We narrowed to 29,083 results for: Tat
-
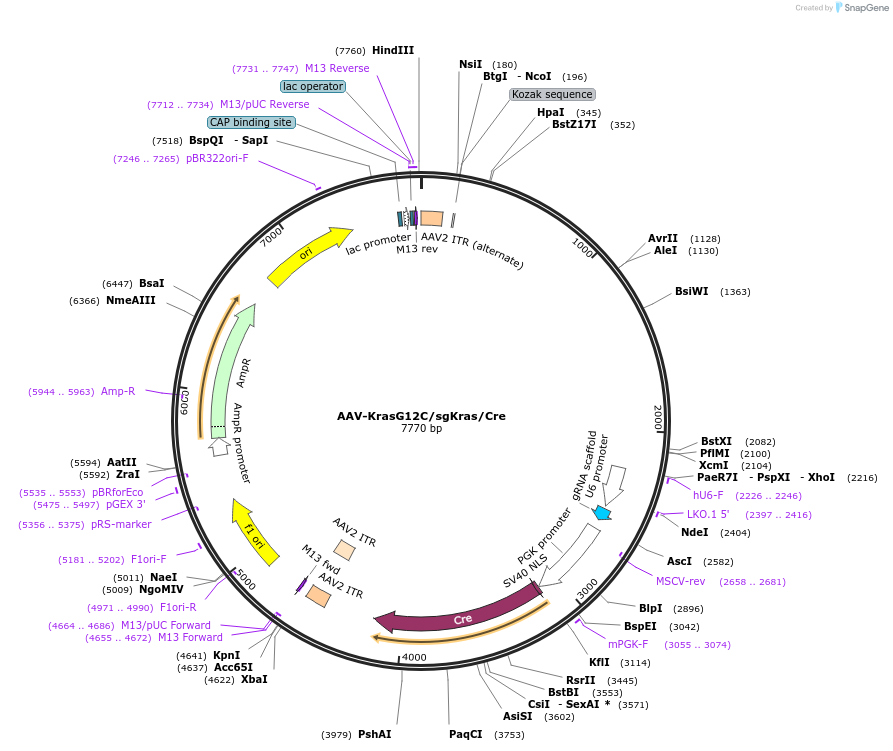
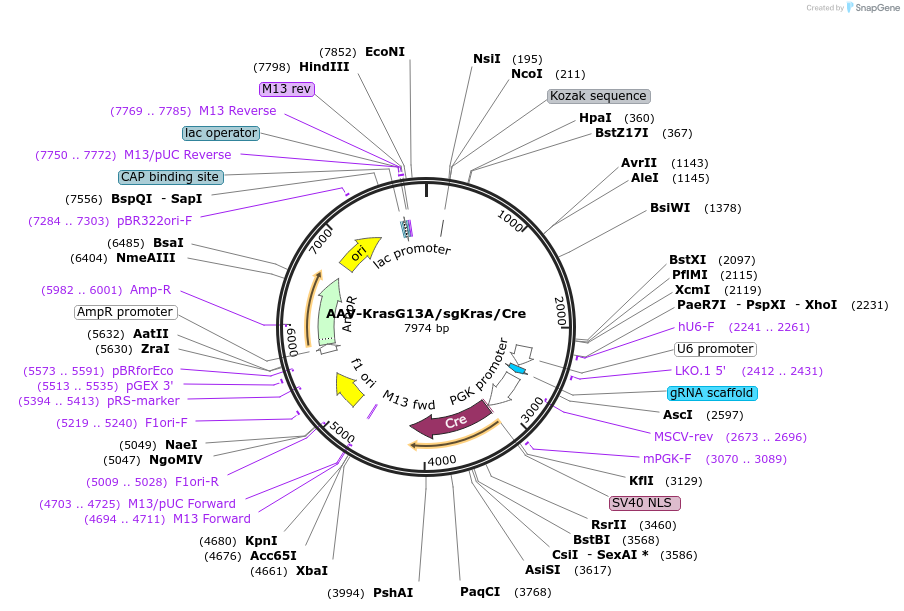
Plasmid#99850PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G12C mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
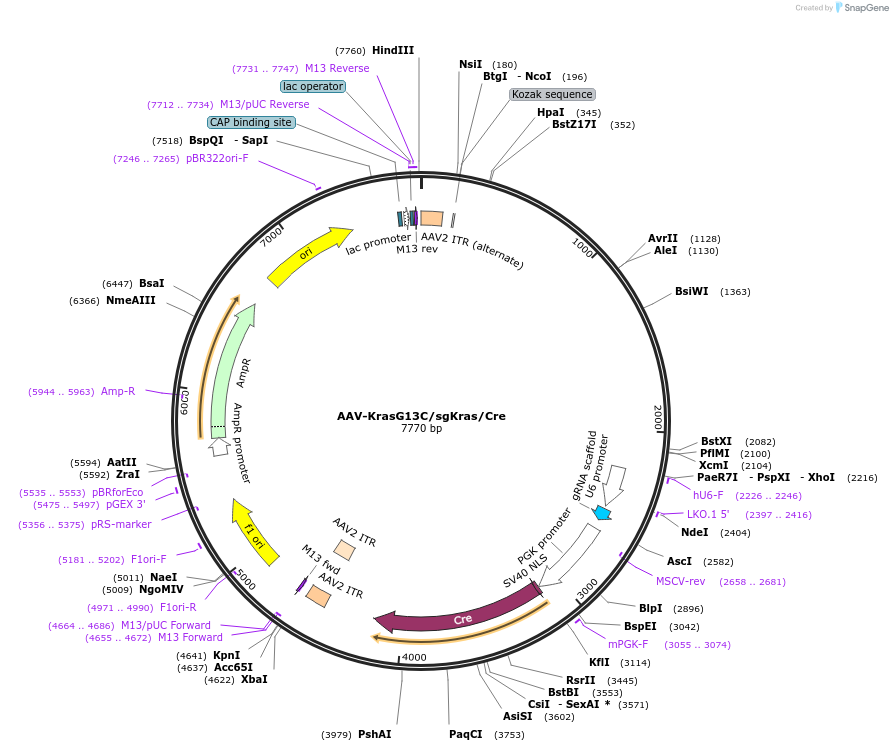
AAV-KrasG13C/sgKras/Cre
Plasmid#99856PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G13C mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
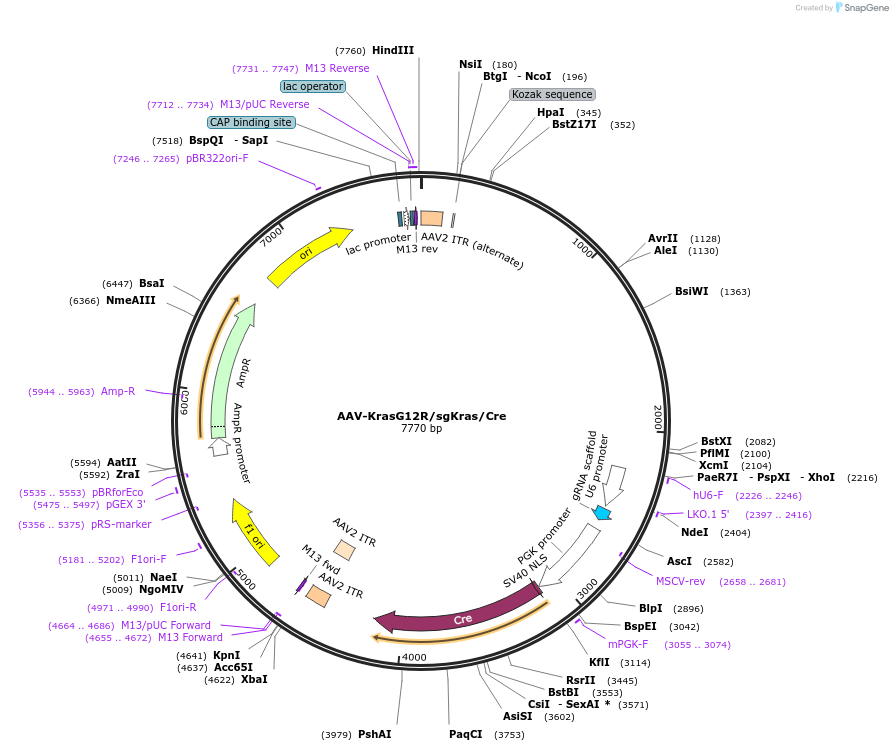
AAV-KrasG12R/sgKras/Cre
Plasmid#99852PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G12R mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
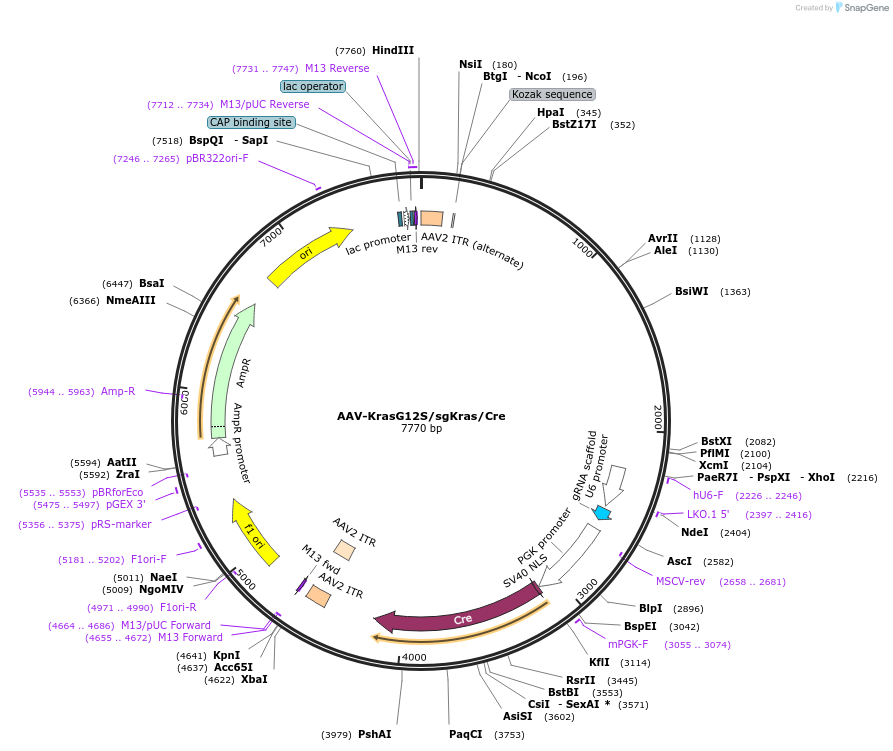
AAV-KrasG12S/sgKras/Cre
Plasmid#99853PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G12S mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
AAV-KrasG12A/sgKras/Cre
Plasmid#99849PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G12A mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
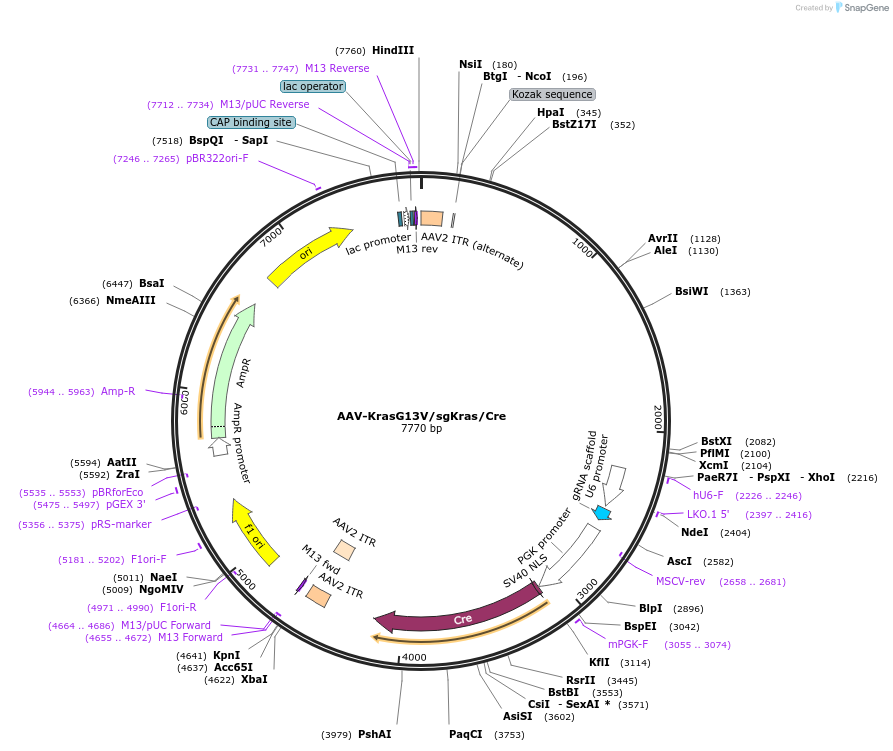
AAV-KrasG13V/sgKras/Cre
Plasmid#99860PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G13V mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceDec. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
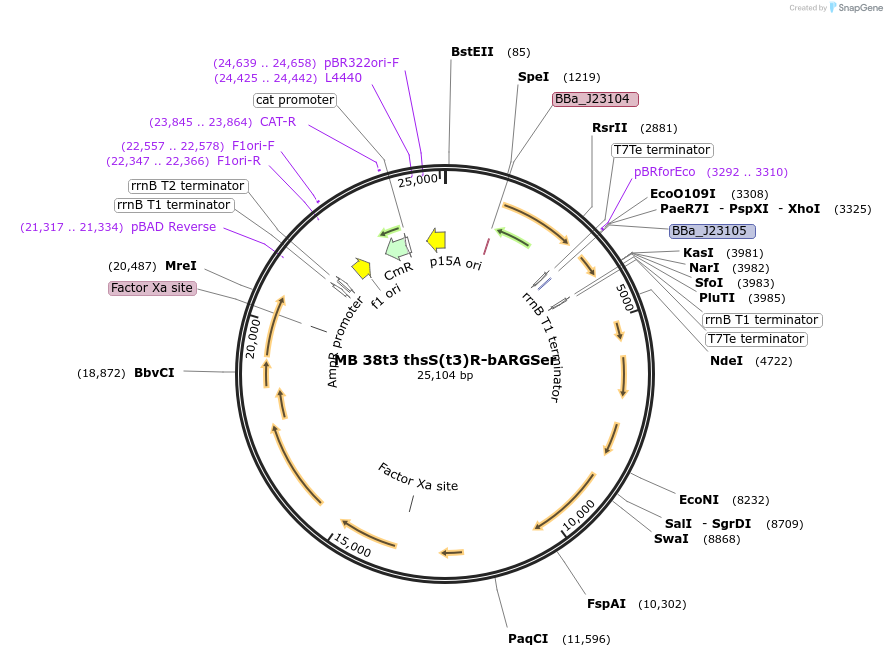
MB 38t3 thsS(t3)R-bARGSer
Plasmid#232468PurposeOptimized thiosulfate sensor with acoustic reporter genes (bARGSer)DepositorInsertsthsS(t3)
thsR
PphsA342
bARGSer
AxeTxe
ExpressionBacterialMutationContains the following mutations K286Q, Q350H, I…PromoterJ23104 (ttgacagctagctcagtcctaggtattgtgctagc), J23…Available SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
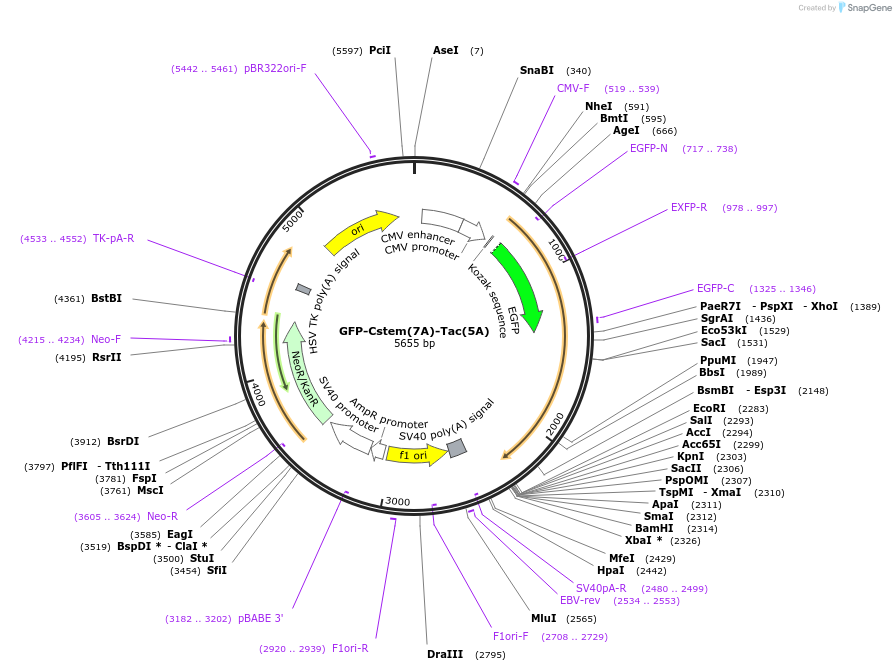
GFP-Cstem(7A)-Tac(5A)
Plasmid#162499PurposeExpresses GFP tagged Tac mutant in mammalian cells. The five O-glycosylation sites in the stem region are mutated, and the stem region of CD8a with mutations is inserted between GFP and Tac(5A).DepositorInsertTagsGFPExpressionMammalianMutationThe five theronine and two serine residues are mu…Available SinceJan. 29, 2021AvailabilityAcademic Institutions and Nonprofits only -
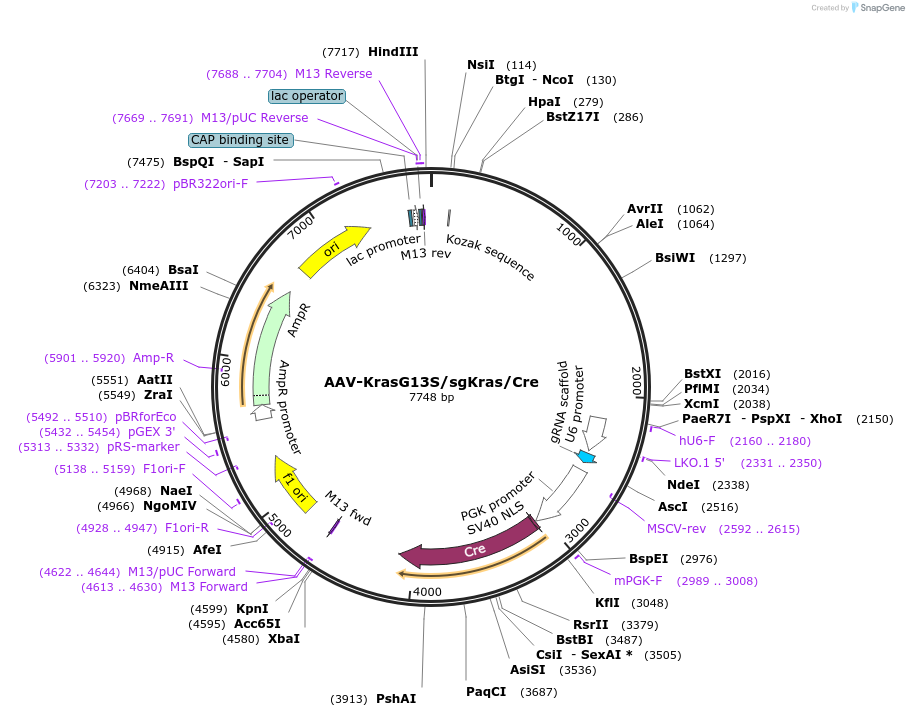
AAV-KrasG13S/sgKras/Cre
Plasmid#99859PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G13S mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceMay 10, 2018AvailabilityAcademic Institutions and Nonprofits only -
AAV-KrasG13A/sgKras/Cre
Plasmid#99855PurposeExpresses Cre-recombinase, a barcoded HDR template to introduce a G13A mutation into the endogenous Kras locus, and a Kras-targeting sgRNA to enhance HDR.DepositorInsertKras (Kras Mouse)
UseAAV and Cre/LoxExpressionMammalianMutationAvrII site (CAAAGG>CCTAGG), PAM/sgRNA mutation…Available SinceMay 10, 2018AvailabilityAcademic Institutions and Nonprofits only -
J23100_PheS*_codon
Plasmid#247070Purposeexpress codon optimized PheS* gene with constitutive promoter BBa_J23100DepositorInsertmutated (A294G) E.coli PheS of which codon is changed
ExpressionBacterialAvailable SinceMarch 6, 2026AvailabilityAcademic Institutions and Nonprofits only -
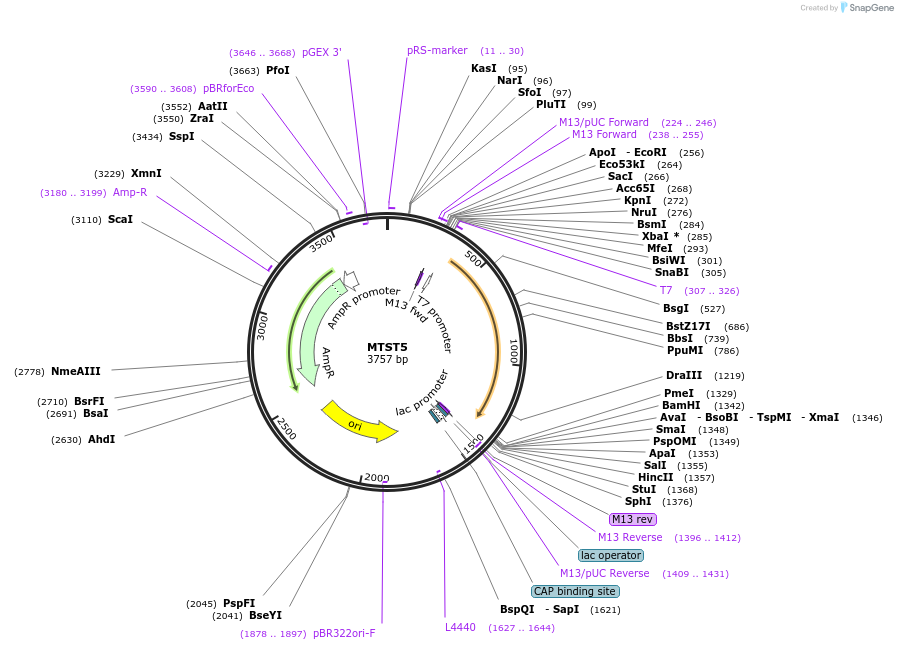
MTST5
Plasmid#86602PurposeCustom designed synthetic DNA fragment lacking homology to natural sequences for preparation of internal standards for RNA-seq in metatranscriptomics.DepositorInsertMTST5
UseInternal standard reverse transcriptionPromoterT7Available SinceSept. 15, 2017AvailabilityIndustry, Academic Institutions, and Nonprofits -
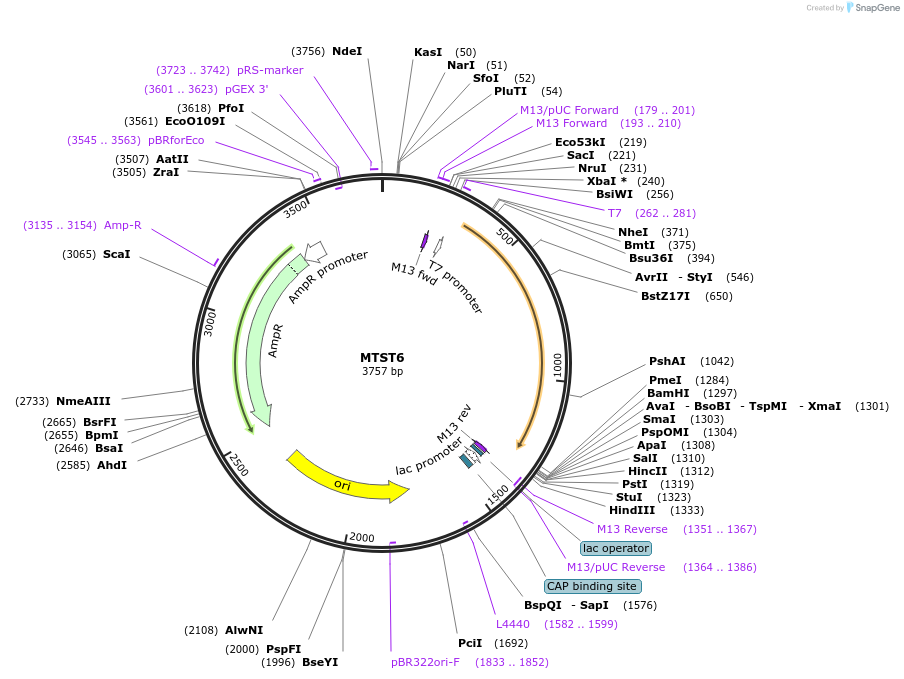
MTST6
Plasmid#86603PurposeCustom designed synthetic DNA fragment lacking homology to natural sequences for preparation of internal standards for RNA-seq in metatranscriptomics.DepositorInsertMTST6
ExpressionBacterialPromoterT7Available SinceSept. 15, 2017AvailabilityIndustry, Academic Institutions, and Nonprofits -
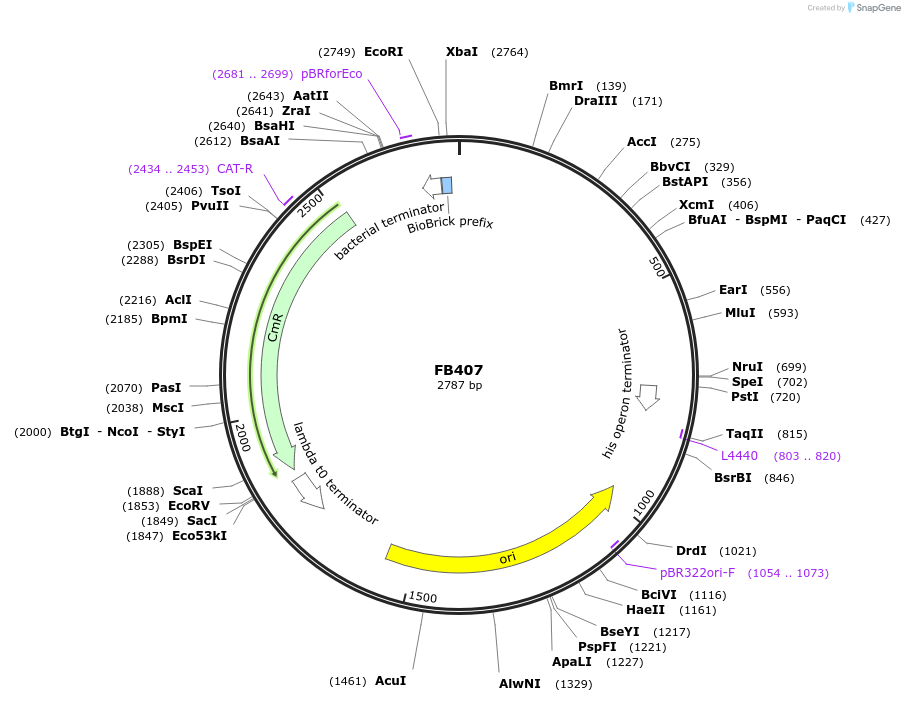
FB407
Plasmid#203628PurposePromoter of the elongation factor 1-alpha gene from P. digitatum (PDIG_59570).DepositorInsertPef1A
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
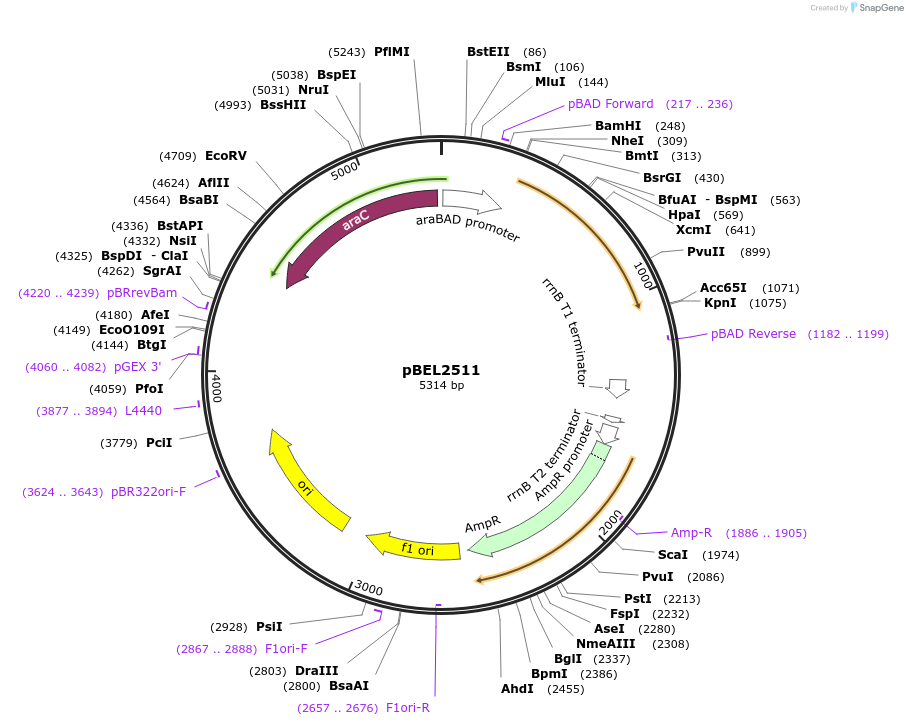
pBEL2511
Plasmid#195625PurposeComplementation of MlaA(WT)DepositorInsertMlaA
ExpressionBacterialAvailable SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
-
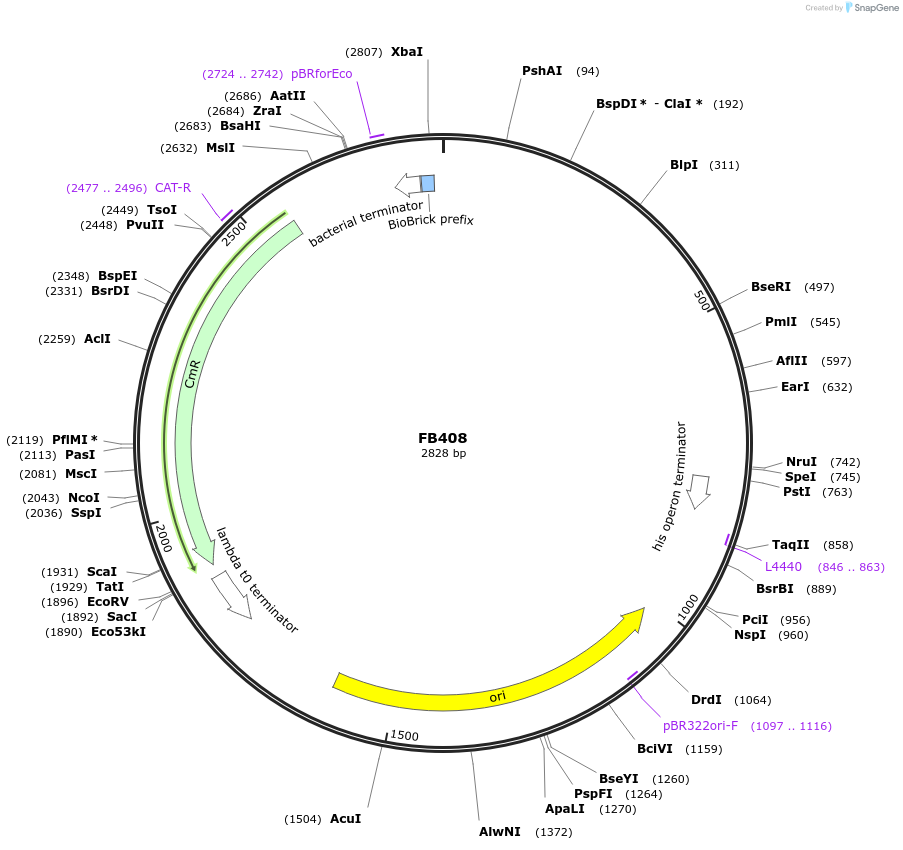
FB408
Plasmid#203629PurposePromoter of the ubiquitin ligase gene from P. digitatum (PDIG_07760).DepositorInsertP07760
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
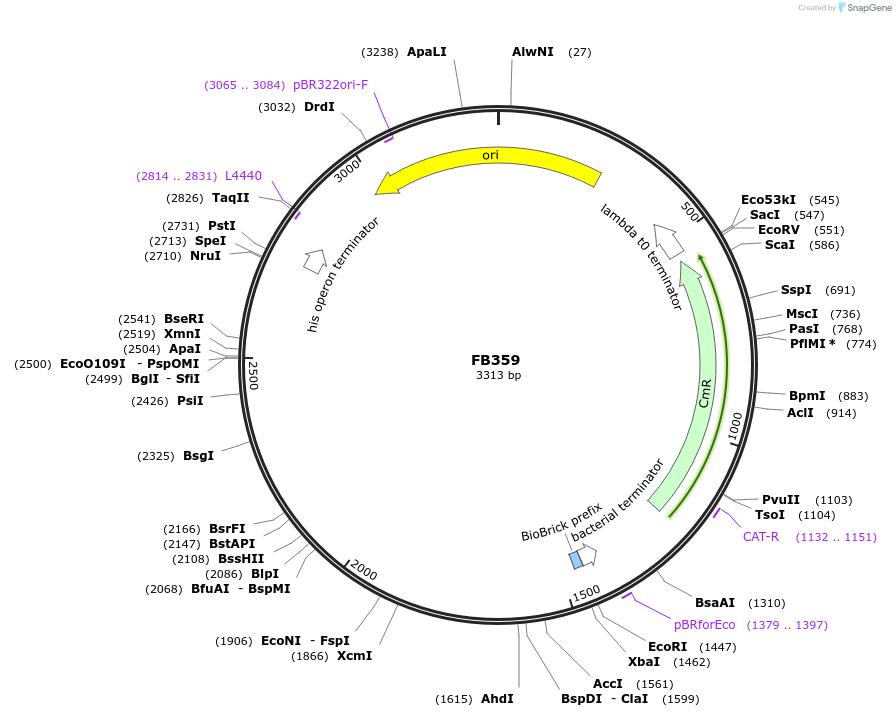
FB359
Plasmid#203614Purpose5' upstream region of pyrG gene in P. digitatum.DepositorInsert5' upstream Pdig pyrG
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
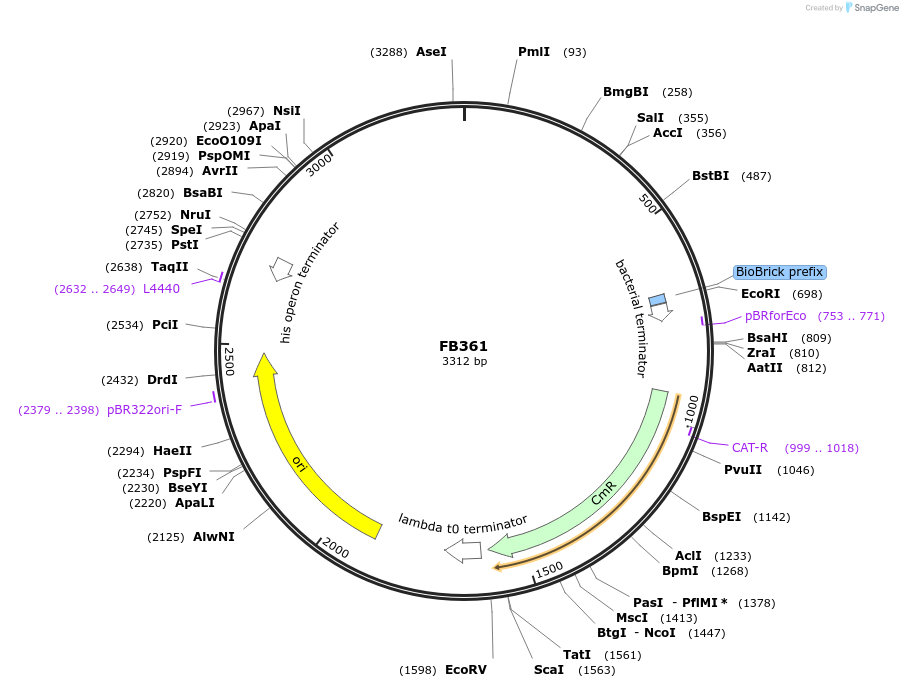
FB361
Plasmid#203615Purpose3' downstream region of pyrG gene in P. digitatum.DepositorInsert3' downstream Pdig pyrG
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
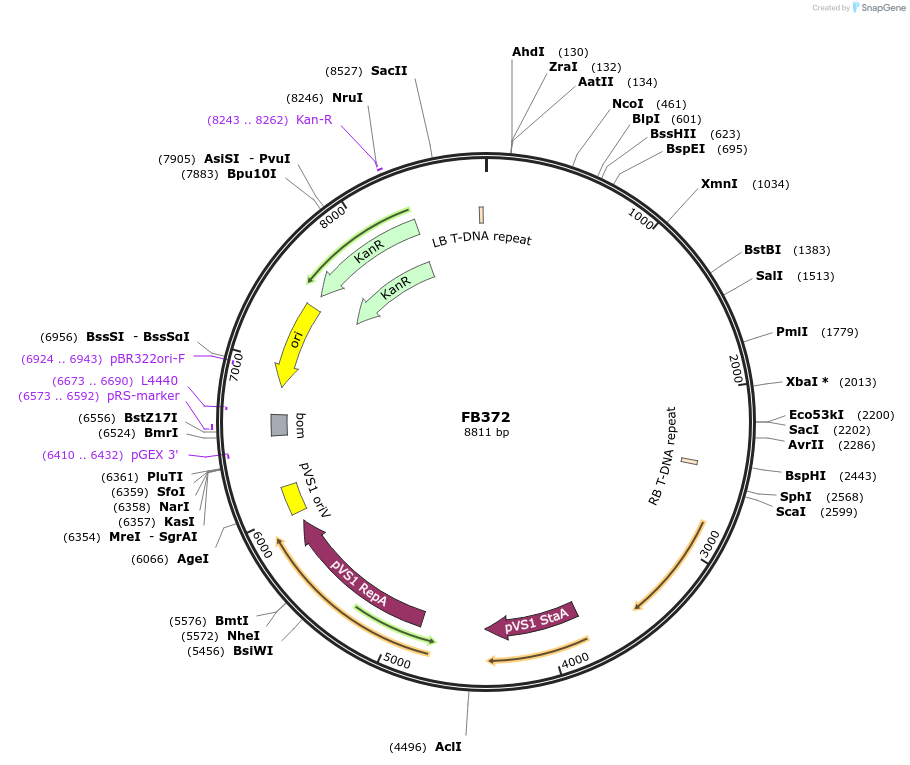
FB372
Plasmid#203616PurposeAssembly for pyrG deletion in P. digitatum.DepositorInsertFB359+FB361
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
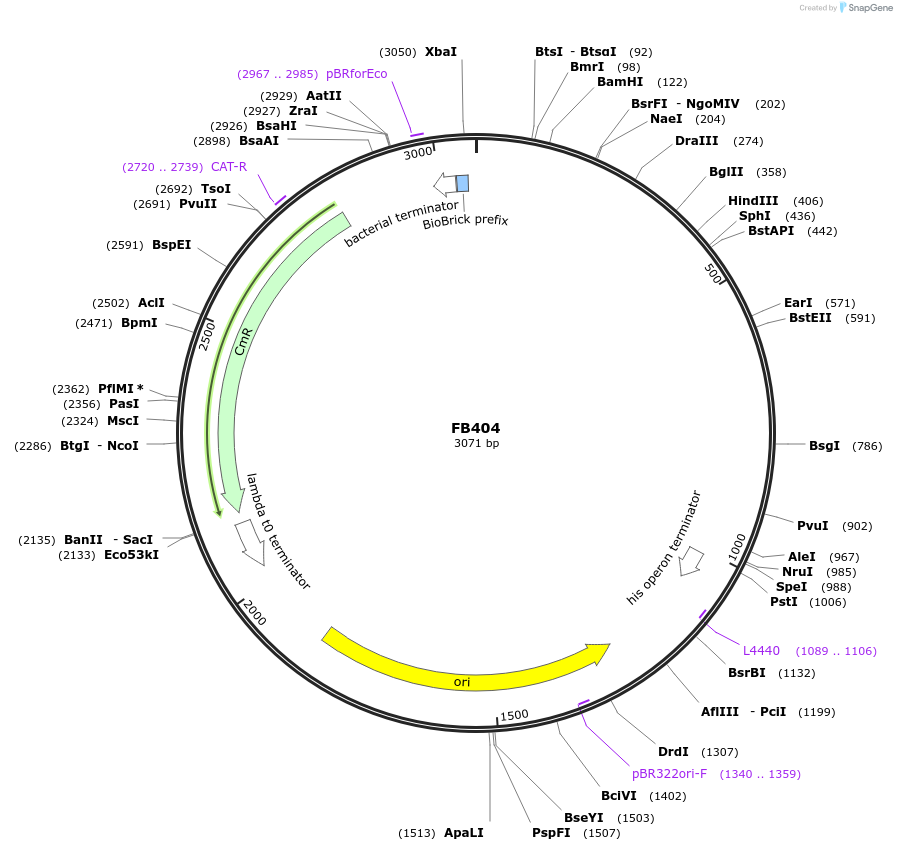
FB404
Plasmid#203625PurposePomoter of antifungal protein afpB gene from P. digitatum.DepositorInsertPafpB
UseSynthetic BiologyAvailable SinceSept. 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
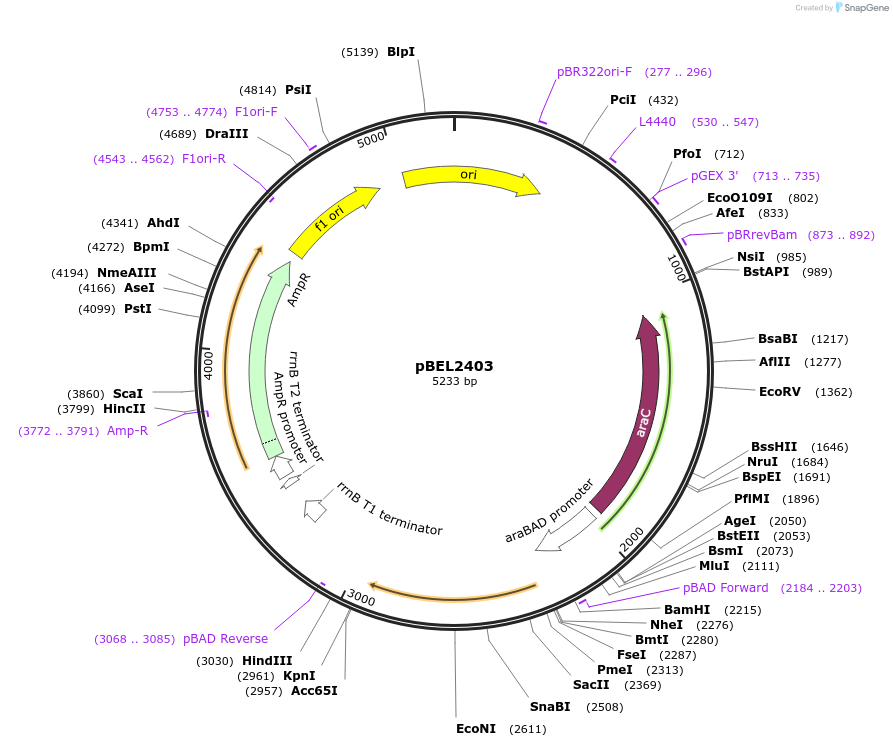
pBEL2403
Plasmid#195642PurposeComplementation of MlaC(WT)DepositorInsertMlaC
ExpressionBacterialAvailable SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
RSRF-Luc-2mut
Plasmid#31819DepositorInsertMutant MEF2-dependent promoter (-307 to -242 of Nur77/NR4A1)
UseLuciferaseExpressionMammalianMutationtwo CTATATTTAG RSRF sites mutated to CGATATTTCGPromotermutated MEF2-dependent (-307 to -242 of Nur77/NR4…Available SinceAug. 29, 2011AvailabilityAcademic Institutions and Nonprofits only -
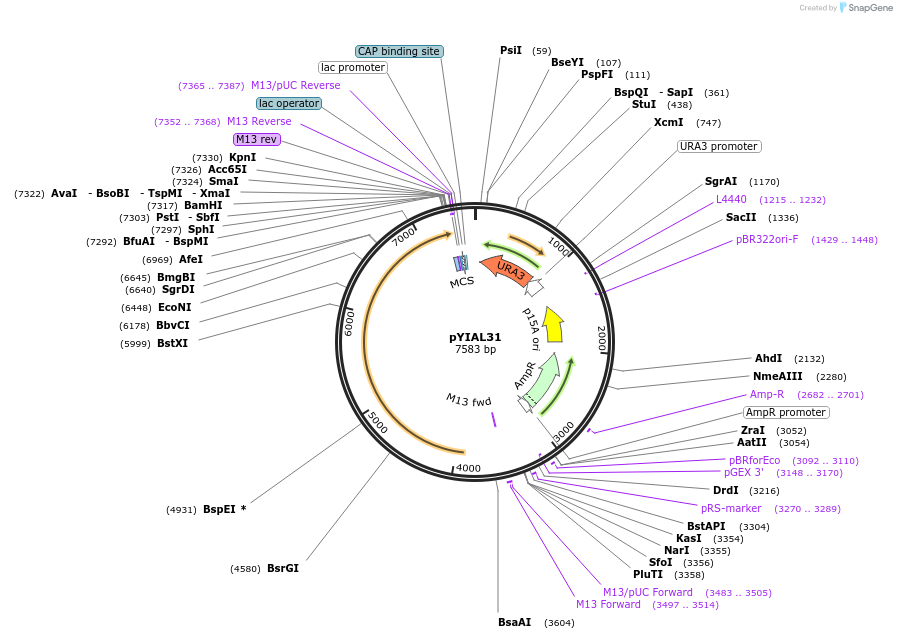
pYIAL31
Plasmid#60807Purposeused to make mutations in yeast POL1DepositorAvailable SinceFeb. 6, 2015AvailabilityAcademic Institutions and Nonprofits only -
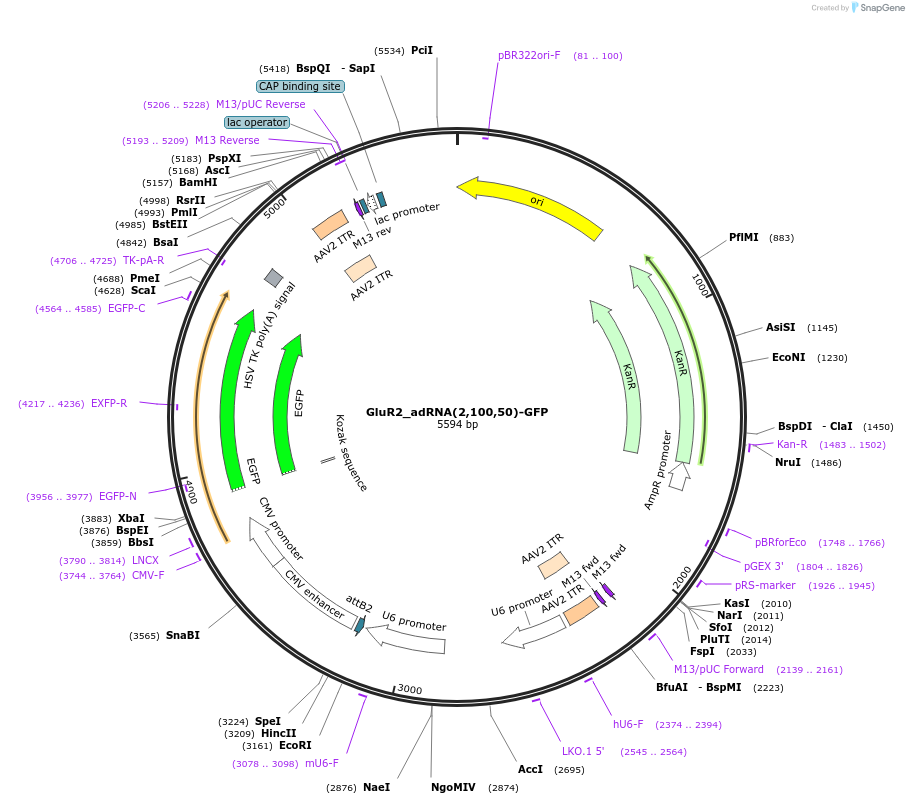
GluR2_adRNA(2,100,50)-GFP
Plasmid#124622PurposeExpresses adRNA(2,100,50) targeting the RAB7A transcript along with GFPDepositorInsertGluR2_adRNA(2,100,50)
UseAAVExpressionMammalianPromoterU6Available SinceMay 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
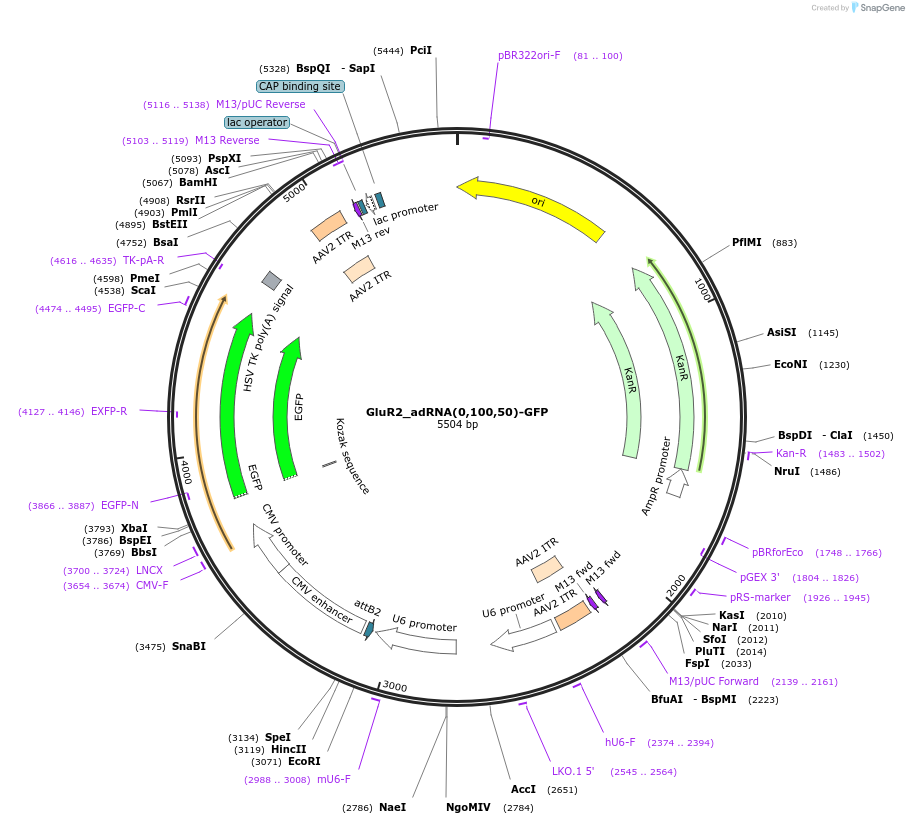
GluR2_adRNA(0,100,50)-GFP
Plasmid#124623PurposeExpresses adRNA(0,100,50) targeting the RAB7A transcript along with GFPDepositorInsertGluR2_adRNA(0,100,50)
UseAAVExpressionMammalianPromoterU6Available SinceMay 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
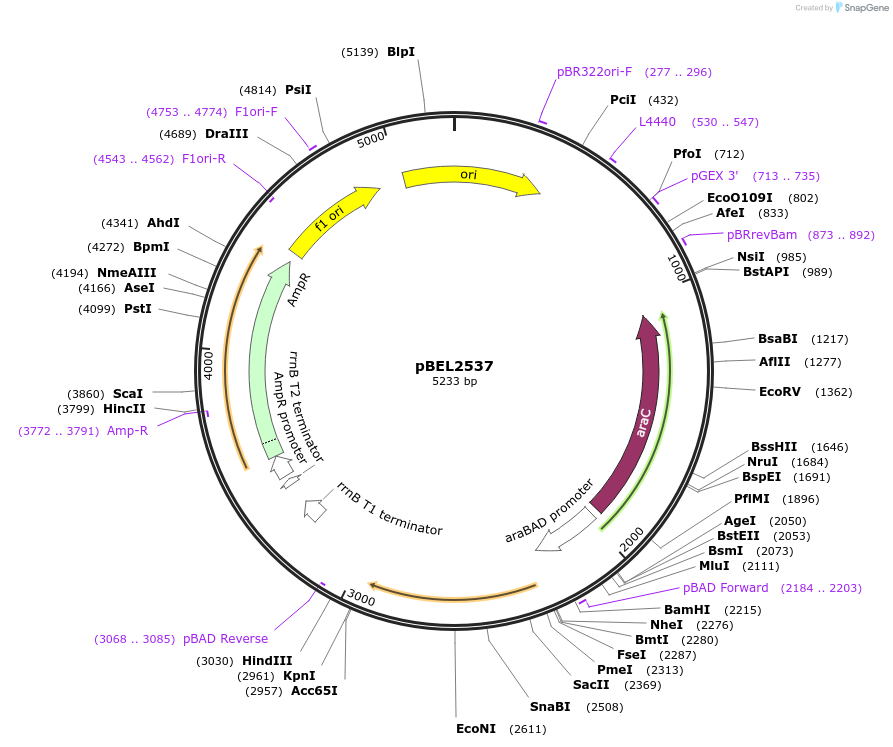
pBEL2537
Plasmid#195658PurposeComplementation of MlaC(R153A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R153A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
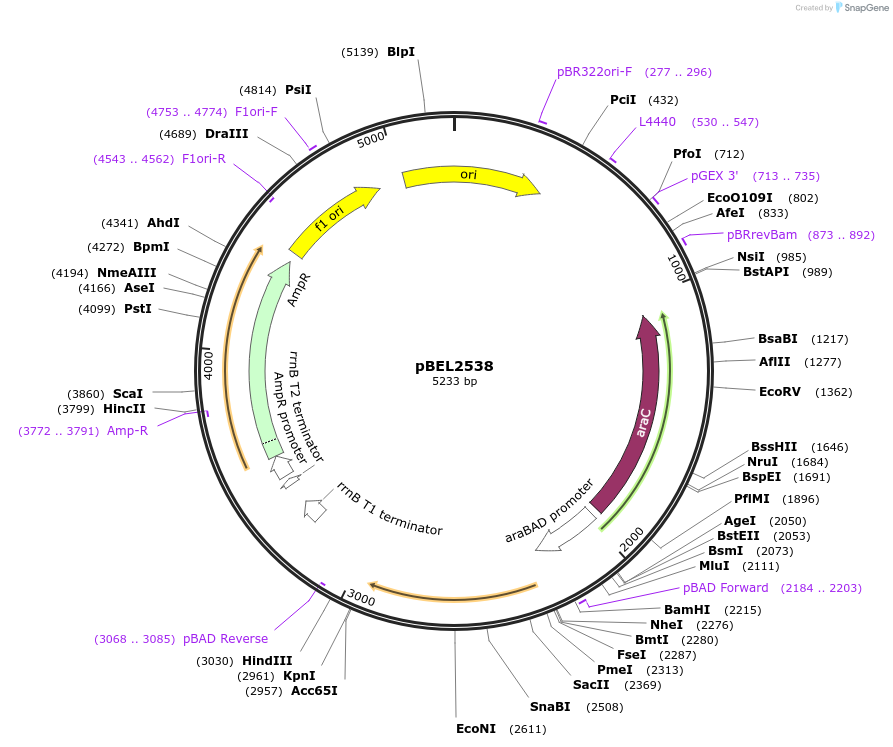
pBEL2538
Plasmid#195659PurposeComplementation of MlaC(R186A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R186A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2539
Plasmid#195660PurposeComplementation of MlaC(T116D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(T116D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
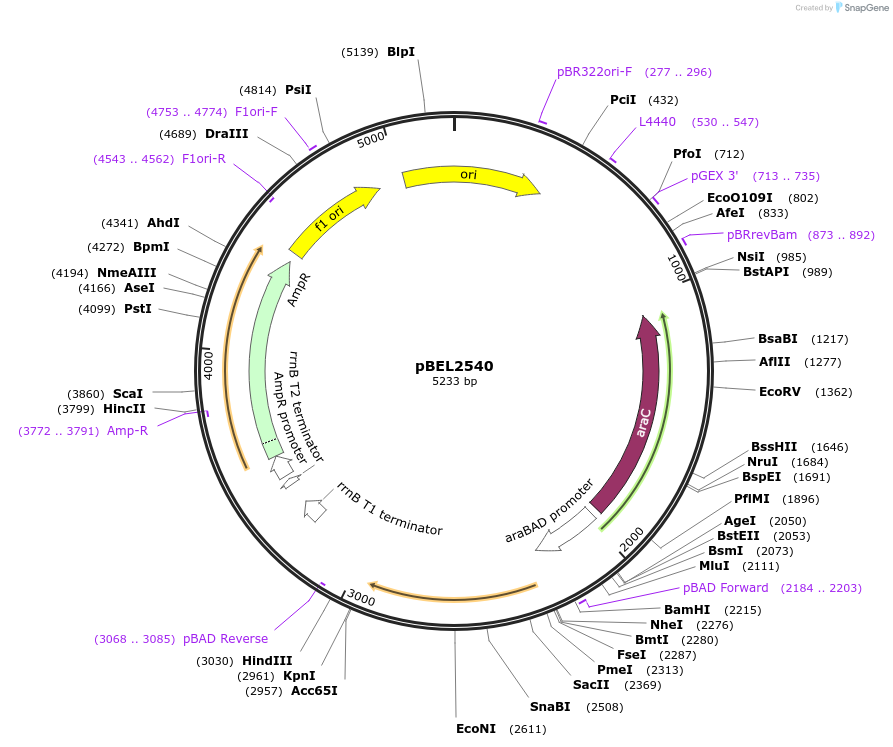
pBEL2540
Plasmid#195661PurposeComplementation of MlaC(T176R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(T176R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2541
Plasmid#195662PurposeComplementation of MlaC(V68R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V68R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
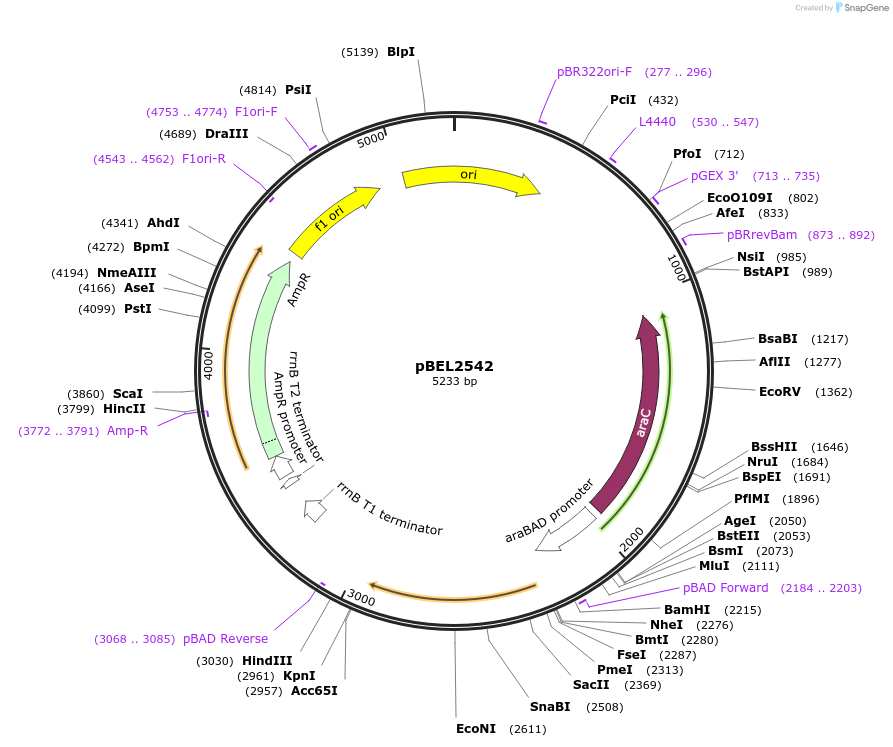
pBEL2542
Plasmid#195663PurposeComplementation of MlaC(V135R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V135R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
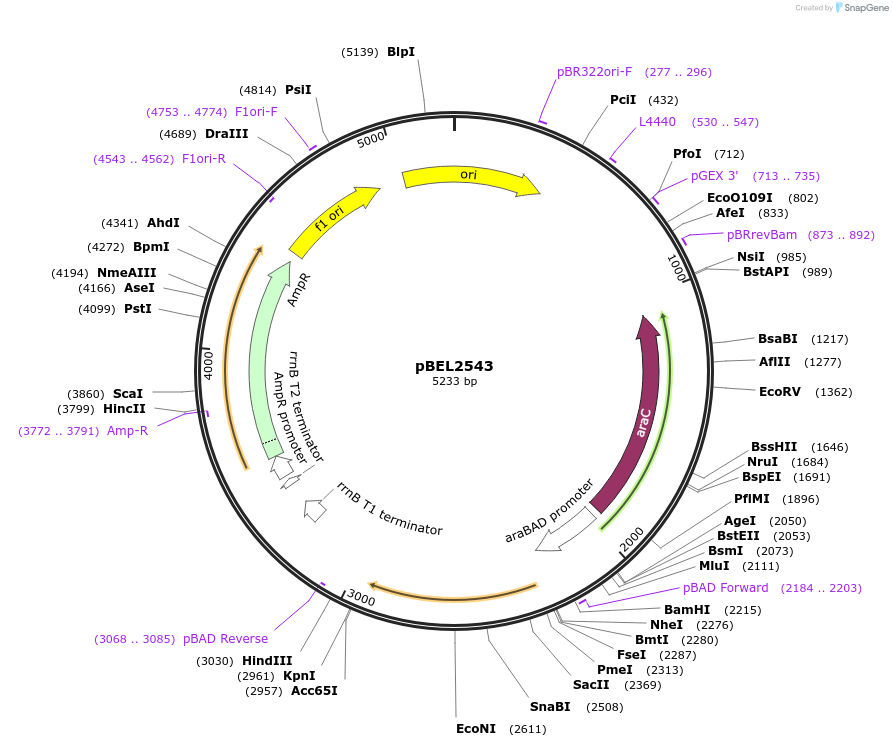
pBEL2543
Plasmid#195664PurposeComplementation of MlaC(V146R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V146R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
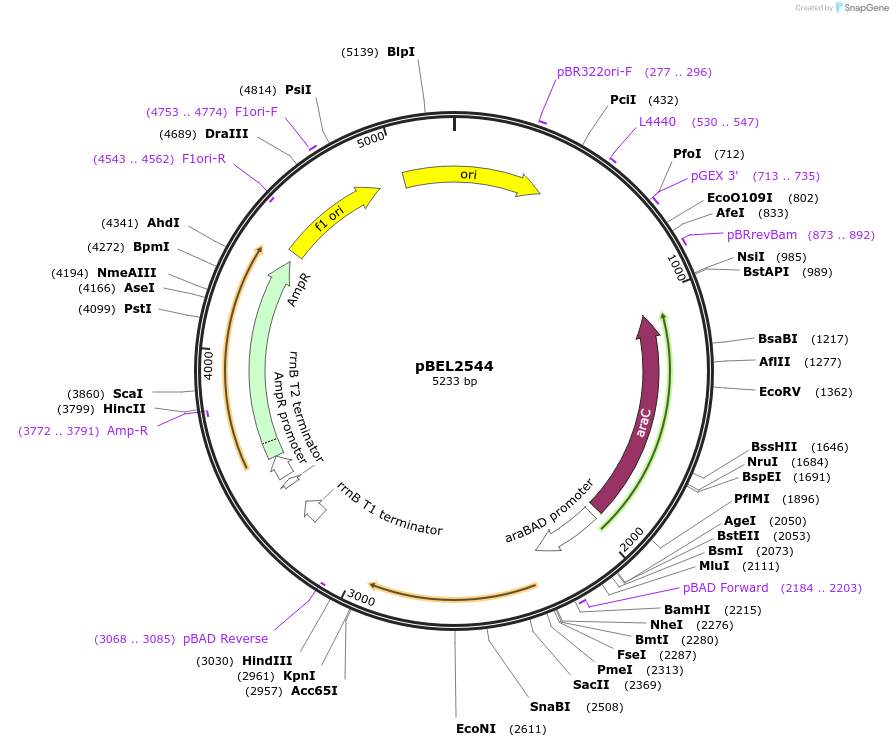
pBEL2544
Plasmid#195665PurposeComplementation of MlaC(V171R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V171R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
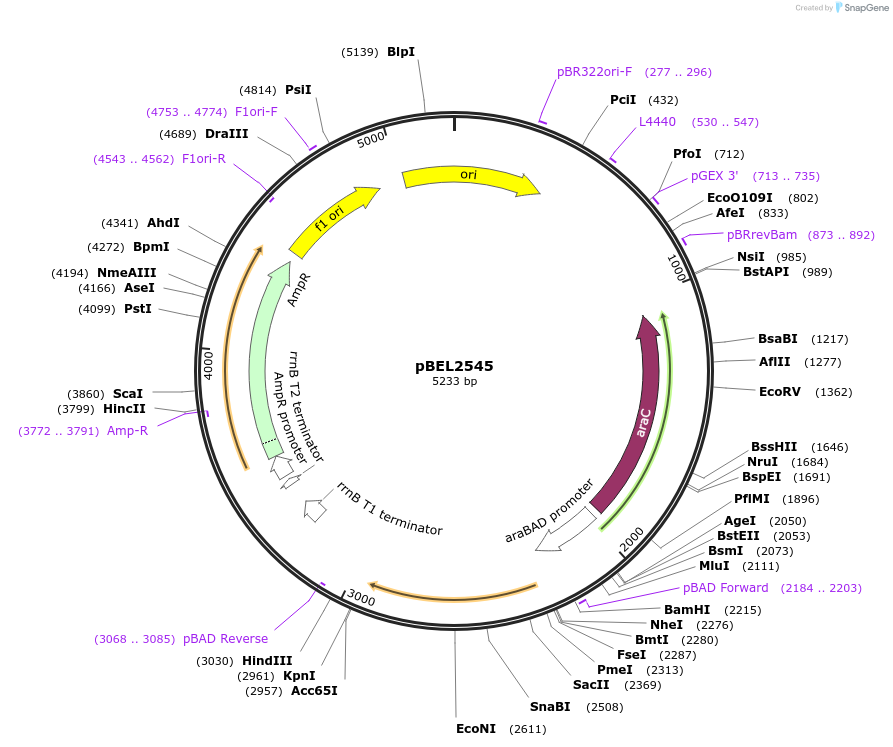
pBEL2545
Plasmid#195666PurposeComplementation of MlaC(Y72A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y72A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
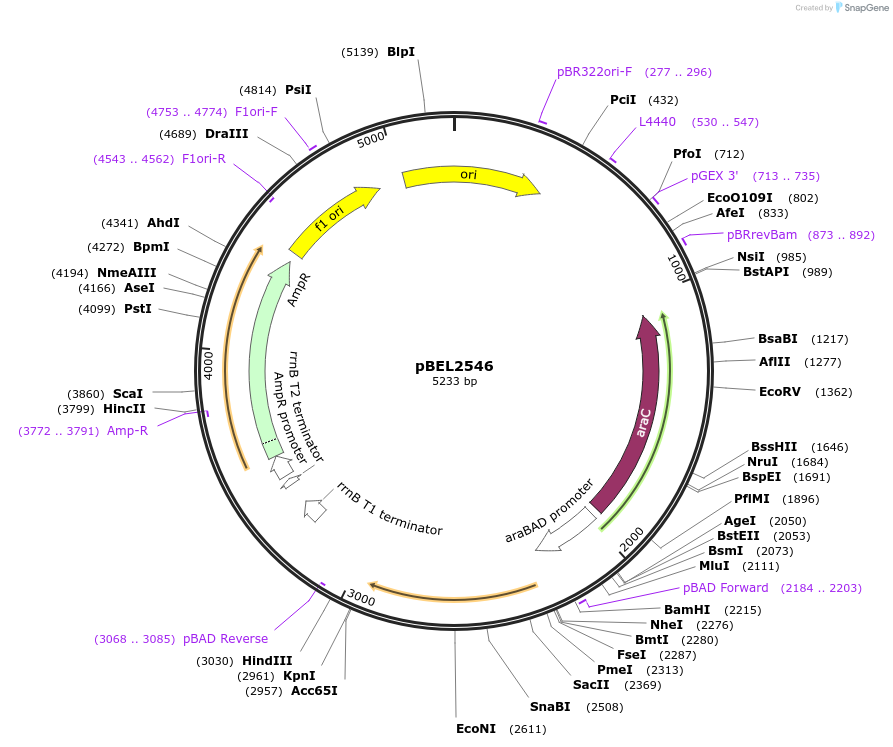
pBEL2546
Plasmid#195667PurposeComplementation of MlaC(Y82K)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y82K)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
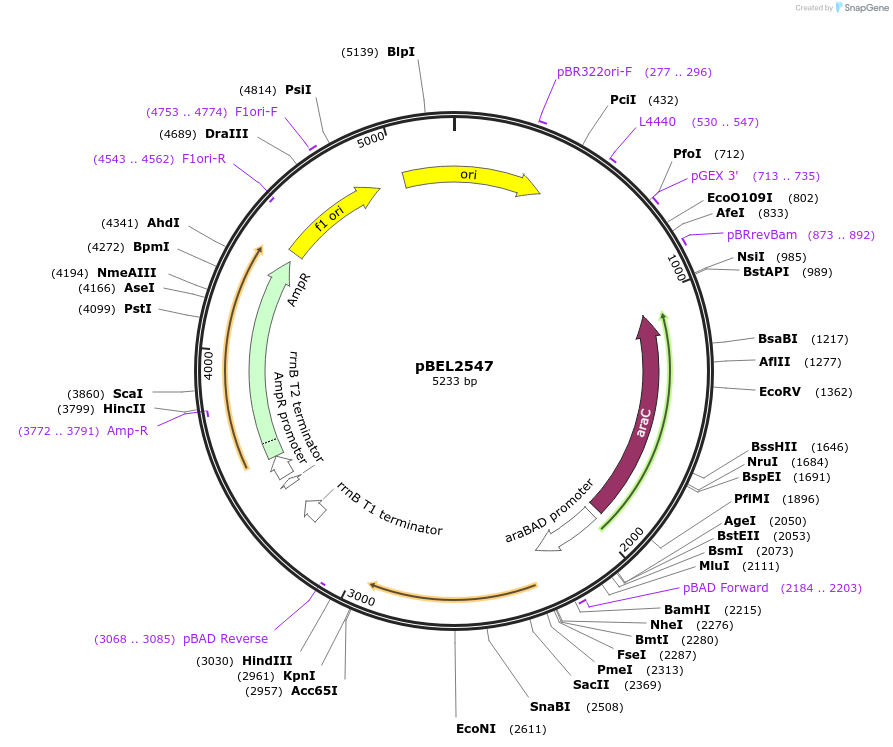
pBEL2547
Plasmid#195668PurposeComplementation of MlaC(Y105A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y105A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
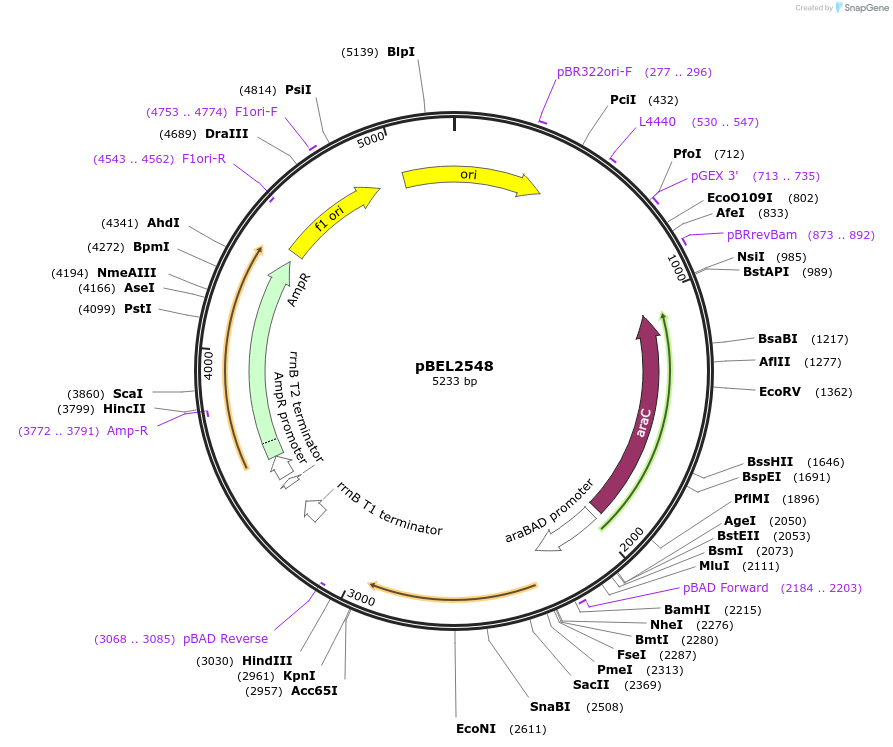
pBEL2548
Plasmid#195669PurposeComplementation of MlaC(Y164R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y164R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
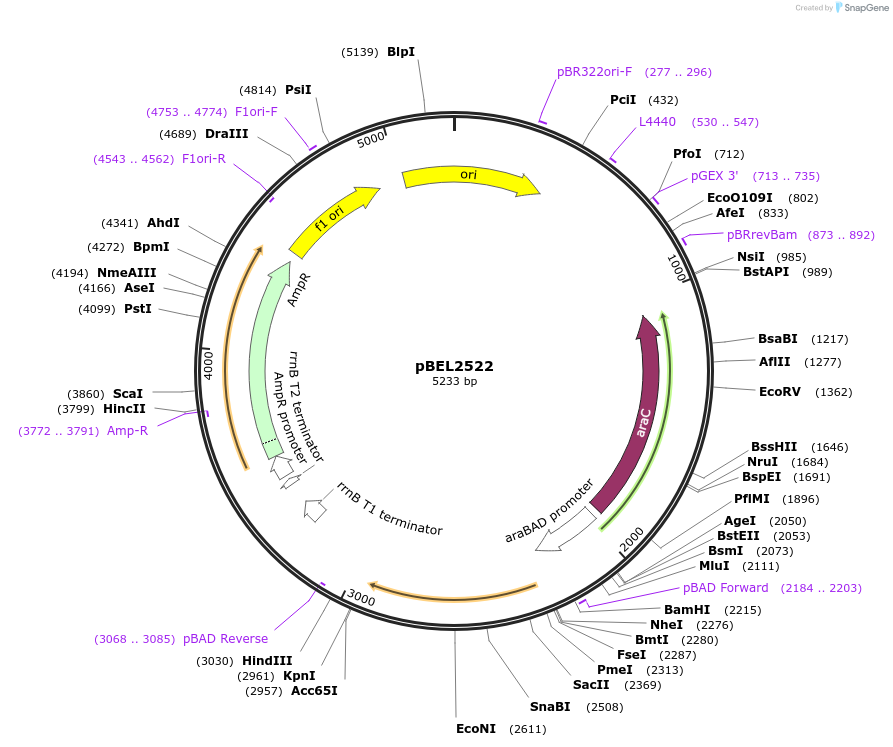
pBEL2522
Plasmid#195644PurposeComplementation of MlaC(A108R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A108R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
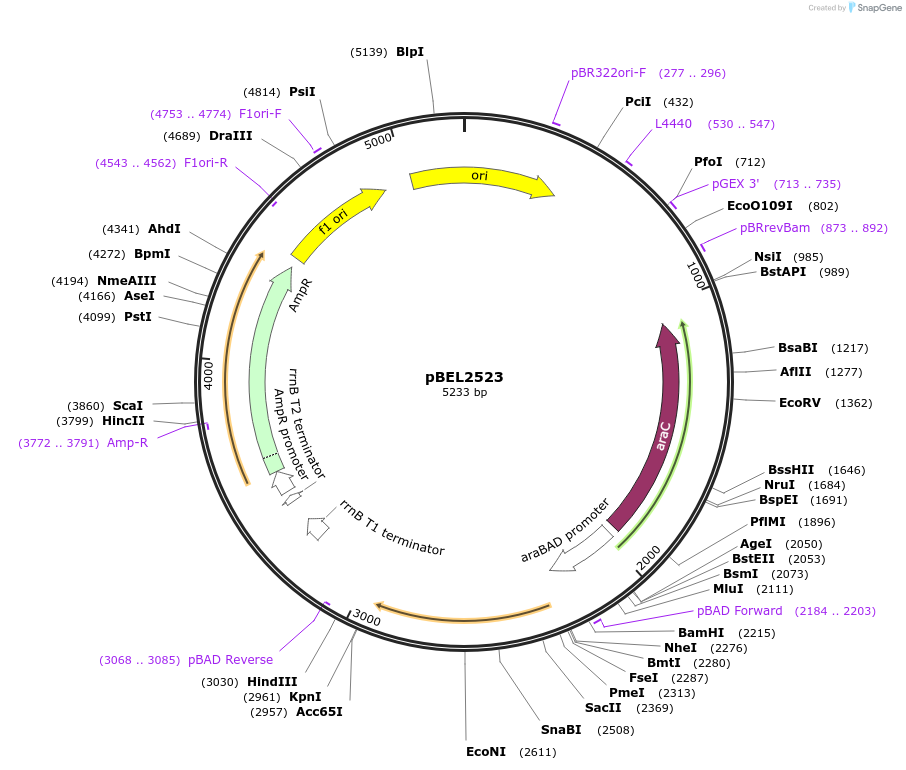
pBEL2523
Plasmid#195645PurposeComplementation of MlaC(A163R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A163R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only