We narrowed to 10,951 results for: T7
-
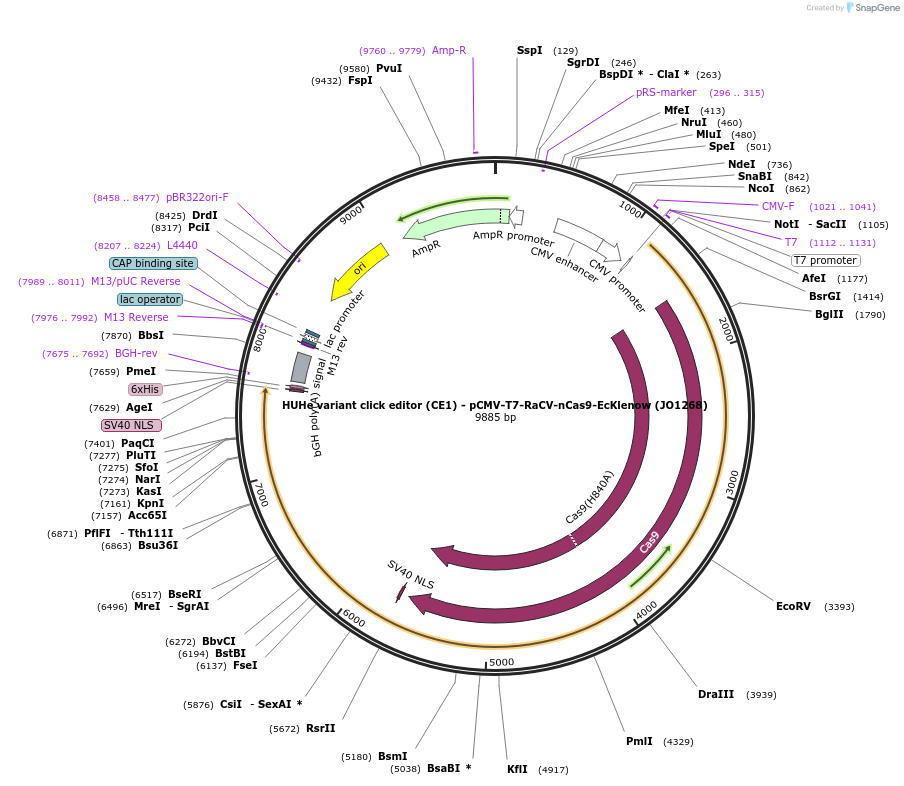
Plasmid#217795PurposeVariant CE1 construct with RaCV HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertRaCV-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
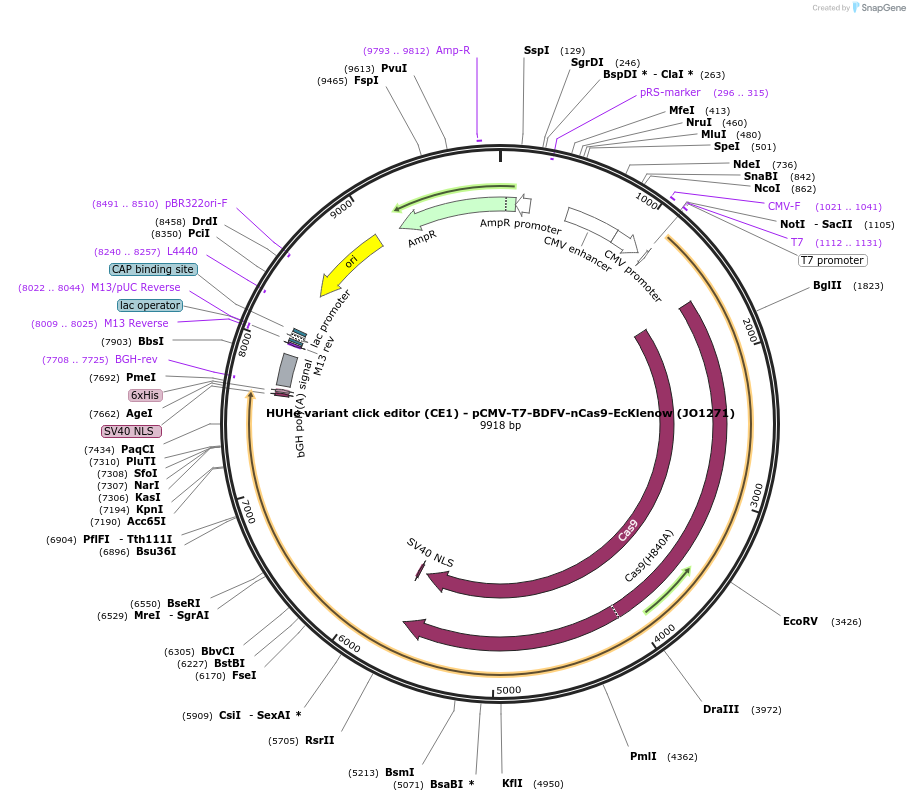
HUHe variant click editor (CE1) - pCMV-T7-BDFV-nCas9-EcKlenow (JO1271)
Plasmid#217785PurposeVariant CE1 construct with BDFV HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertBDFV-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
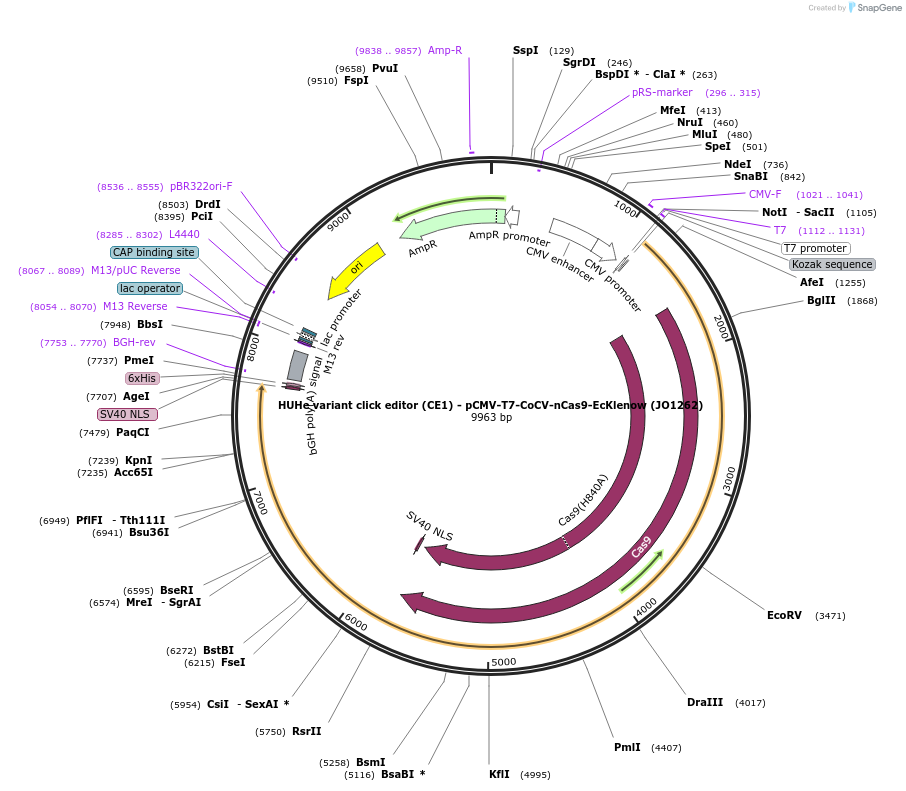
HUHe variant click editor (CE1) - pCMV-T7-CoCV-nCas9-EcKlenow (JO1262)
Plasmid#217786PurposeVariant CE1 construct with CoCV HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertCoCV-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
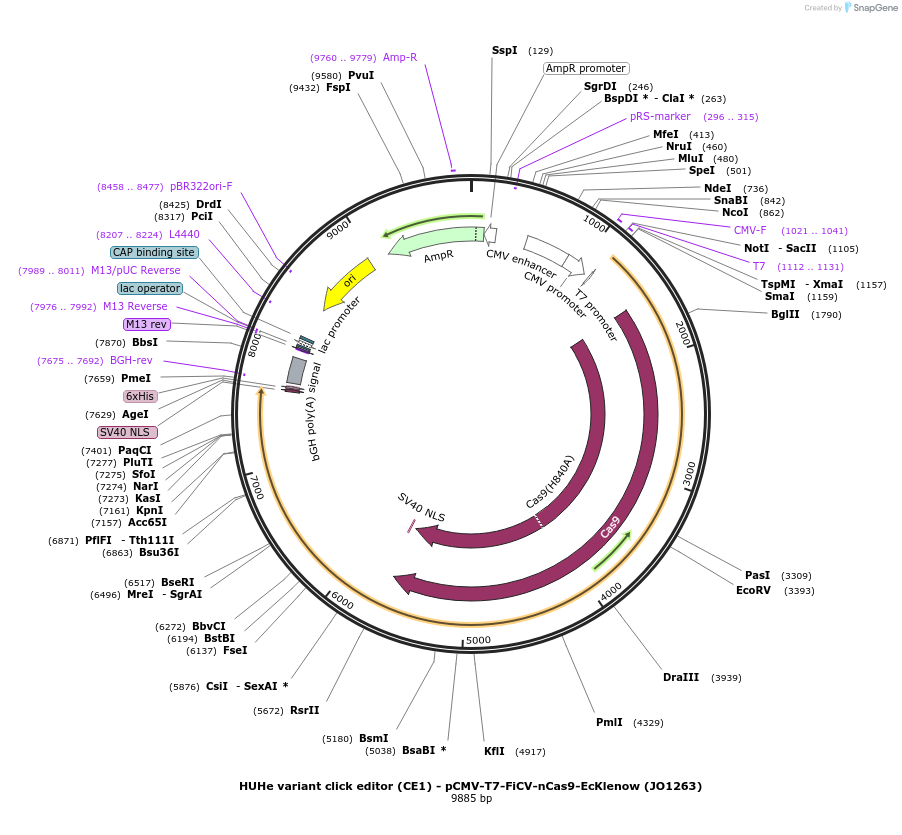
HUHe variant click editor (CE1) - pCMV-T7-FiCV-nCas9-EcKlenow (JO1263)
Plasmid#217787PurposeVariant CE1 construct with FiCV HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertFiCV-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
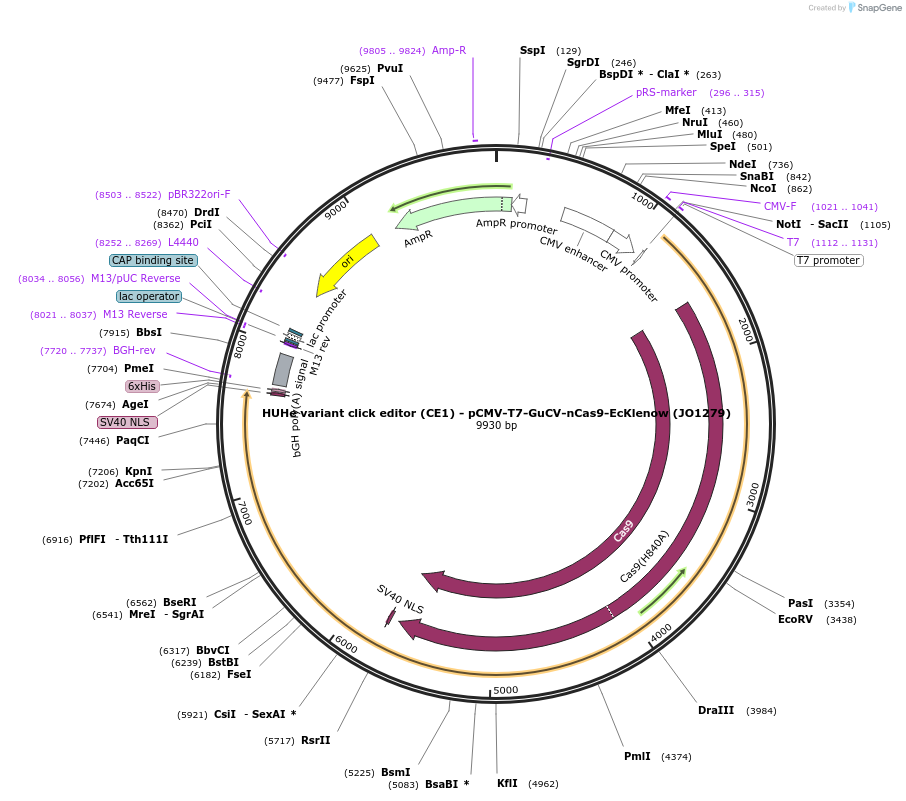
HUHe variant click editor (CE1) - pCMV-T7-GuCV-nCas9-EcKlenow (JO1279)
Plasmid#217788PurposeVariant CE1 construct with GuCV HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertGuCV-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
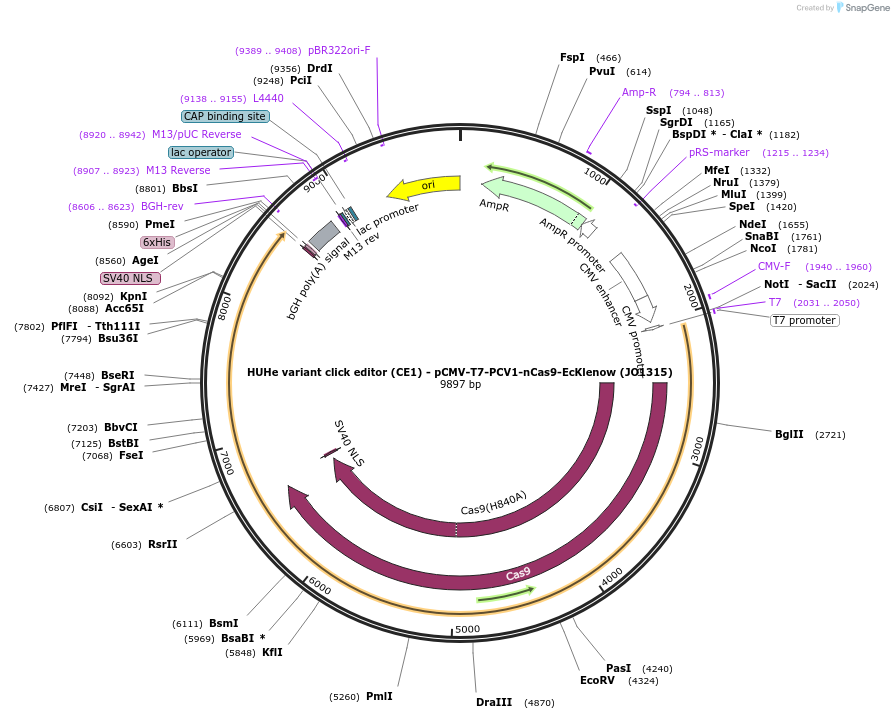
HUHe variant click editor (CE1) - pCMV-T7-PCV1-nCas9-EcKlenow (JO1315)
Plasmid#217789PurposeVariant CE1 construct with PCV1 HUH endonuclease, expressed from CMV or T7 promoters.DepositorInsertPCV1-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
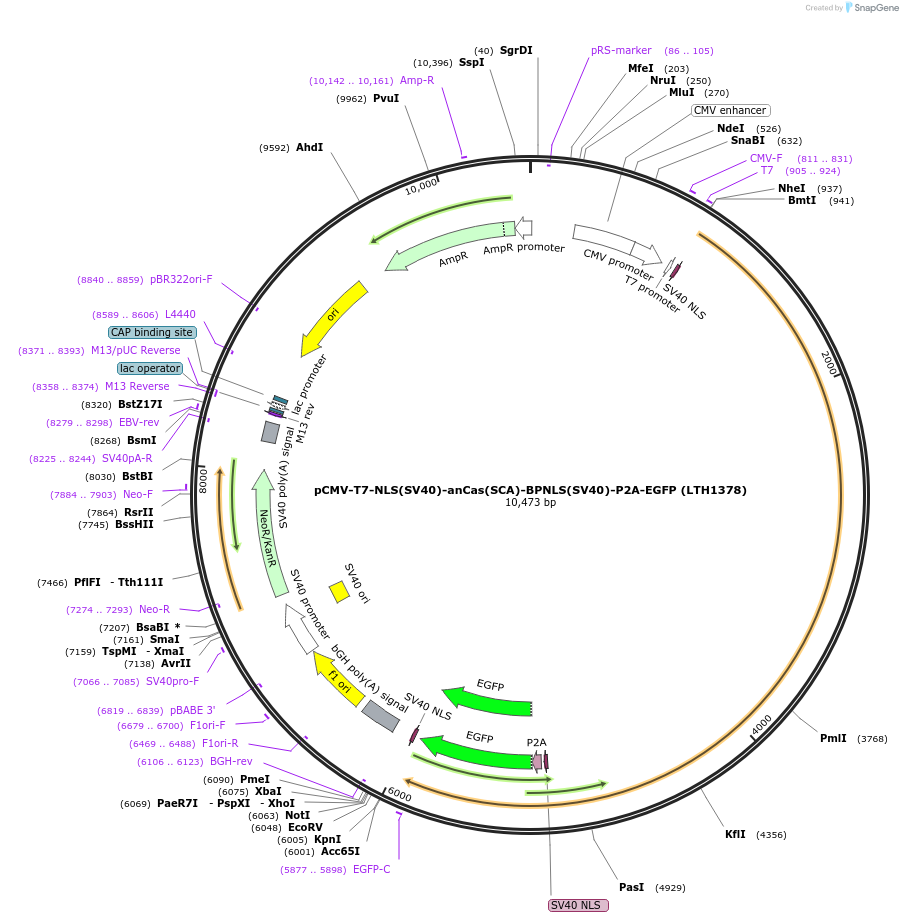
pCMV-T7-NLS(SV40)-anCas(SCA)-BPNLS(SV40)-P2A-EGFP (LTH1378)
Plasmid#185484PurposeCMV and T7 promoter expression plasmid for human codon usage anCas(SCA) with an N-terminal NLS, C-terminal bi-partite NLS, and P2A-eGFPDepositorInserthuman codon usage NLS-anCas(SCA)-BPNLS-P2A-eGFP
UseIn vitro transcription; t7 promoterTagsBPNLS-P2A-eGFP and NLSExpressionMammalianPromoterCMV and T7Available SinceJune 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
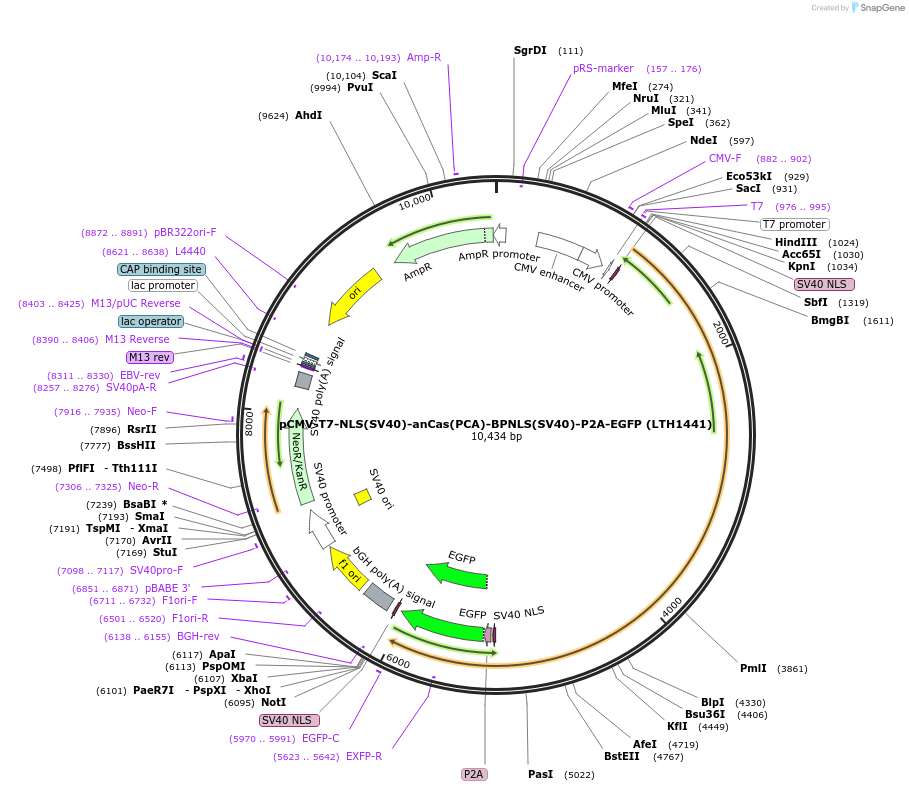
pCMV-T7-NLS(SV40)-anCas(PCA)-BPNLS(SV40)-P2A-EGFP (LTH1441)
Plasmid#185486PurposeCMV and T7 promoter expression plasmid for human codon usage anCas(PCA) with an N-terminal NLS, C-terminal bi-partite NLS, and P2A-eGFPDepositorInserthuman codon usage NLS-anCas(PCA)-BPNLS-P2A-eGFP
UseIn vitro transcription; t7 promoterTagsBPNLS-P2A-eGFP and NLSExpressionMammalianPromoterCMV and T7Available SinceJune 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
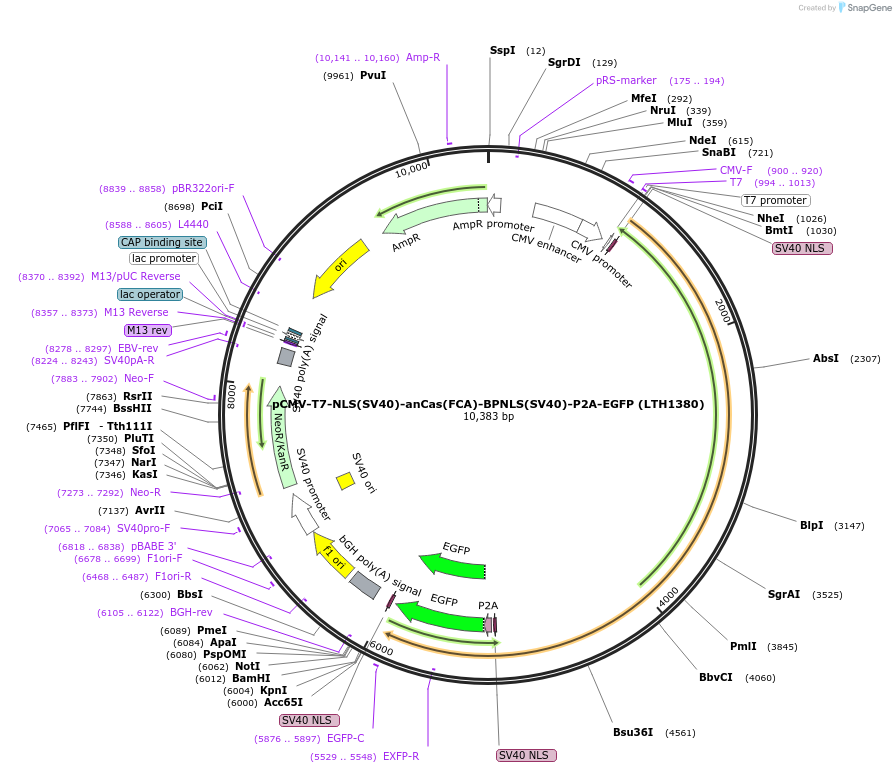
pCMV-T7-NLS(SV40)-anCas(FCA)-BPNLS(SV40)-P2A-EGFP (LTH1380)
Plasmid#185487PurposeCMV and T7 promoter expression plasmid for human codon usage anCas(FCA) with an N-terminal NLS, C-terminal bi-partite NLS, and P2A-eGFPDepositorInserthuman codon usage NLS-anCas(FCA)-BPNLS-P2A-eGFP
UseIn vitro transcription; t7 promoterTagsBPNLS-P2A-eGFP and NLSExpressionMammalianPromoterCMV and T7Available SinceJune 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
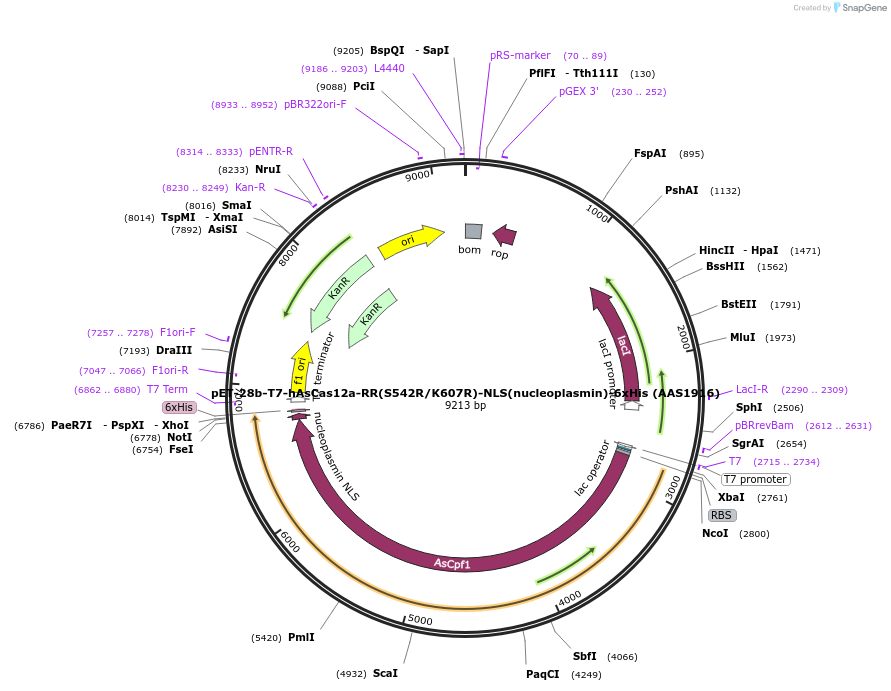
pET-28b-T7-hAsCas12a-RR(S542R/K607R)-NLS(nucleoplasmin)-6xHis (AAS1916)
Plasmid#114076PurposeT7 promoter bacterial expression plasmid for human codon optimized AsCas12a-RR(S542R/K607R) with C-terminal NLS and His-tagDepositorInserthuman codon optimized AsCas12a-RR (S542R/K607R) with C-terminal NLS and His-tag
TagsNLS(nucleoplasmin) and 6xHisExpressionBacterialMutationS542R and K607RAvailable SinceDec. 11, 2018AvailabilityAcademic Institutions and Nonprofits only -
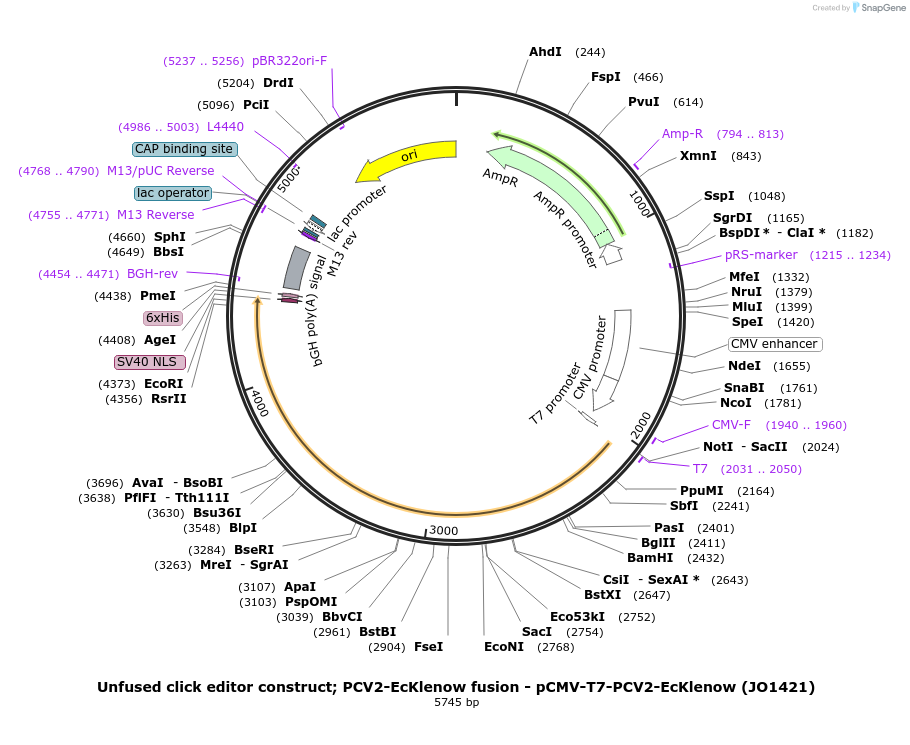
Unfused click editor construct; PCV2-EcKlenow fusion - pCMV-T7-PCV2-EcKlenow (JO1421)
Plasmid#217804PurposeUnfused click editor construct expressing PCV2-EcKlenow, expressed from CMV or T7 promoters.DepositorInsertPCV2-linker-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationEcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
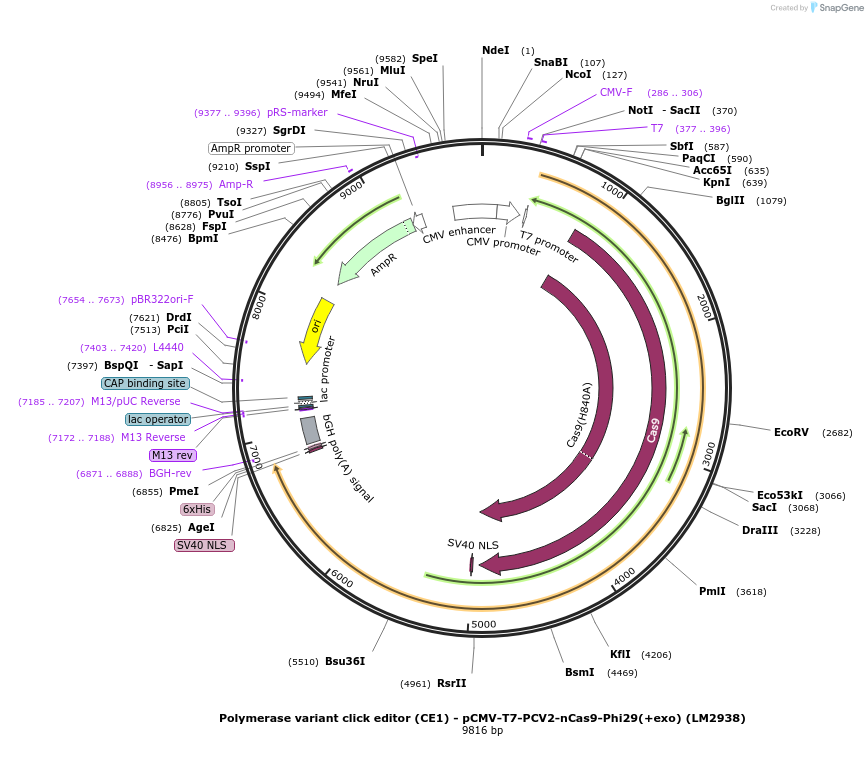
Polymerase variant click editor (CE1) - pCMV-T7-PCV2-nCas9-Phi29(+exo) (LM2938)
Plasmid#208957PurposeA variant CE1 construct with Phi29 DNA polymerase (exo active), expressed from CMV or T7 promoters.DepositorInsertPCV2-XTEN-nSpCas9-BPNLS-Phi29(+exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A)PromoterCMV and T7Available SinceNov. 27, 2023AvailabilityAcademic Institutions and Nonprofits only -
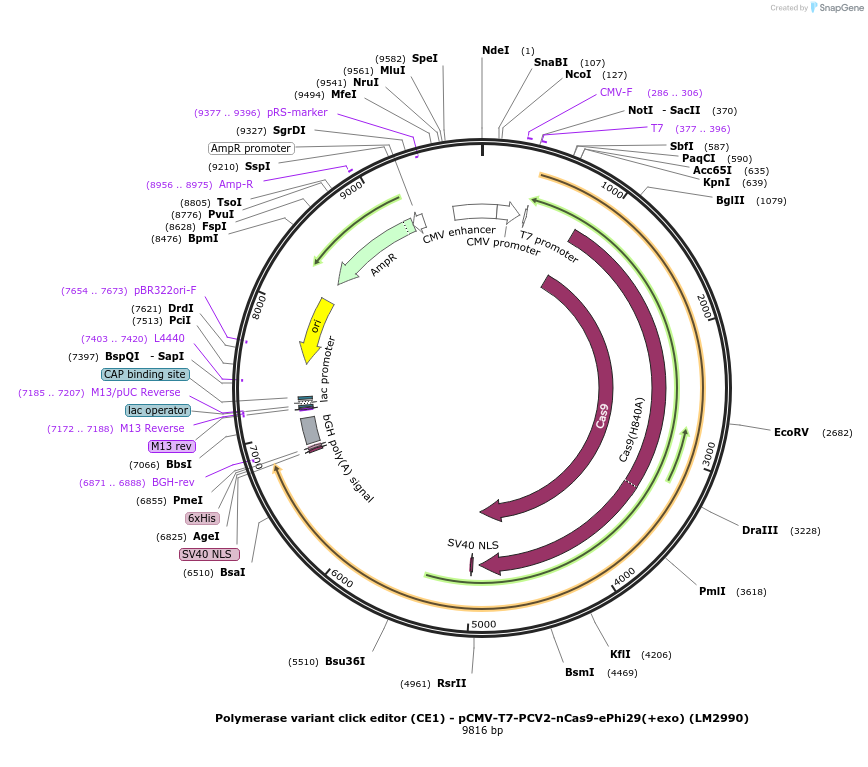
Polymerase variant click editor (CE1) - pCMV-T7-PCV2-nCas9-ePhi29(+exo) (LM2990)
Plasmid#208958PurposeA variant CE1 construct with ePhi29 DNA polymerase (exo active), expressed from CMV or T7 promoters.DepositorInsertPCV2-XTEN-nSpCas9-BPNLS-ePhi29(+exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); ePhi29(M8R/V51A/M97T/G197D/E221K/…PromoterCMV and T7Available SinceNov. 27, 2023AvailabilityAcademic Institutions and Nonprofits only -
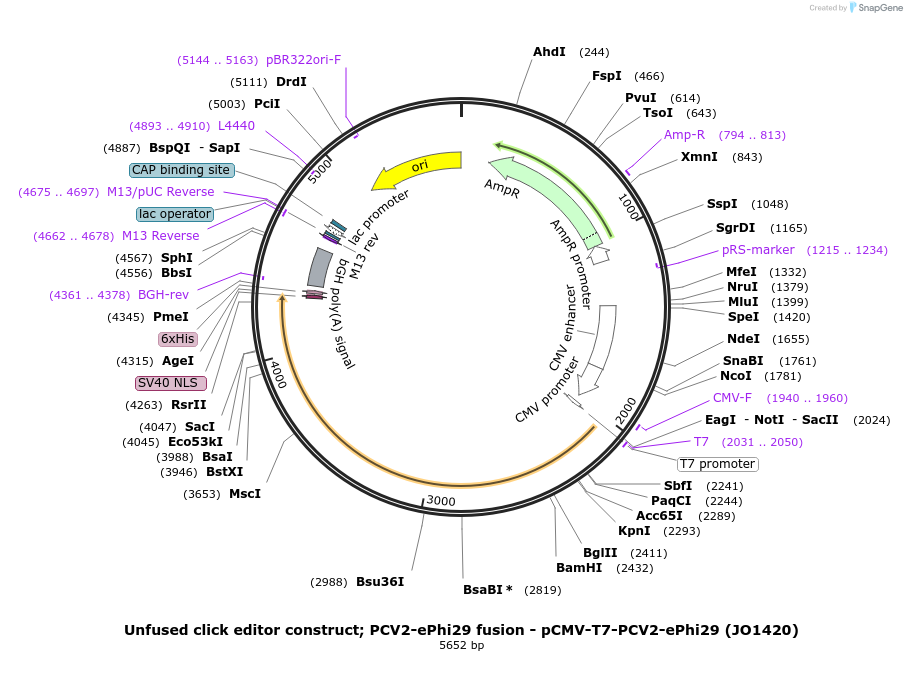
Unfused click editor construct; PCV2-ePhi29 fusion - pCMV-T7-PCV2-ePhi29 (JO1420)
Plasmid#217805PurposeUnfused click editor construct expressing PCV2-ePhi29(D169A), expressed from CMV or T7 promoters.DepositorInsertPCV2-linker-ePhi29-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationPhi29(D169A/M8R/V51A/M97T/G197D/E221K/Q497P/K512E…PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
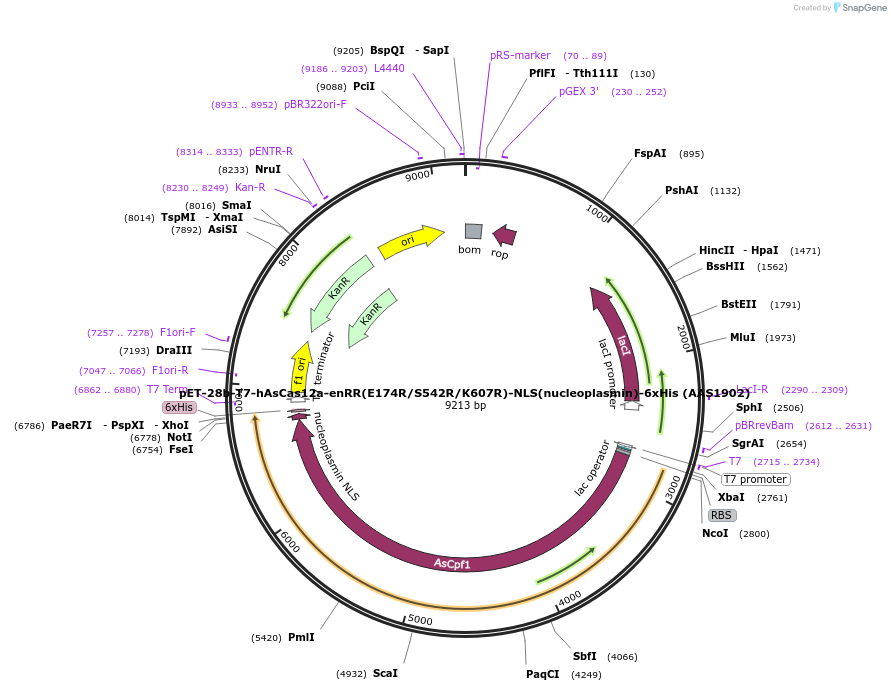
pET-28b-T7-hAsCas12a-enRR(E174R/S542R/K607R)-NLS(nucleoplasmin)-6xHis (AAS1902)
Plasmid#114077PurposeT7 promoter bacterial expression plasmid for human codon optimized AsCas12a-enRR(E174R/S542R/K607R) with C-terminal NLS and His-tagDepositorInserthuman codon optimized AsCas12a-enRR (E174R/S542R/K607R) with C-terminal NLS and His-tag
TagsNLS(nucleoplasmin) and 6xHisExpressionBacterialMutationE174R/S542R/K607RAvailable SinceDec. 11, 2018AvailabilityAcademic Institutions and Nonprofits only -
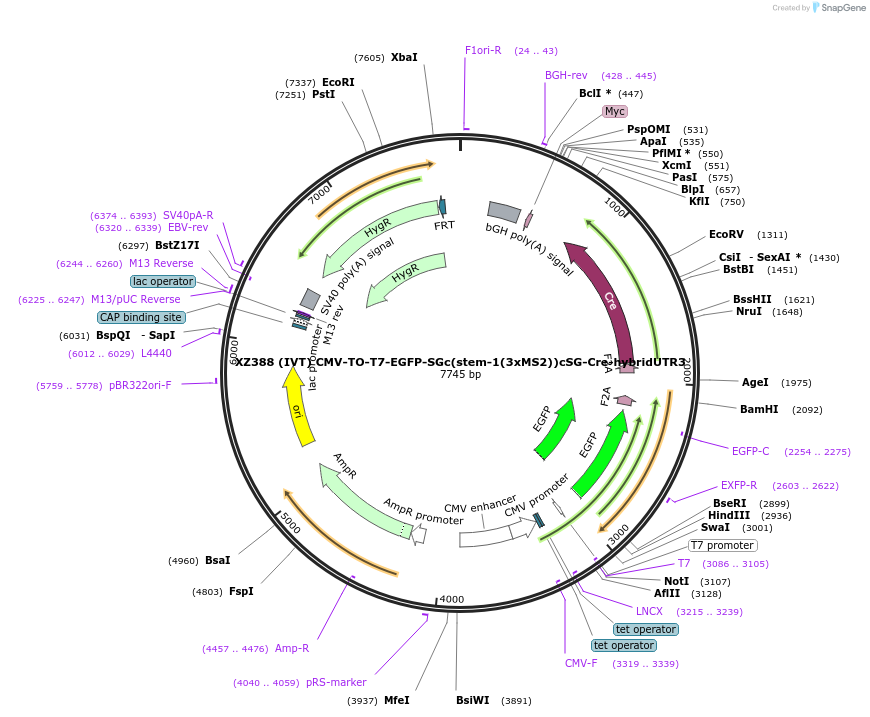
XZ388 (IVT) CMV-TO-T7-EGFP-SGc(stem-1(3xMS2))cSG-Cre-hybridUTR3
Plasmid#233424PurposeIVT template for LIDAR reporter mRNA; EGFP as marker, Cre recombinase as outputDepositorInsertT7-EGFP-SGc(stem-1(3xMS2))cSG-Cre-hybridUTR3
ExpressionMammalianMutationWTPromoterCMVAvailable SinceApril 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
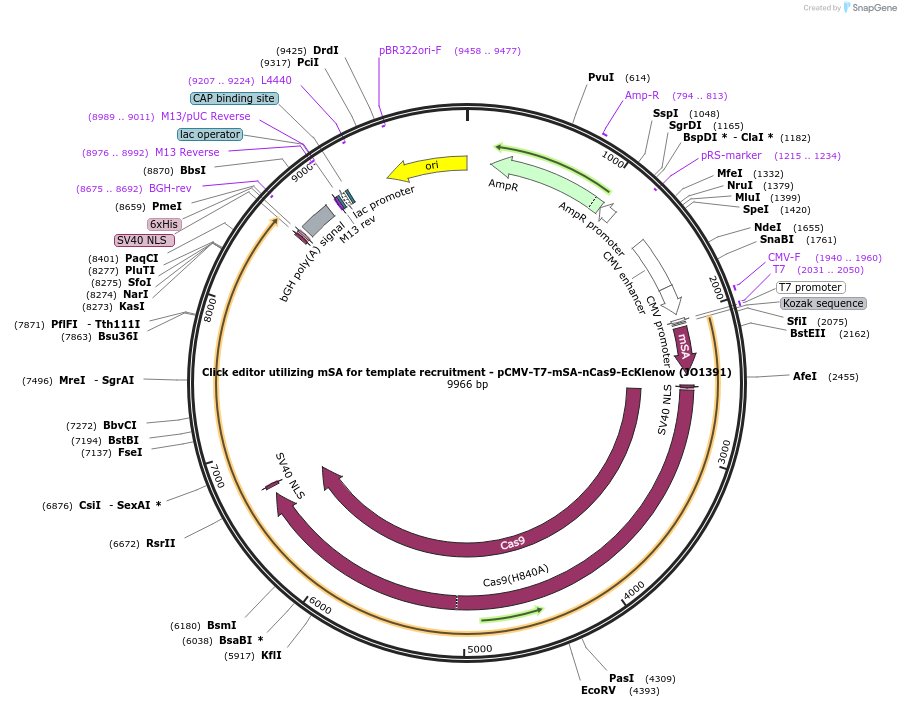
Click editor utilizing mSA for template recruitment - pCMV-T7-mSA-nCas9-EcKlenow (JO1391)
Plasmid#217802PurposeVariant CE1 construct with mSA for template recruitment, expressed from CMV or T7 promoters.DepositorInsertmSA-linker-SV40NLS-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
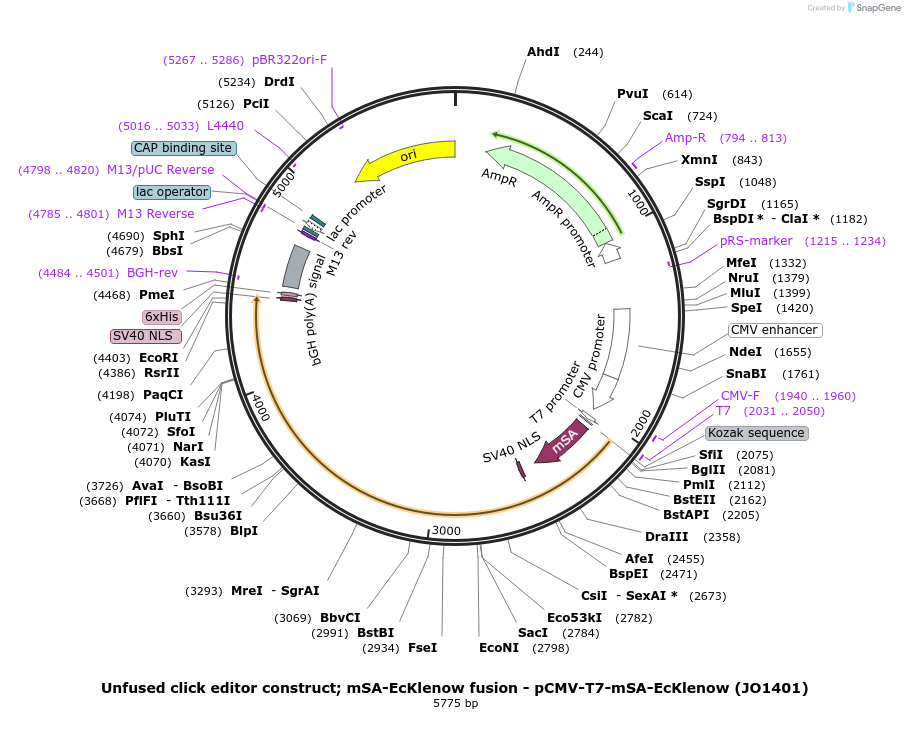
Unfused click editor construct; mSA-EcKlenow fusion - pCMV-T7-mSA-EcKlenow (JO1401)
Plasmid#217806PurposeUnfused click editor construct expressing mSA-EcKlenow, expressed from CMV or T7 promoters.DepositorInsertmSA-linker-SV40NLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationEcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
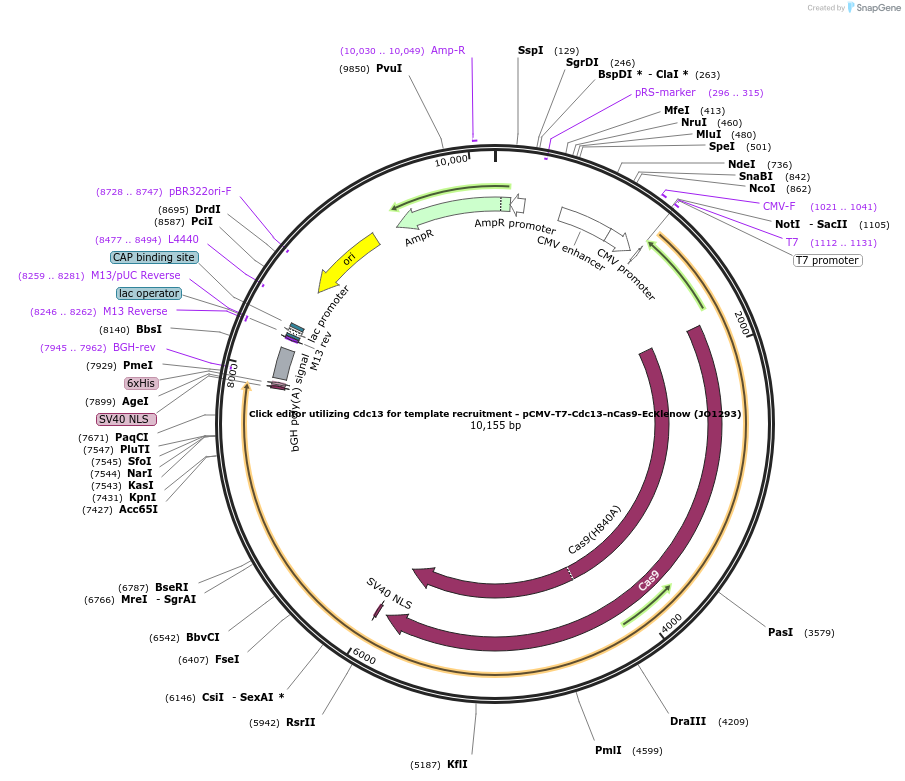
Click editor utilizing Cdc13 for template recruitment - pCMV-T7-Cdc13-nCas9-EcKlenow (JO1293)
Plasmid#217796PurposeVariant CE1 construct with Cdc13 telomere binding protein for template recruitment, expressed from CMV or T7 promoters.DepositorInsertCdc13-XTEN-nSpCas9-BPNLS-EcKlenow(-exo)-BPNLS
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLSExpressionMammalianMutationnSpCas9(H840A); EcKlenow(-exo;D355A/E357A)PromoterCMV and T7Available SinceSept. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
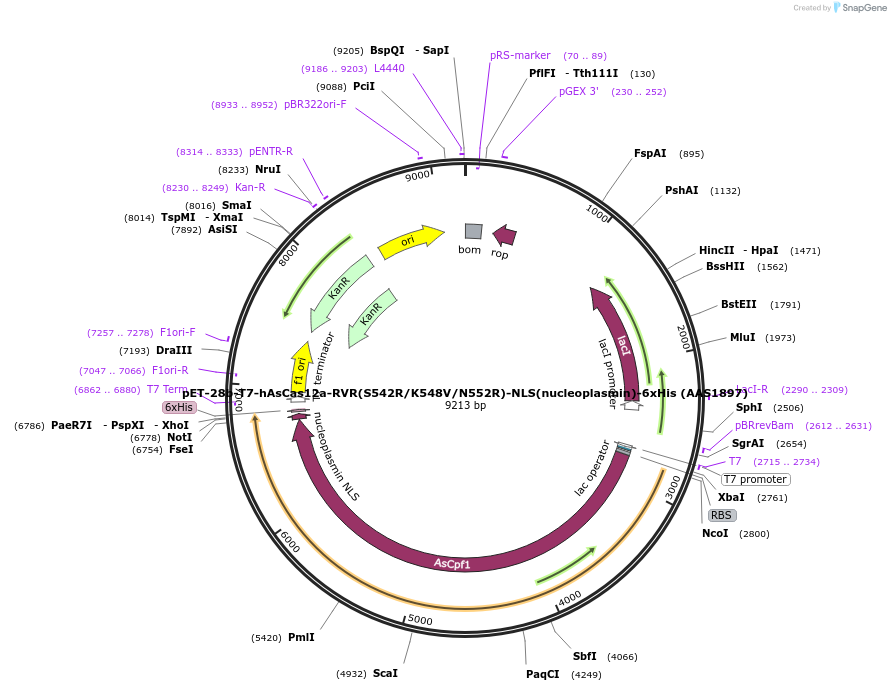
pET-28b-T7-hAsCas12a-RVR(S542R/K548V/N552R)-NLS(nucleoplasmin)-6xHis (AAS1897)
Plasmid#114074PurposeT7 promoter bacterial expression plasmid for human codon optimized AsCas12a-RVR(S542R/K548V/N552R) with C-terminal NLS and His-tagDepositorInserthuman codon optimized AsCas12a-RVR (S542R/K548V/N552R) with C-terminal NLS and His-tag
TagsNLS(nucleoplasmin) and 6xHisExpressionBacterialMutationS542R, K548V and N552RAvailable SinceDec. 11, 2018AvailabilityAcademic Institutions and Nonprofits only