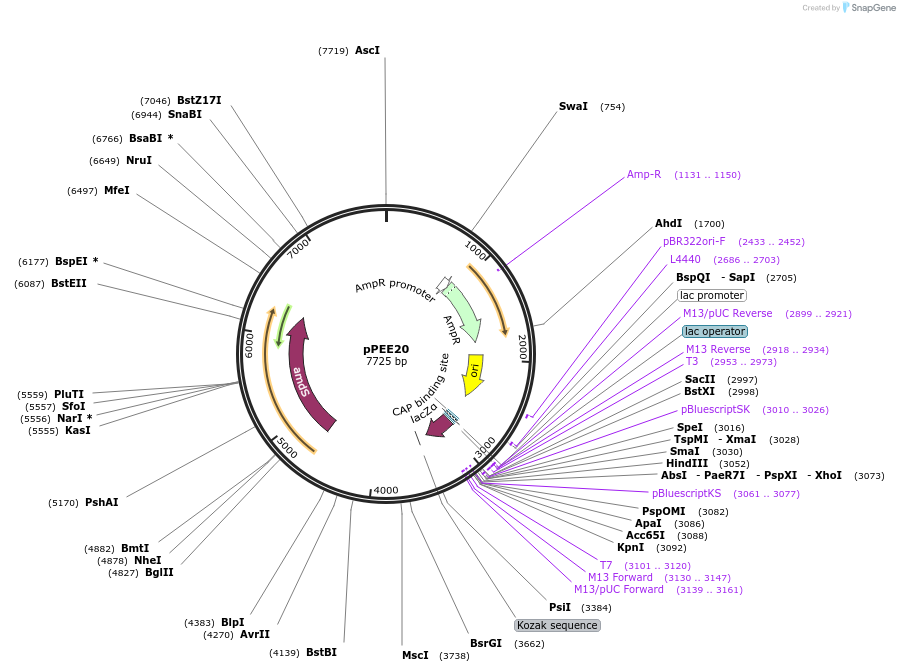
pPEE20
Plasmid#128363PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceOct. 11, 2019AvailabilityAcademic Institutions and Nonprofits only -
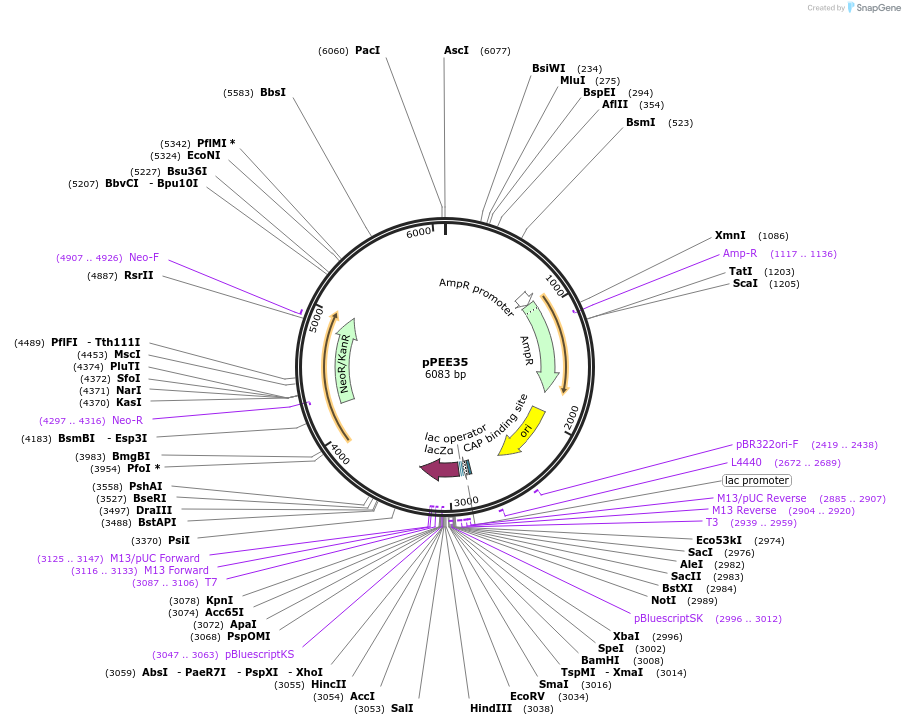
pPEE35
Plasmid#128375PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
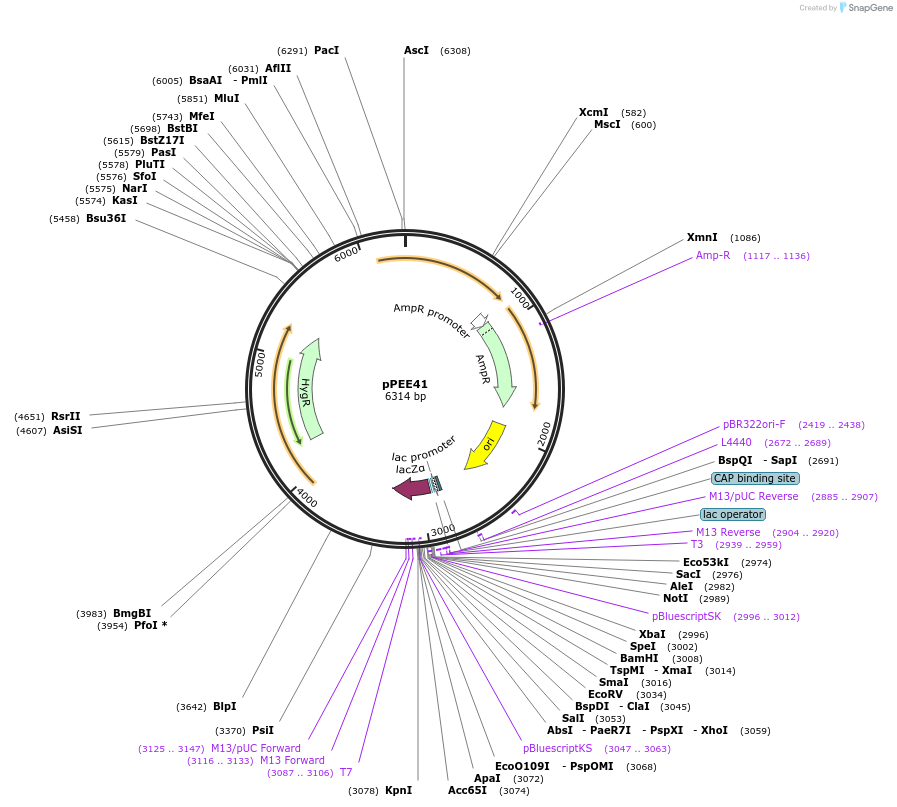
pPEE41
Plasmid#128381PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
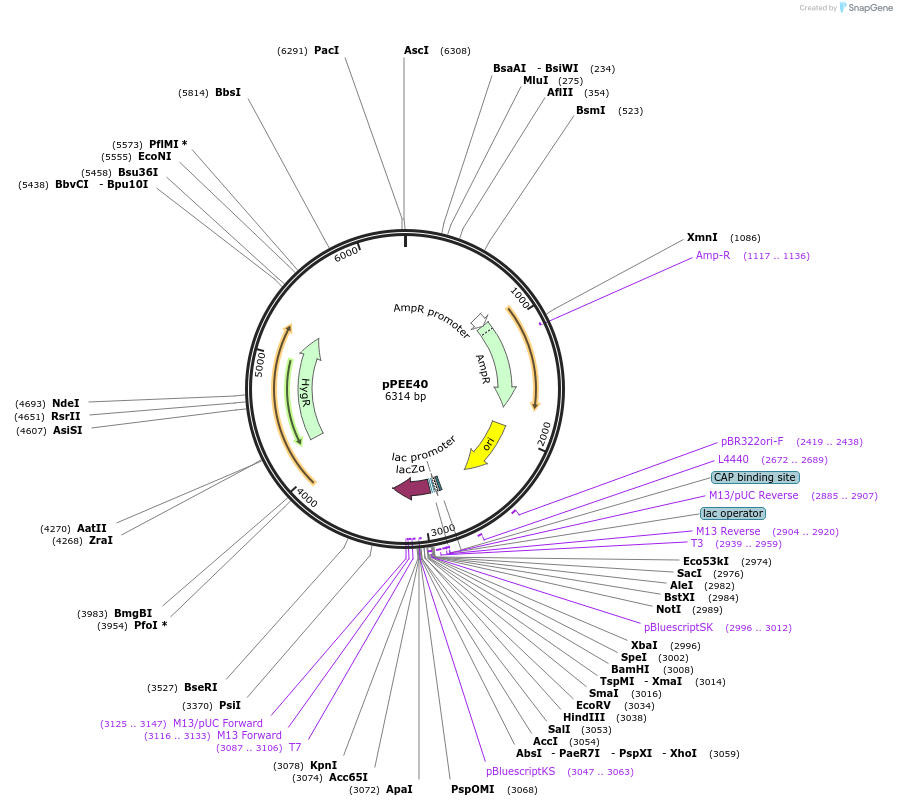
pPEE40
Plasmid#128380PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
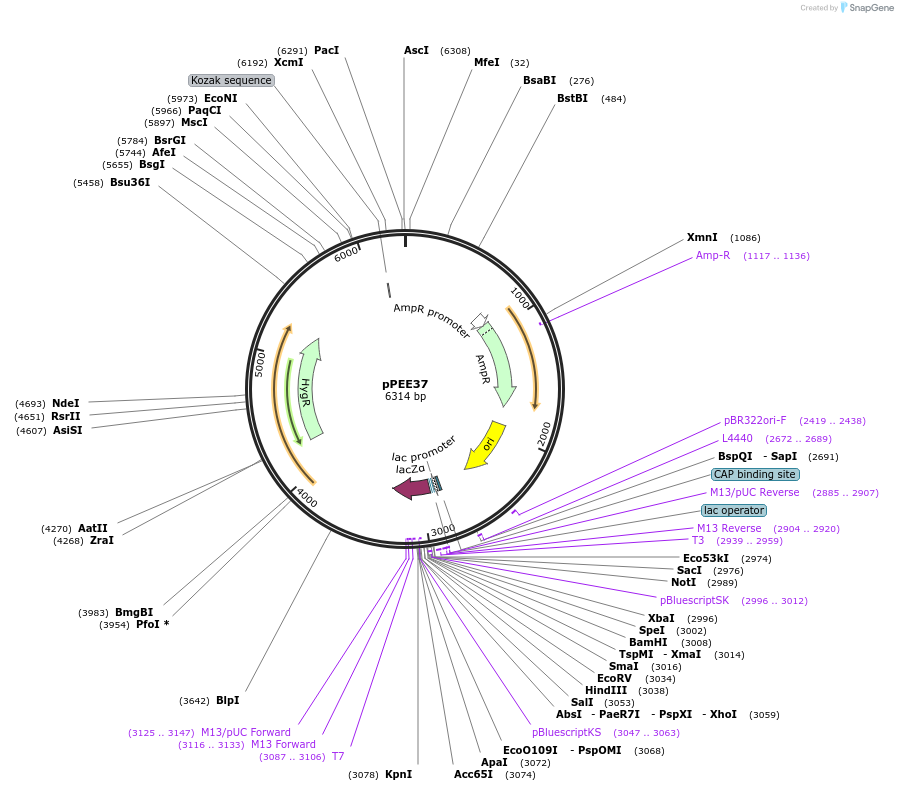
pPEE37
Plasmid#128377PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
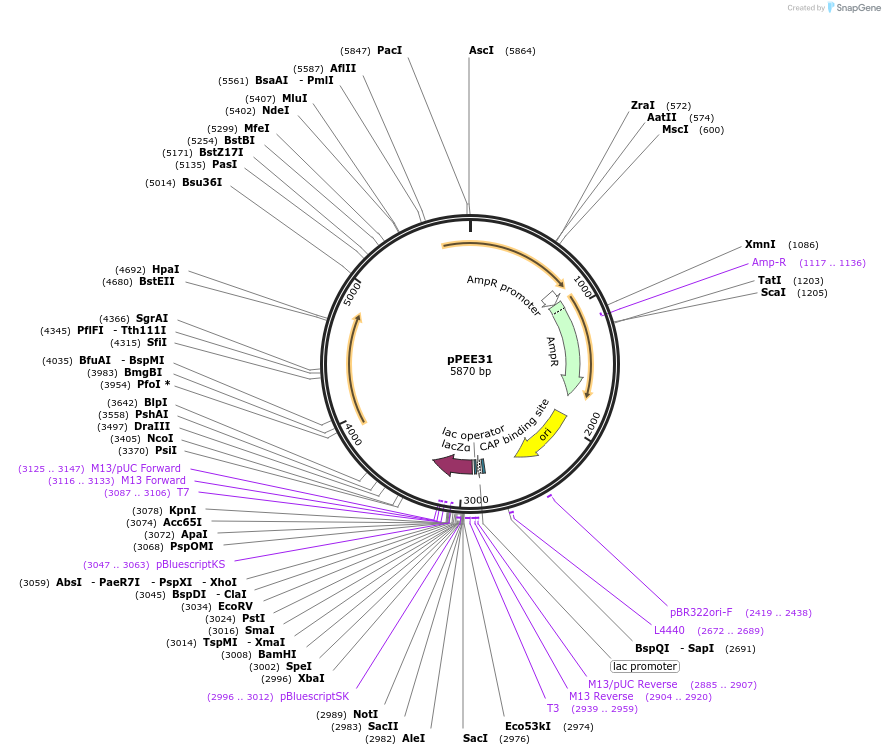
pPEE31
Plasmid#128371PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
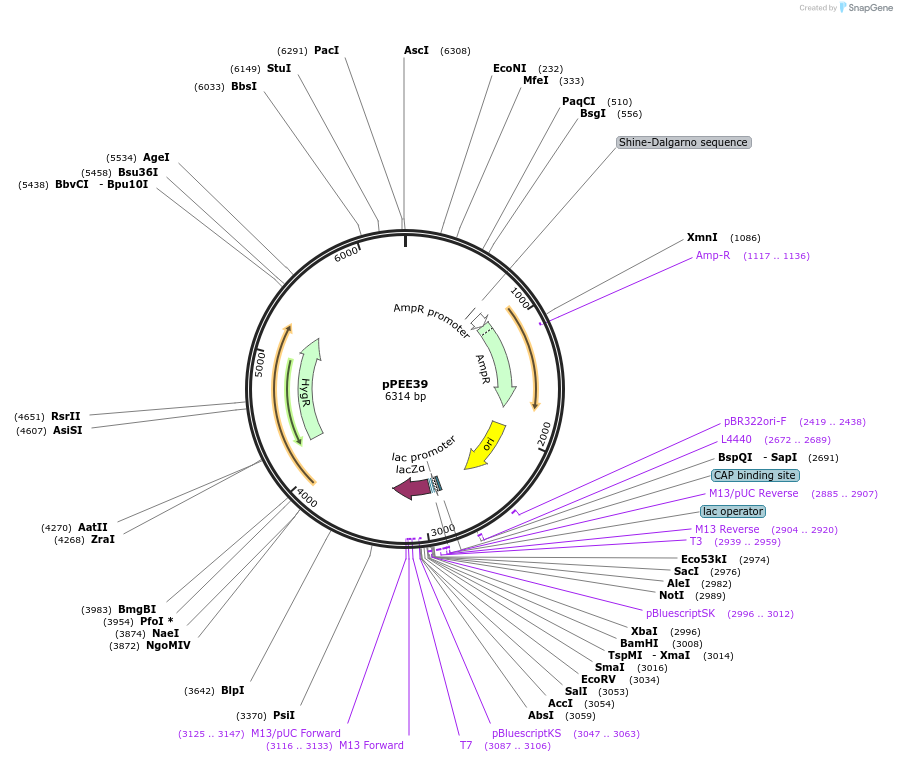
pPEE39
Plasmid#128379PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
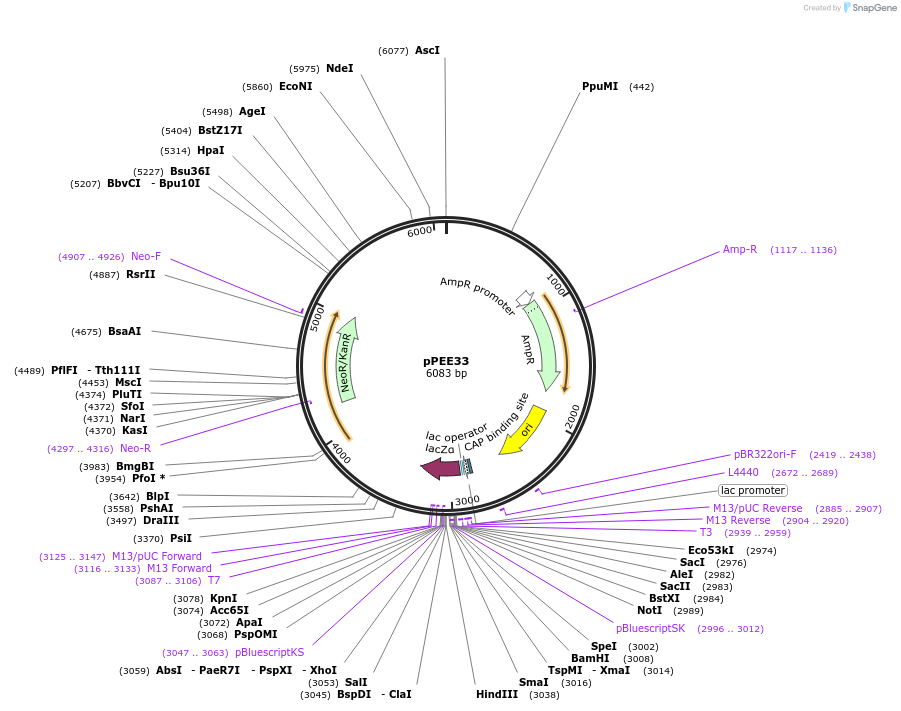
pPEE33
Plasmid#128373PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
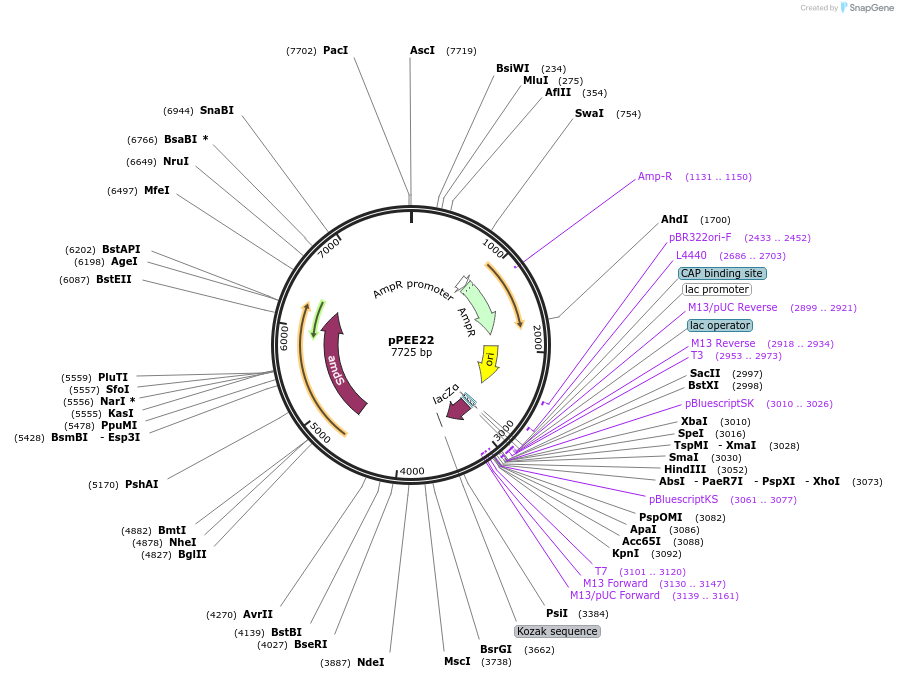
pPEE22
Plasmid#128365PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
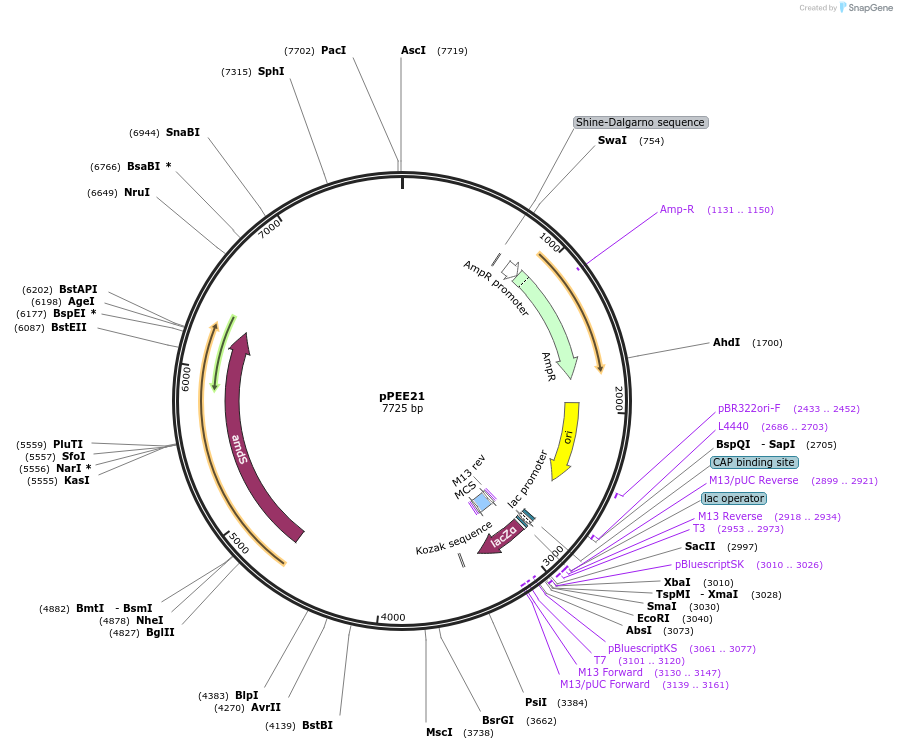
pPEE21
Plasmid#128364PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
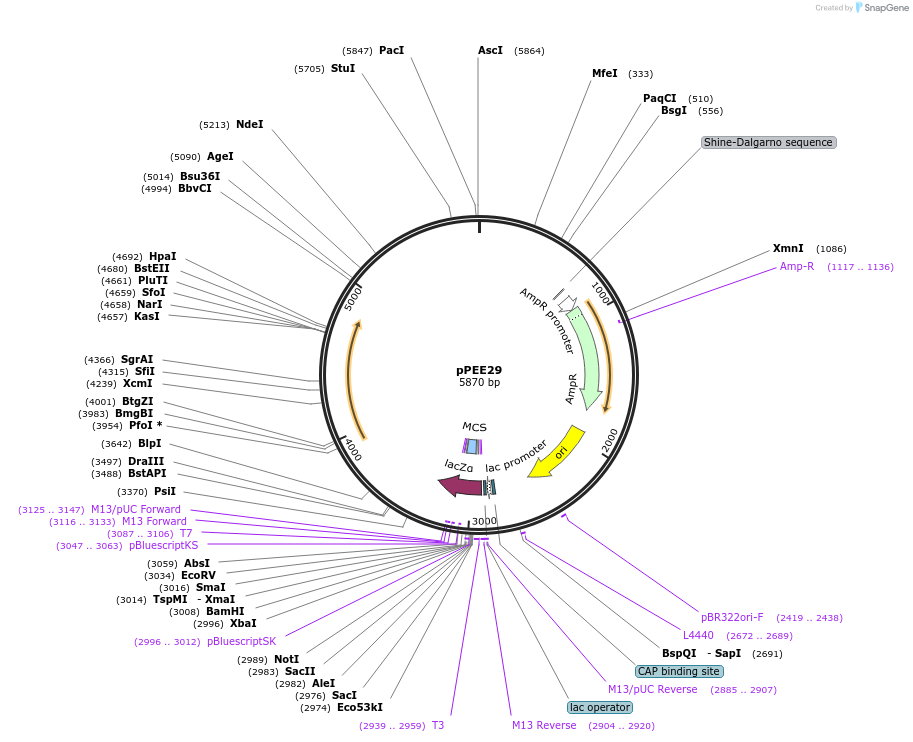
pPEE29
Plasmid#128369PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
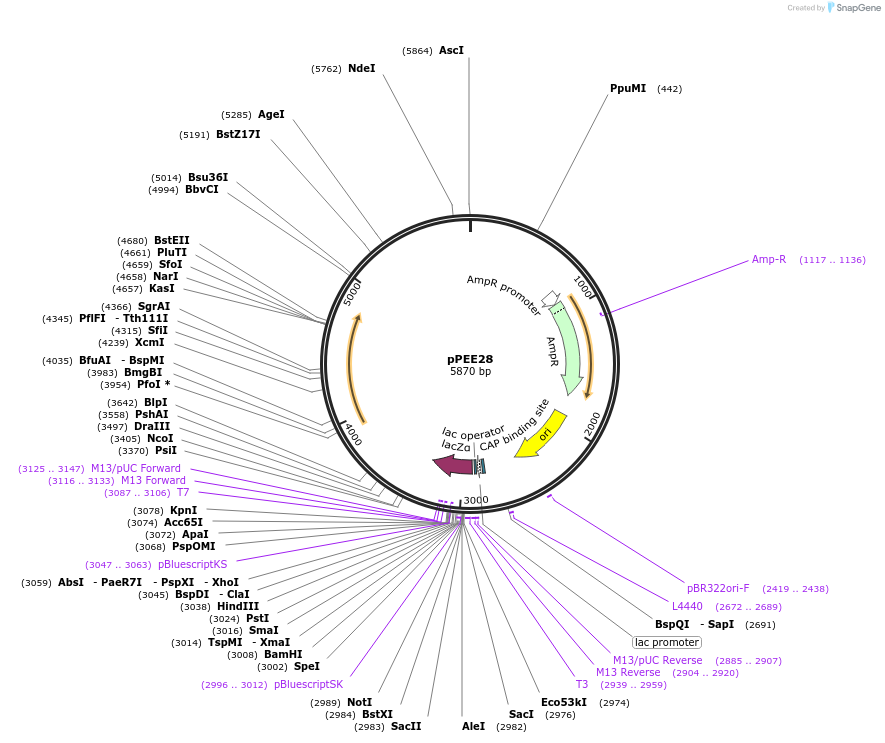
pPEE28
Plasmid#128368PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
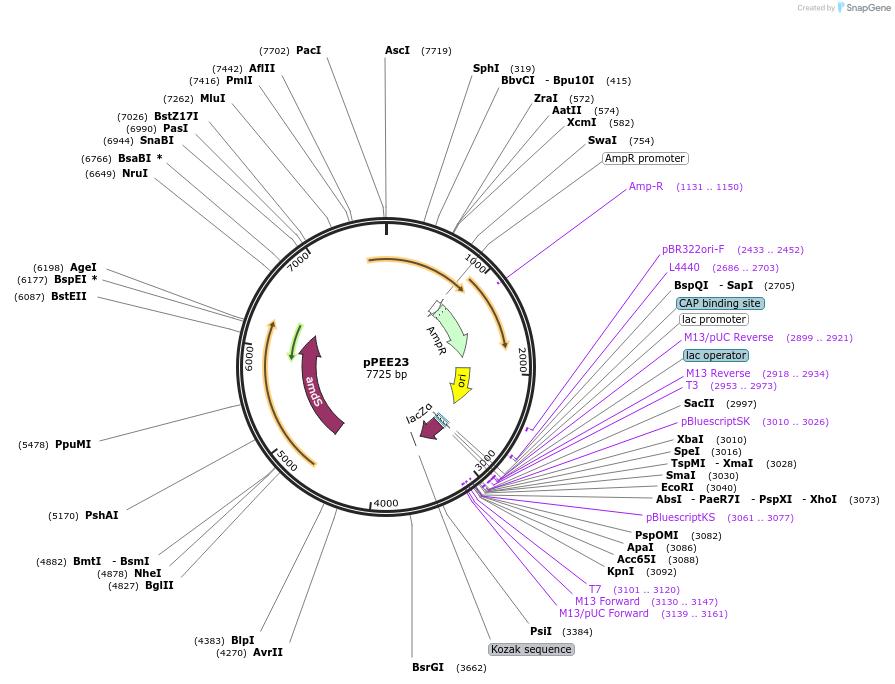
pPEE23
Plasmid#128366PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
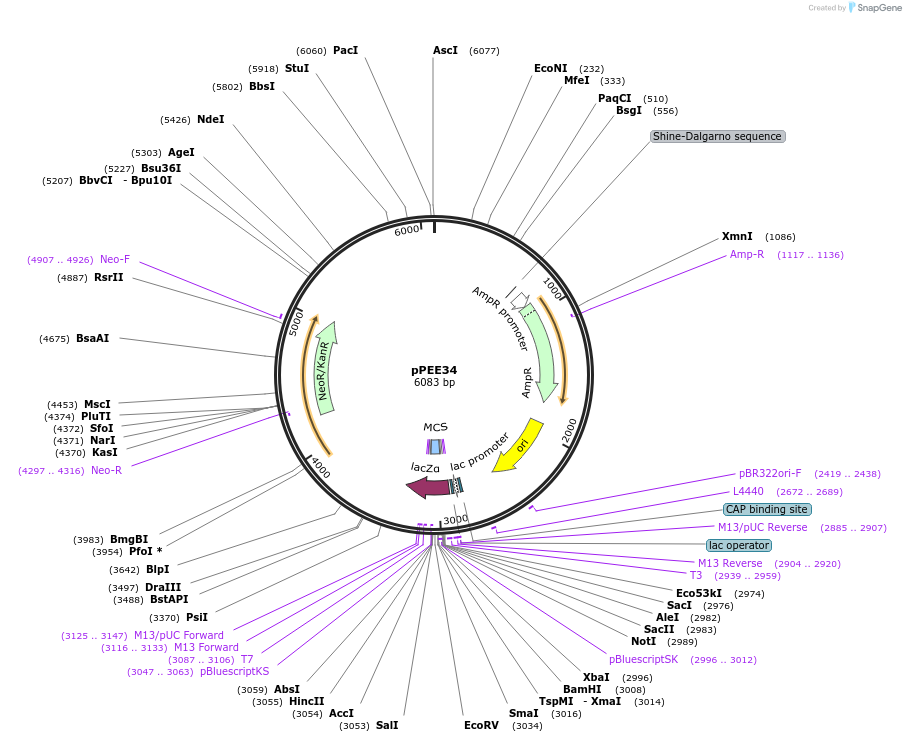
pPEE34
Plasmid#128374PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
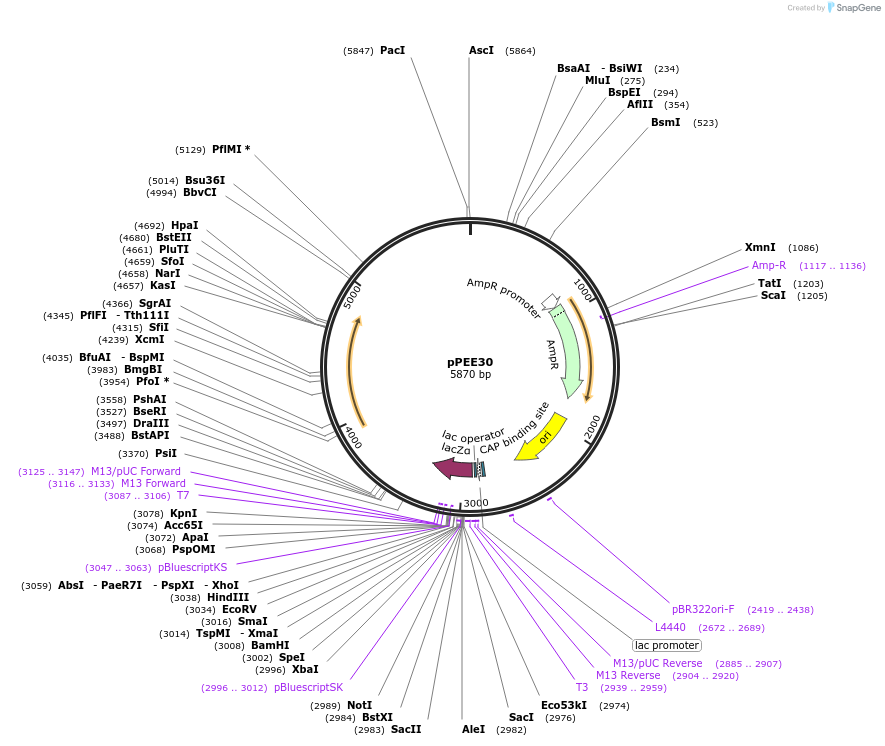
pPEE30
Plasmid#128370PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
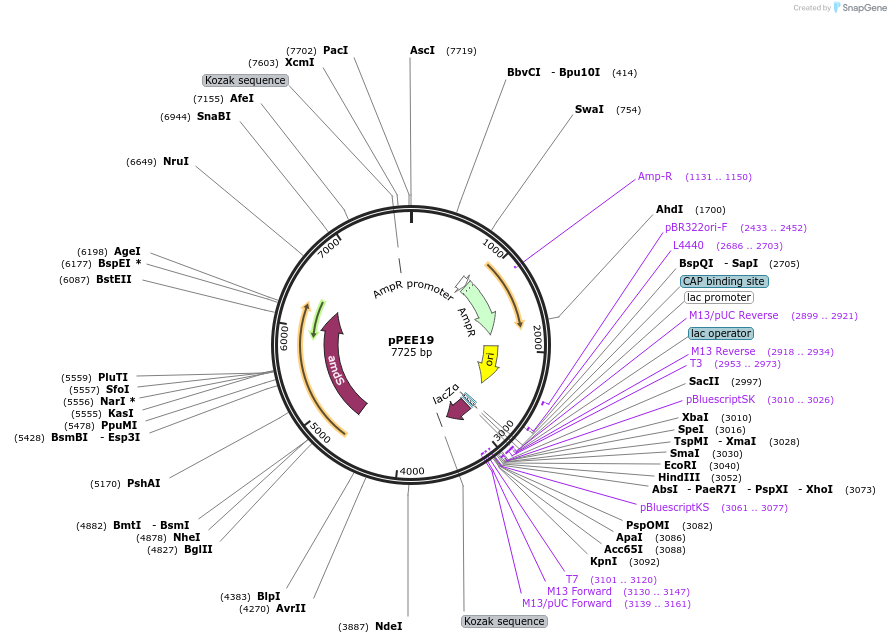
pPEE19
Plasmid#128362PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
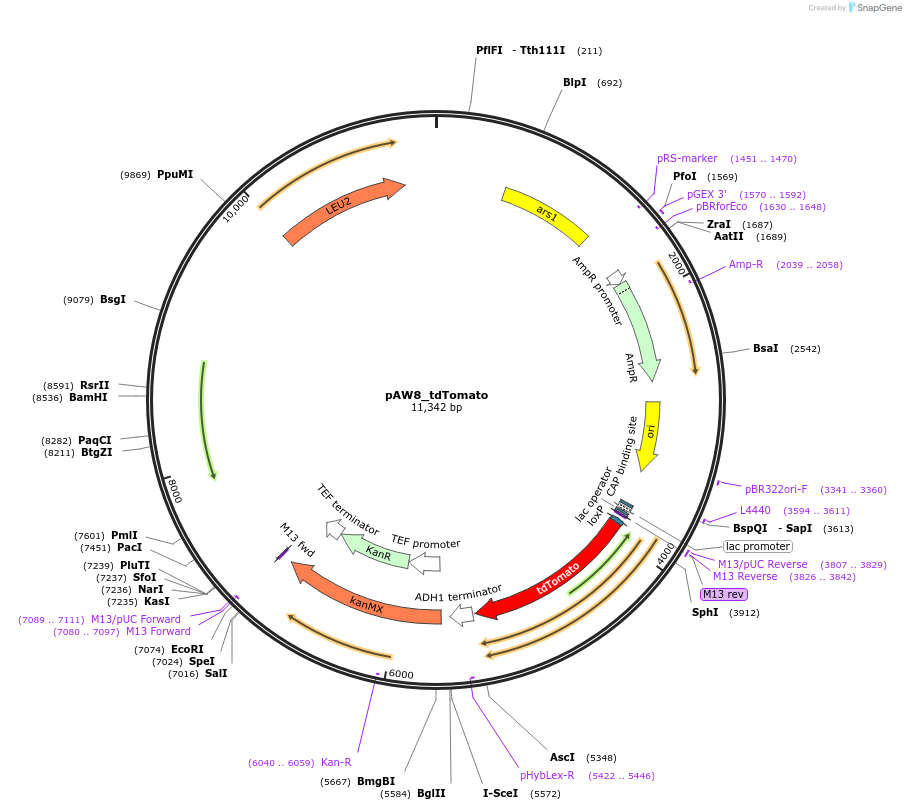
pAW8_tdTomato
Plasmid#110239PurposeCre/Lox C-terminal tagging construct encoding red fluorescent protein tdTomatoDepositorTypeEmpty backboneUseCre/LoxAvailable SinceMay 21, 2019AvailabilityAcademic Institutions and Nonprofits only -
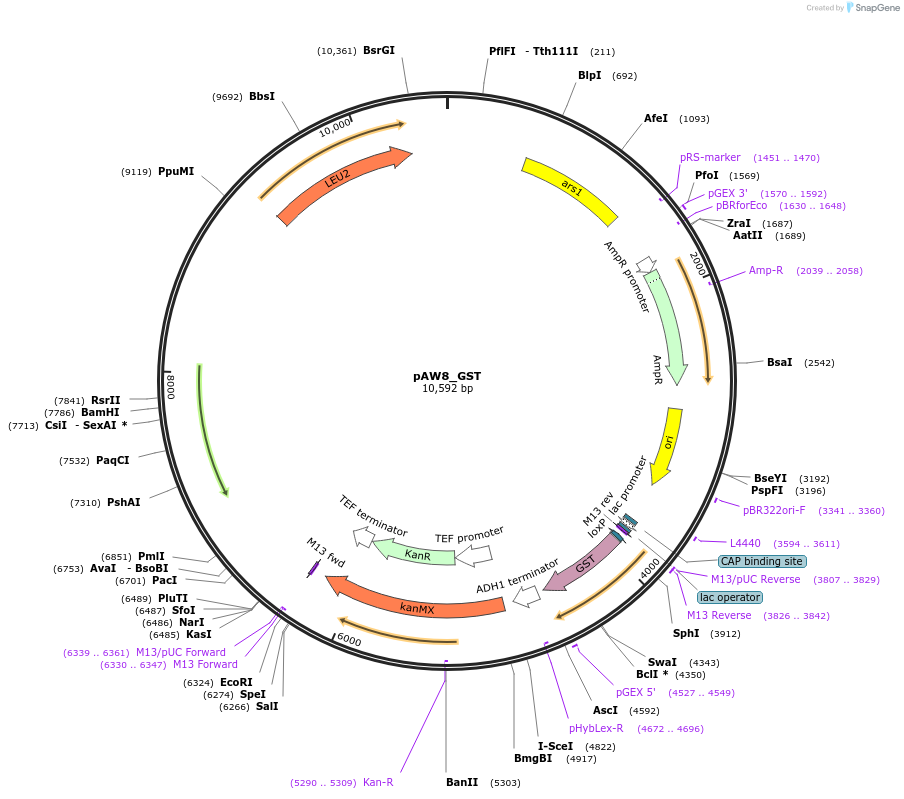
pAW8_GST
Plasmid#110237PurposeCre/Lox C-terminal tagging construct encoding glutathione S-transferaseDepositorTypeEmpty backboneUseCre/LoxAvailable SinceMay 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
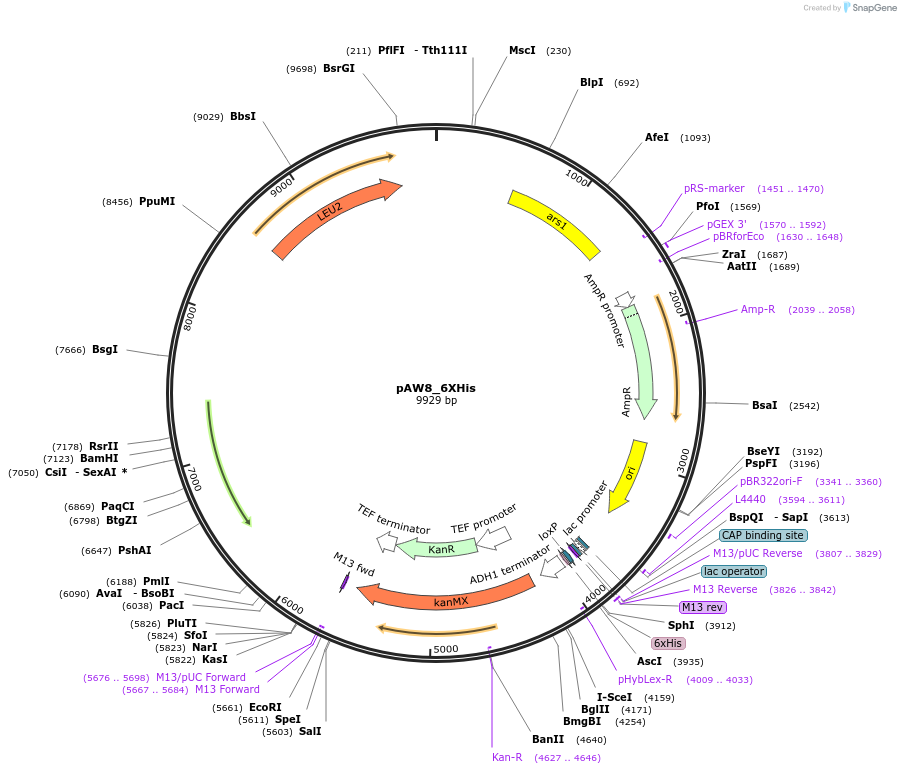
pAW8_6XHis
Plasmid#110236PurposeCre/Lox C-terminal tagging construct encoding six histidine (His) residuesDepositorTypeEmpty backboneUseCre/LoxAvailable SinceApril 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
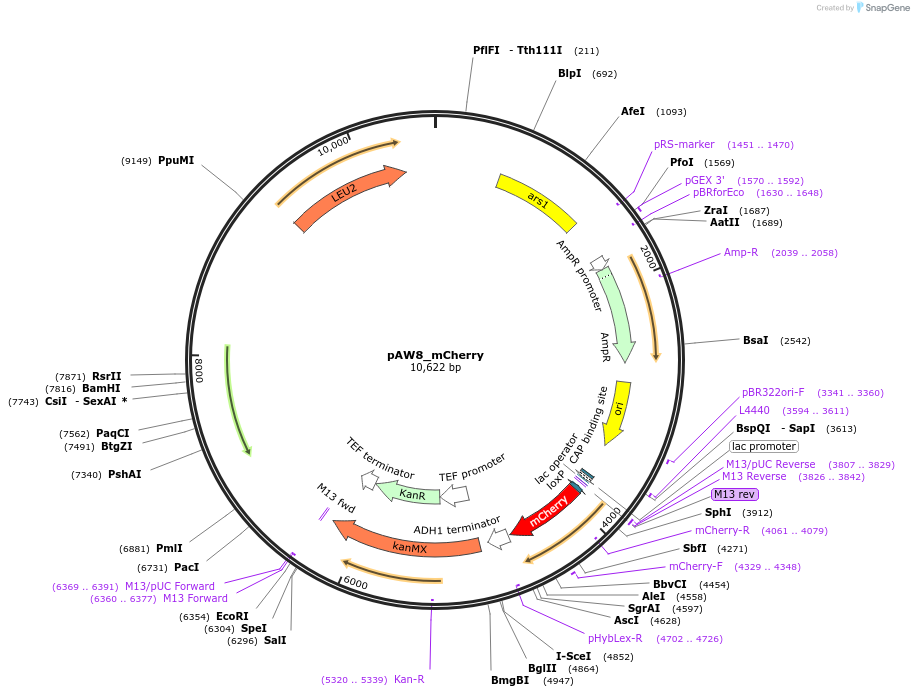
pAW8_mCherry
Plasmid#110238PurposeCre/Lox C-terminal tagging construct encoding red fluorescent protein mCherryDepositorTypeEmpty backboneUseCre/LoxAvailable SinceApril 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
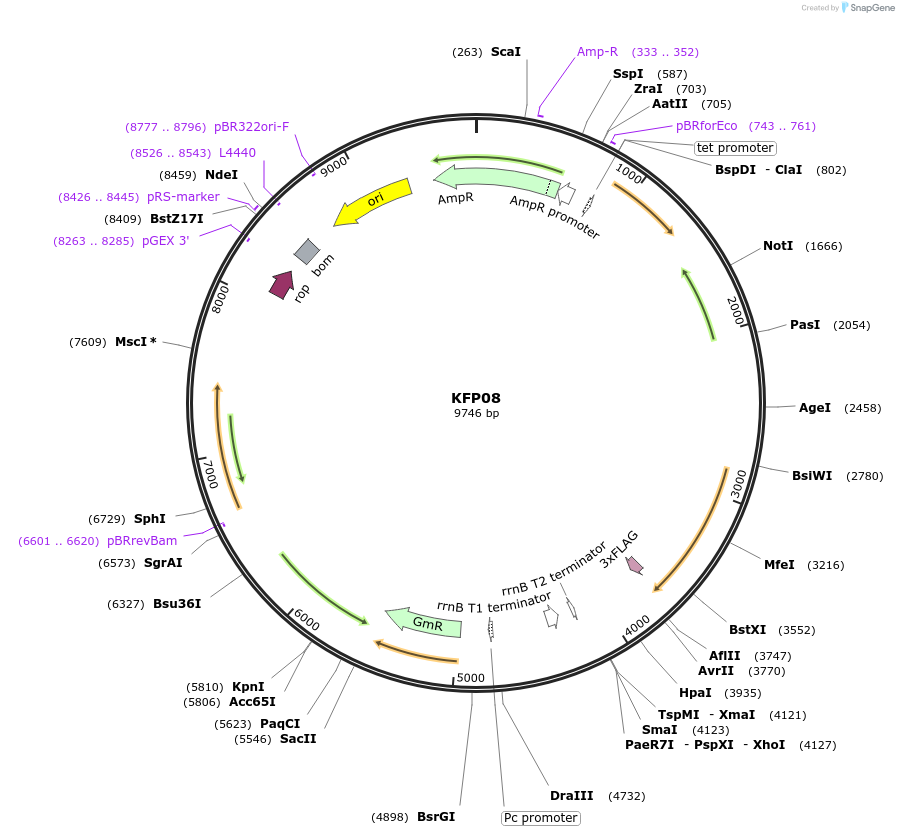
KFP08
Plasmid#119798PurposeSigF2 epitope tagged construct for integration at neutral site 2.2DepositorInsertSynpcc7942_1784
ExpressionBacterialAvailable SinceDec. 21, 2018AvailabilityAcademic Institutions and Nonprofits only -
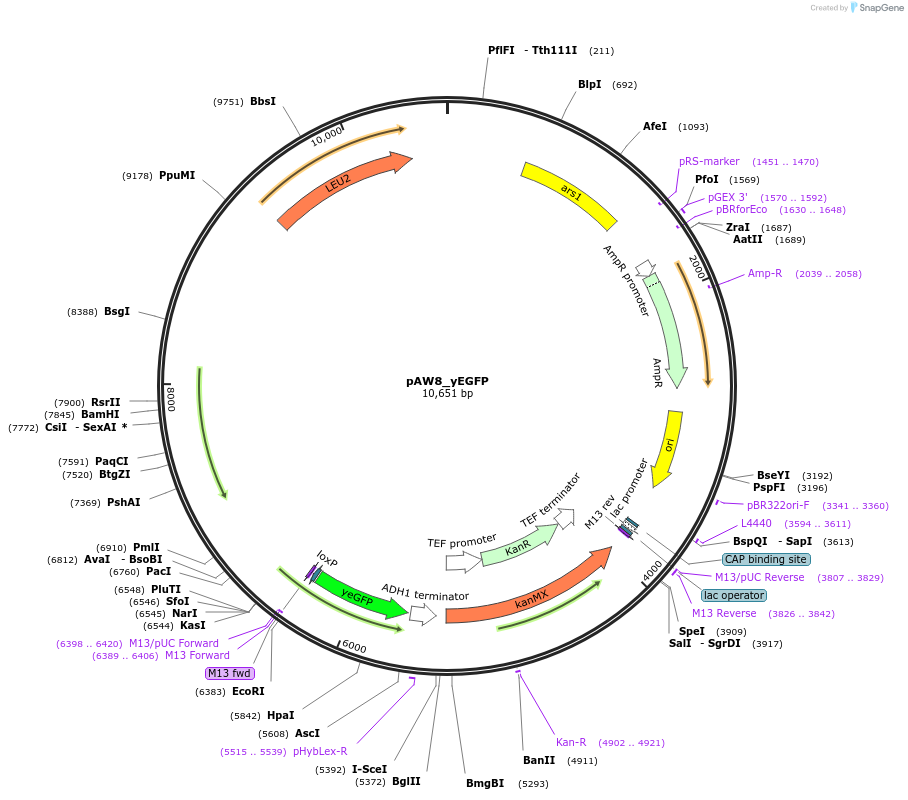
pAW8_yEGFP
Plasmid#110229PurposeCre/Lox C-terminal tagging construct encoding codon-optimized green fluorescent proteinDepositorTypeEmpty backboneUseCre/LoxAvailable SinceOct. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
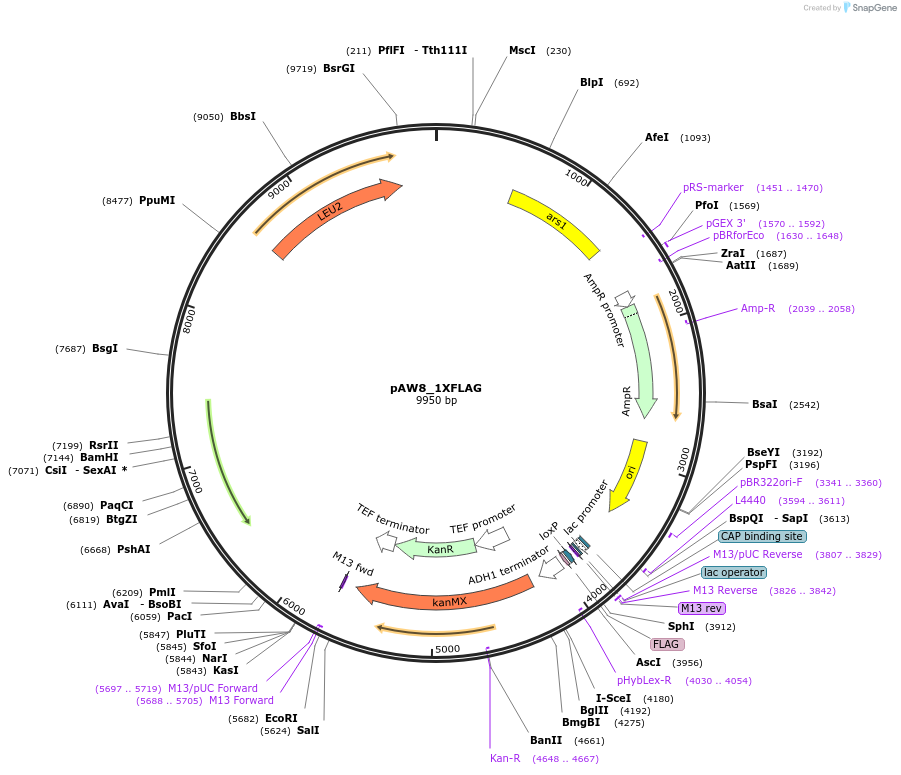
pAW8_1XFLAG
Plasmid#110234PurposeCre/Lox C-terminal tagging construct encoding single copy of the FLAG epitopeDepositorTypeEmpty backboneUseCre/LoxAvailable SinceSept. 18, 2018AvailabilityAcademic Institutions and Nonprofits only -
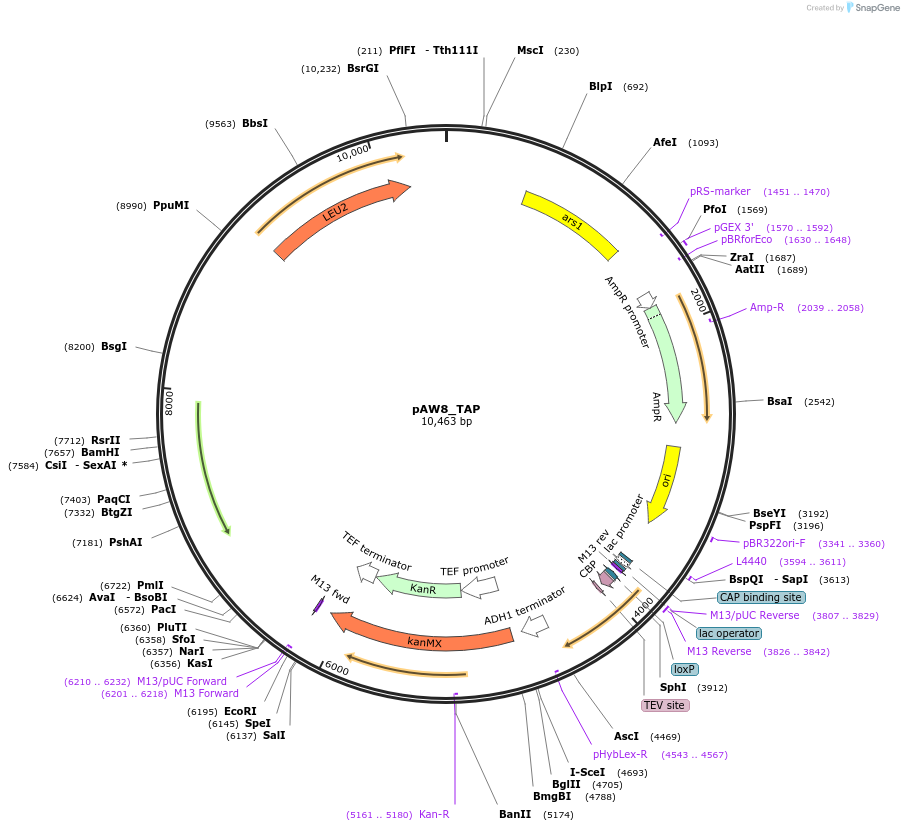
pAW8_TAP
Plasmid#110240PurposeCre/Lox C-terminal tagging construct encoding the Twin Affinity Purification proteinDepositorTypeEmpty backboneUseCre/LoxAvailable SinceAug. 22, 2018AvailabilityAcademic Institutions and Nonprofits only -
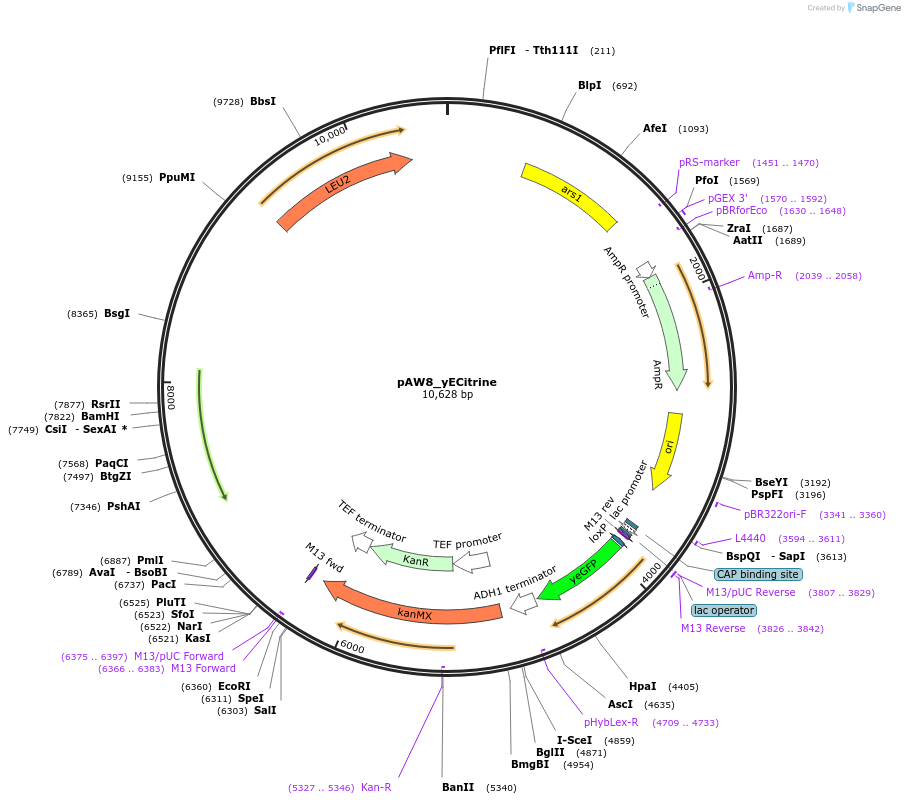
pAW8_yECitrine
Plasmid#110231PurposeCre/Lox C-terminal tagging construct encoding codon-optimized yellow fluorescent proteinDepositorTypeEmpty backboneUseCre/LoxAvailable SinceJuly 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
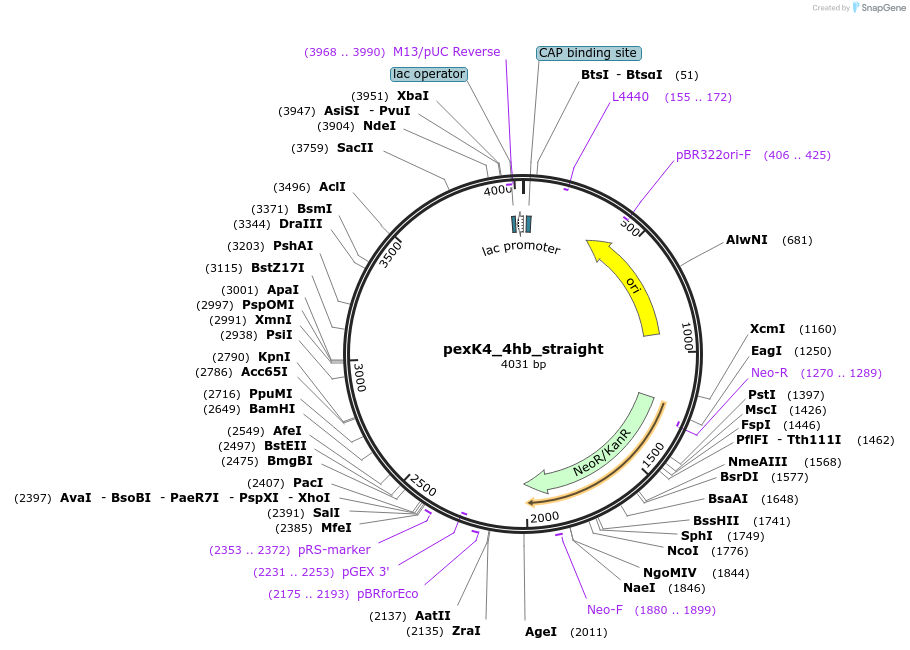
pexK4_4hb_straight
Plasmid#89678PurposePlasmid for the construction of DNA-protein hybrid nanostructures. Template for the straight four-helix-bundle shown in figure 4A of associated article.DepositorInsert4hb_straight
UseUnspecifiedAvailable SinceAug. 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
pexK4_4hb_bent
Plasmid#89679PurposePlasmid for the construction of DNA-protein hybrid nanostructures. Template for the bent four-helix-bundle shown in figure 4E of associated article.DepositorInsert4hb_bent
UseUnspecifiedAvailable SinceAug. 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
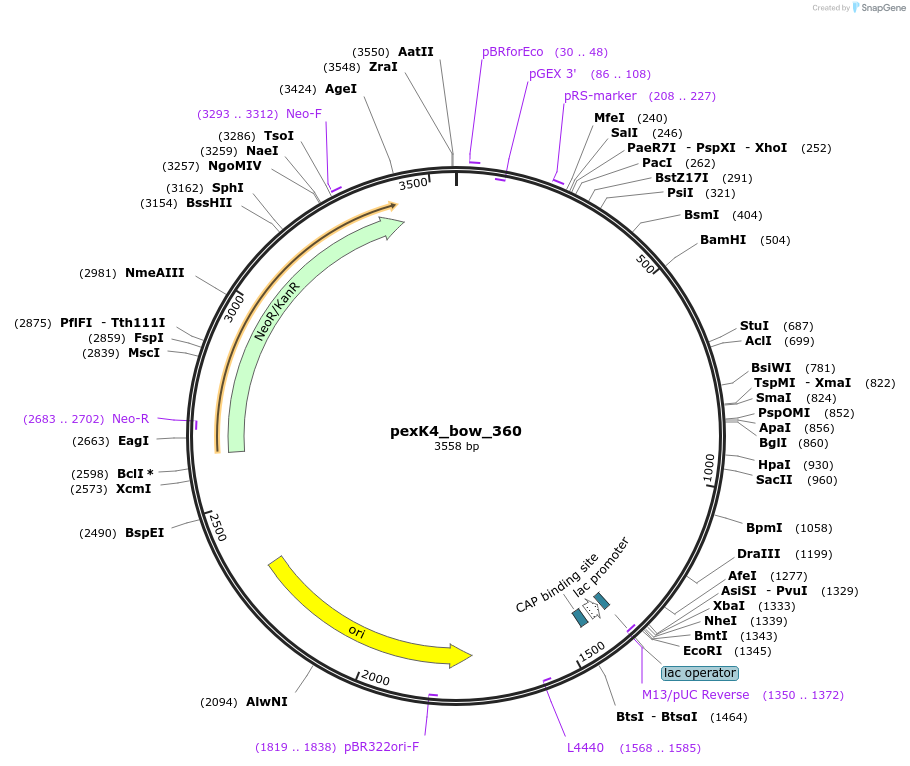
pexK4_bow_360
Plasmid#89673PurposePlasmid for the construction of DNA-protein hybrid nanostructures. Template for the bent two-helix-bundle shown in figure 3B of associated article.DepositorInsertbow_360
UseUnspecifiedAvailable SinceAug. 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
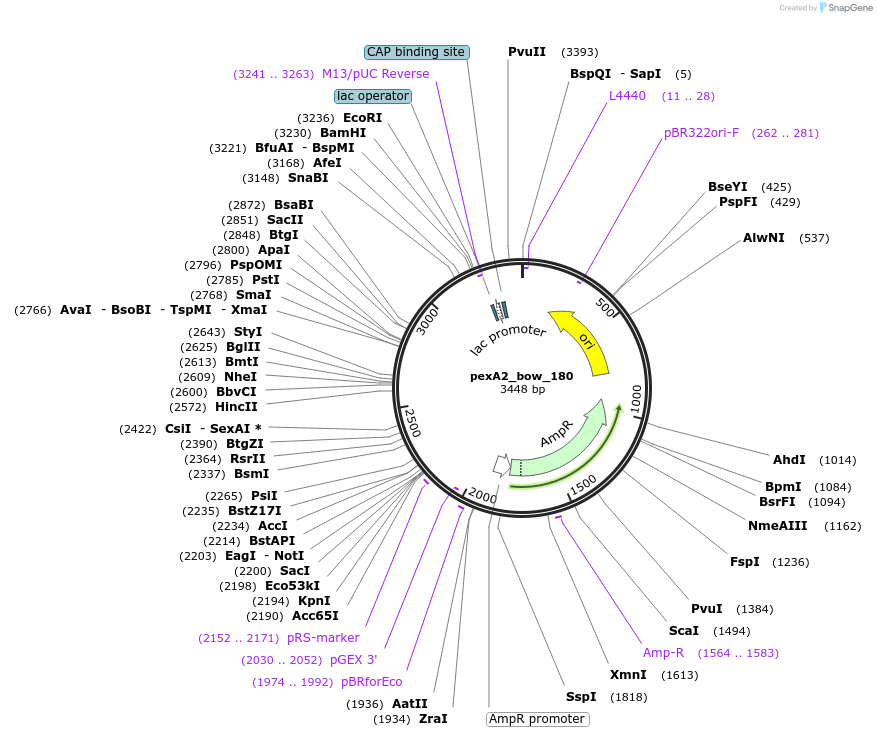
pexA2_bow_180
Plasmid#89671PurposePlasmid for the construction of DNA-protein hybrid nanostructures. Template for the bent two-helix-bundle shown in figure 3A of associated article.DepositorInsertbow_180
UseUnspecifiedAvailable SinceAug. 23, 2017AvailabilityAcademic Institutions and Nonprofits only -
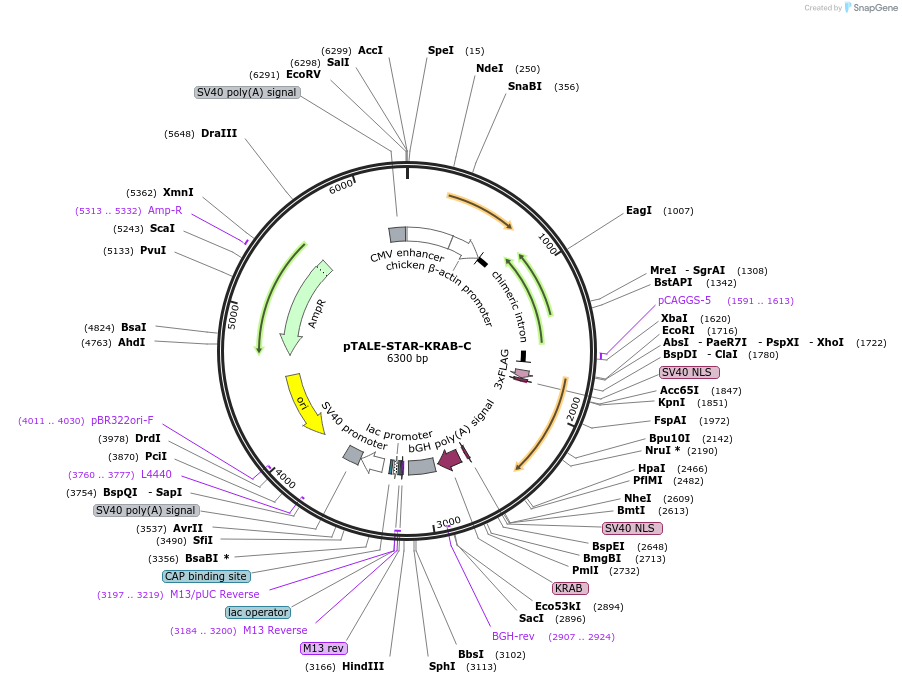
pTALE-STAR-KRAB-C
Plasmid#87540PurposeDestination vector for TALE-KRAB constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (repressor)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
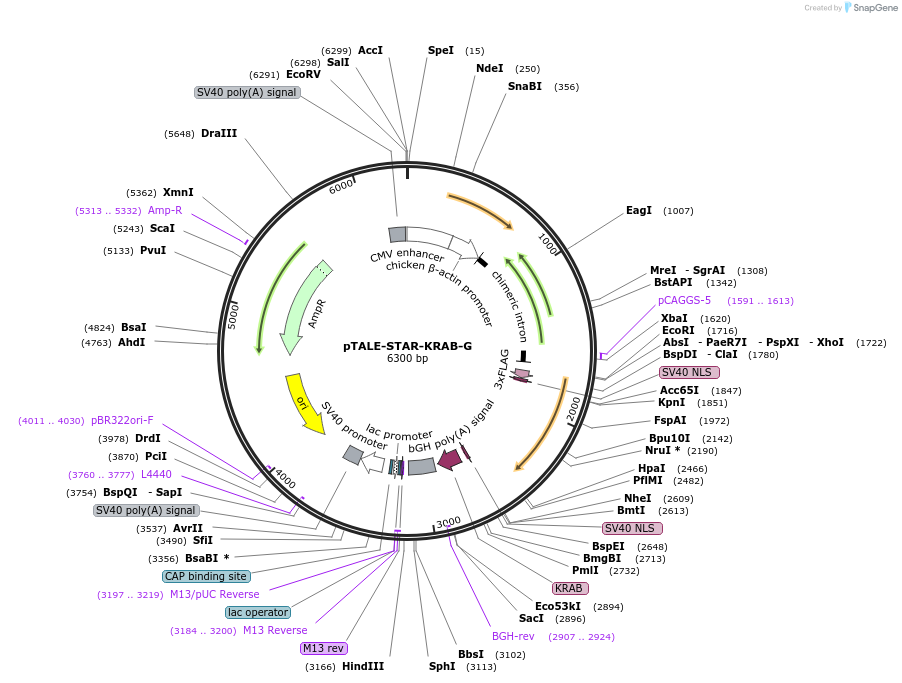
pTALE-STAR-KRAB-G
Plasmid#87541PurposeDestination vector for TALE-KRAB constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (repressor)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
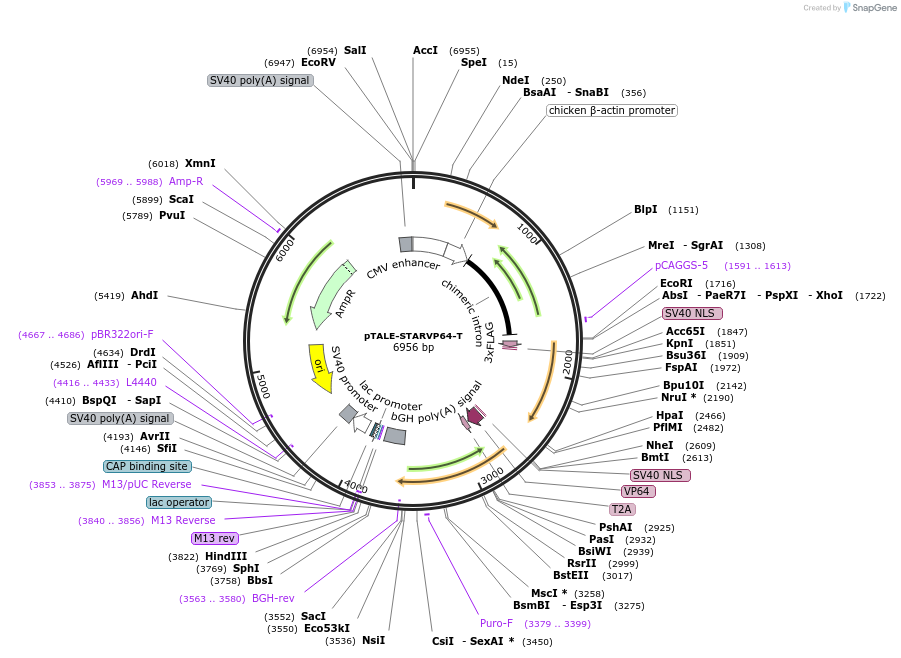
pTALE-STARVP64-T
Plasmid#87530PurposeDestination vector for TALE-VP64 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
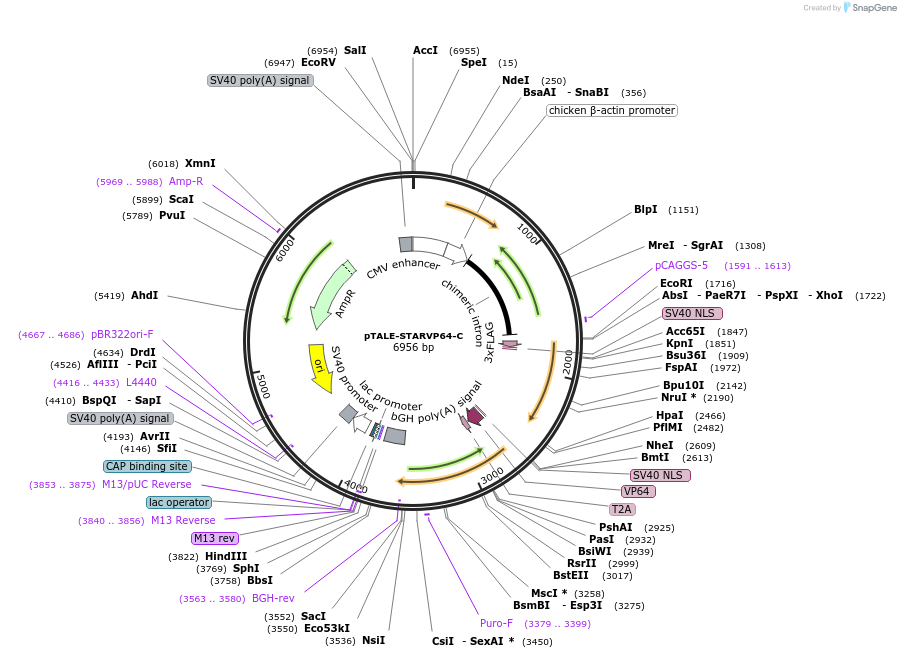
pTALE-STARVP64-C
Plasmid#87532PurposeDestination vector for TALE-VP64 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
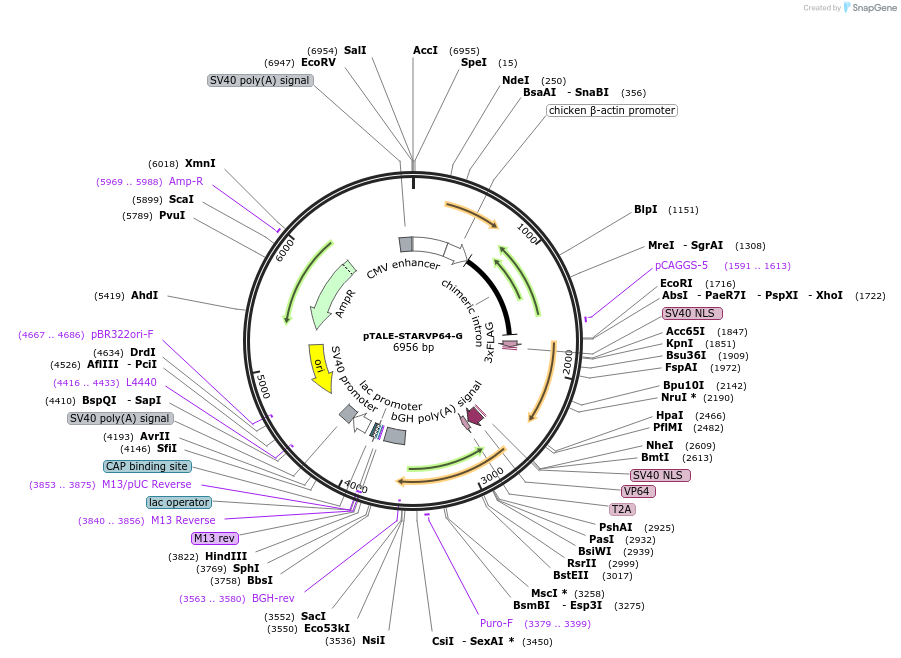
pTALE-STARVP64-G
Plasmid#87533PurposeDestination vector for TALE-VP64 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
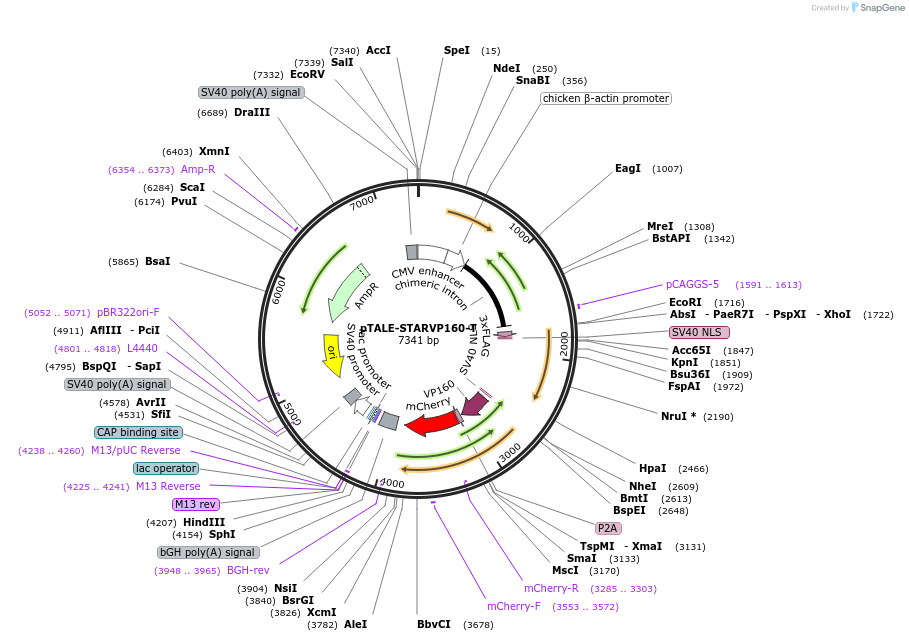
pTALE-STARVP160-T
Plasmid#87534PurposeDestination vector for TALE-VP160 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
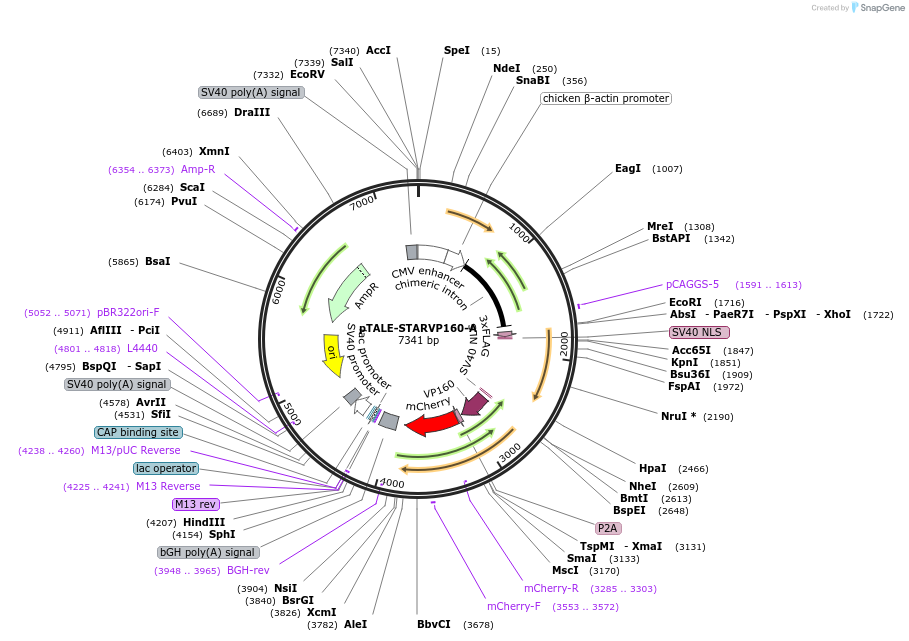
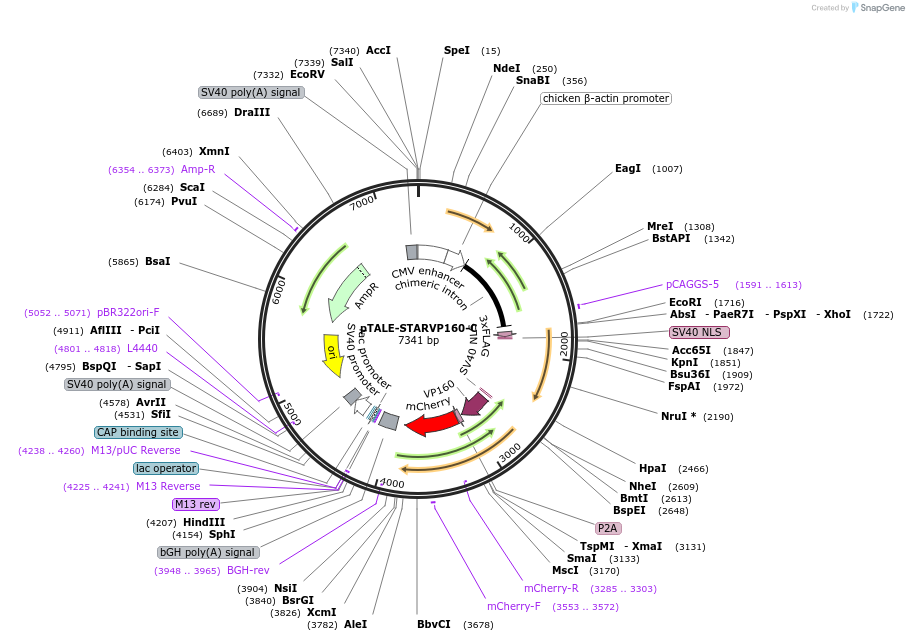
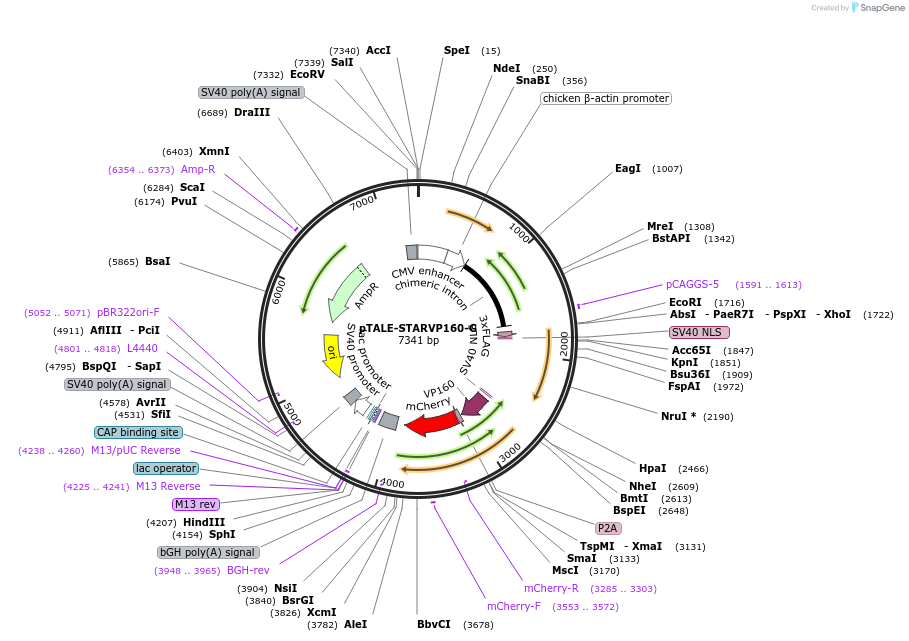
pTALE-STARVP160-A
Plasmid#87535PurposeDestination vector for TALE-VP160 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
pTALE-STARVP160-C
Plasmid#87536PurposeDestination vector for TALE-VP160 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
pTALE-STARVP160-G
Plasmid#87537PurposeDestination vector for TALE-VP160 constructionDepositorTypeEmpty backboneUseSynthetic Biology and TALEN; Tale-tf (activator)Available SinceJuly 13, 2017AvailabilityAcademic Institutions and Nonprofits only