We narrowed to 1,445 results for: pBAD
-
Plasmid#195665PurposeComplementation of MlaC(V171R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(V171R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
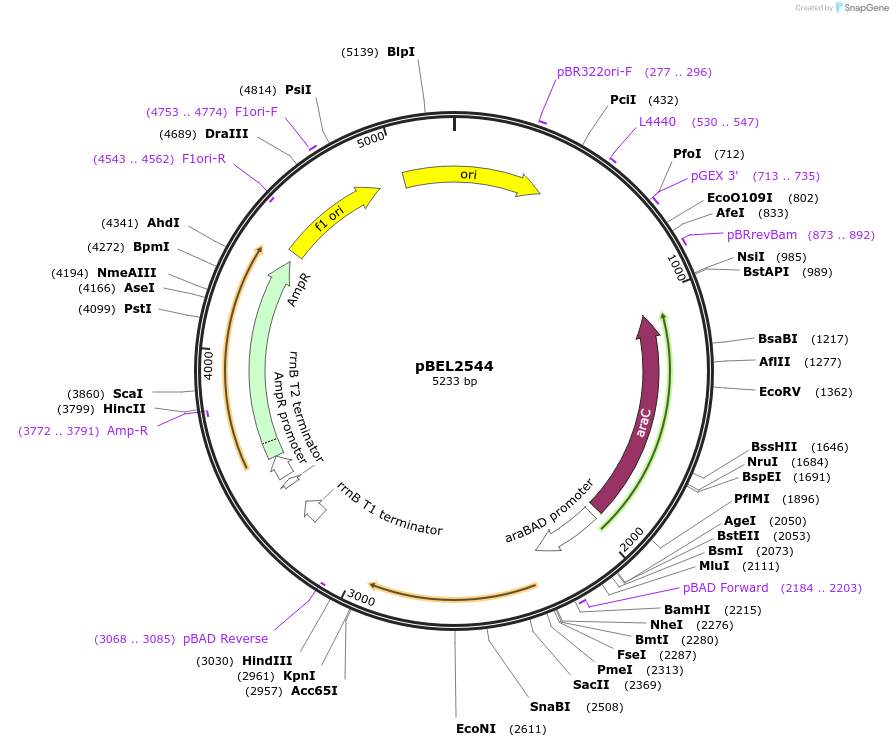
pBEL2545
Plasmid#195666PurposeComplementation of MlaC(Y72A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y72A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
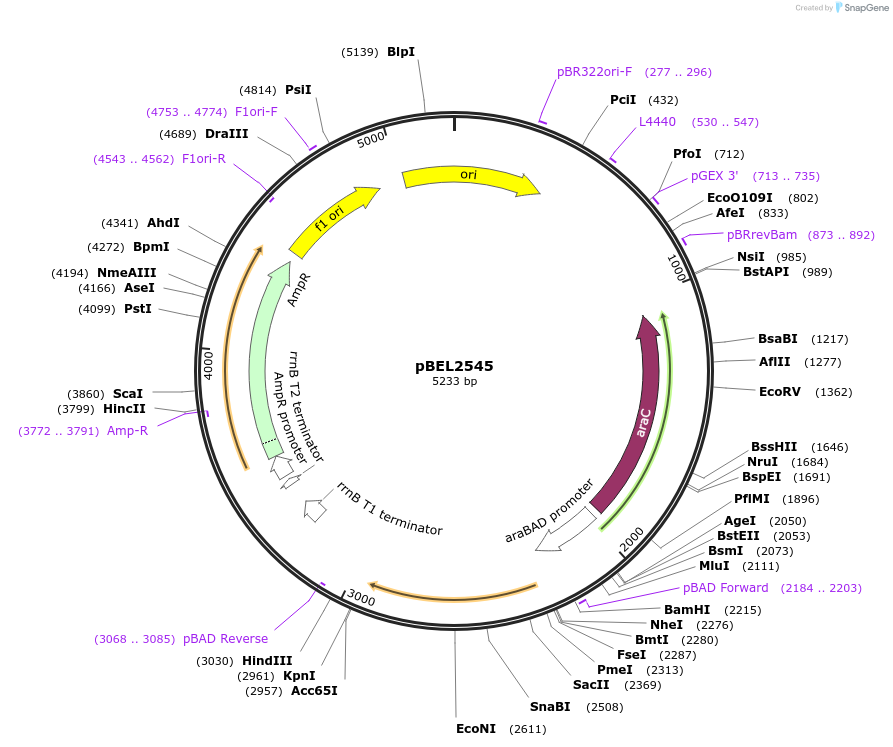
pBEL2546
Plasmid#195667PurposeComplementation of MlaC(Y82K)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y82K)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
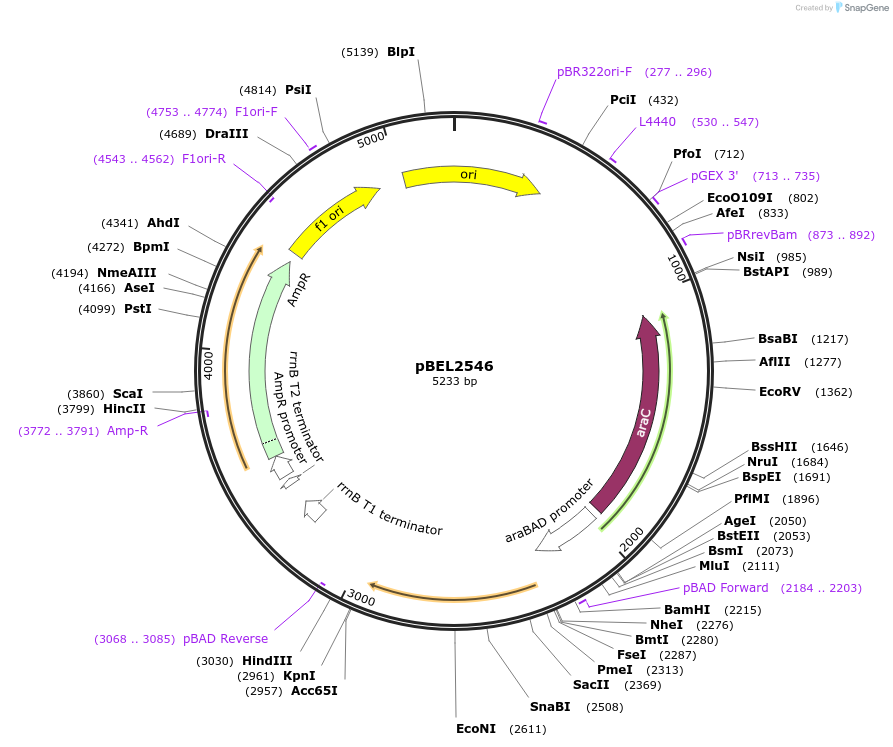
pBEL2547
Plasmid#195668PurposeComplementation of MlaC(Y105A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y105A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
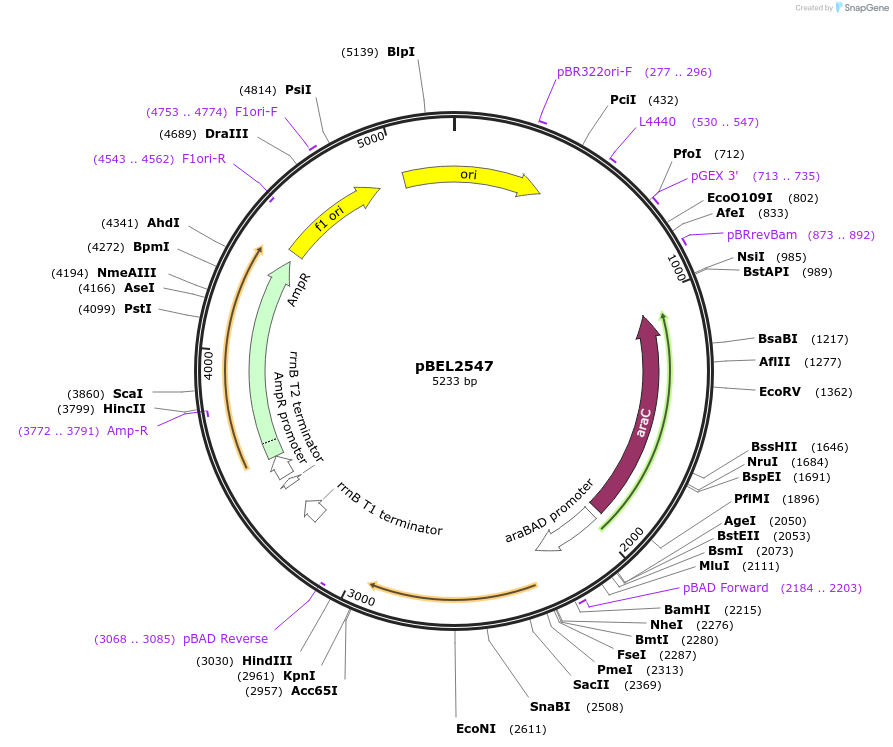
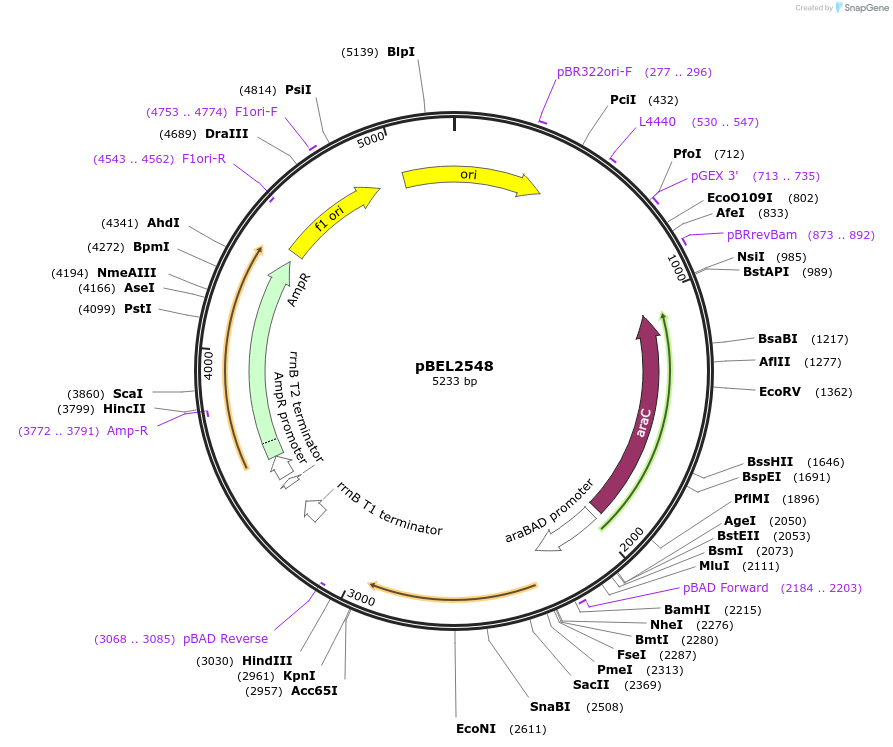
pBEL2548
Plasmid#195669PurposeComplementation of MlaC(Y164R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Y164R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
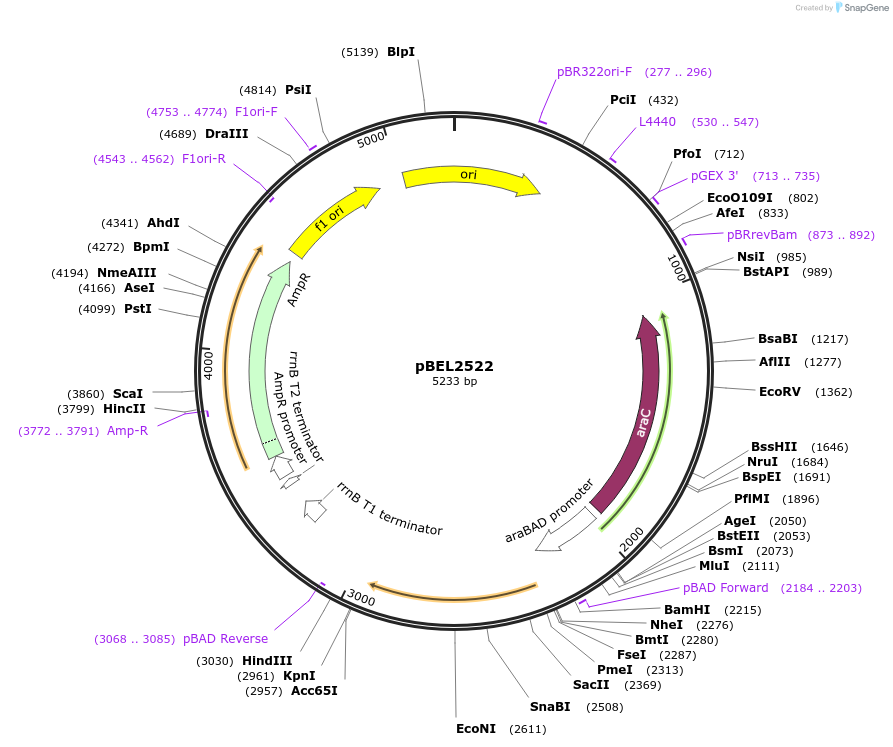
pBEL2522
Plasmid#195644PurposeComplementation of MlaC(A108R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A108R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
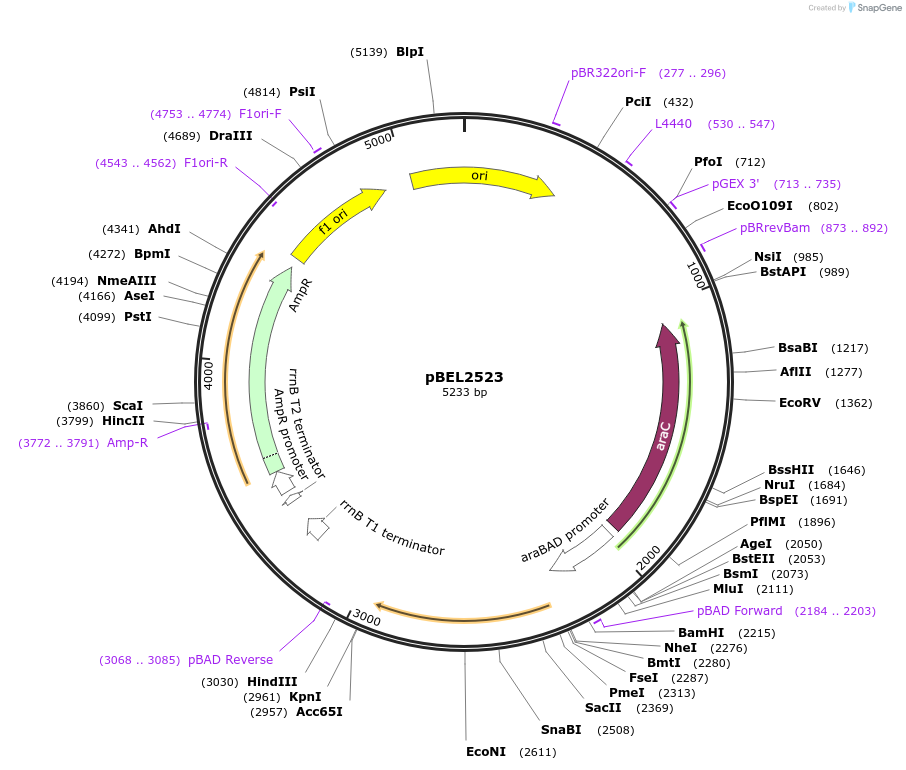
pBEL2523
Plasmid#195645PurposeComplementation of MlaC(A163R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A163R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
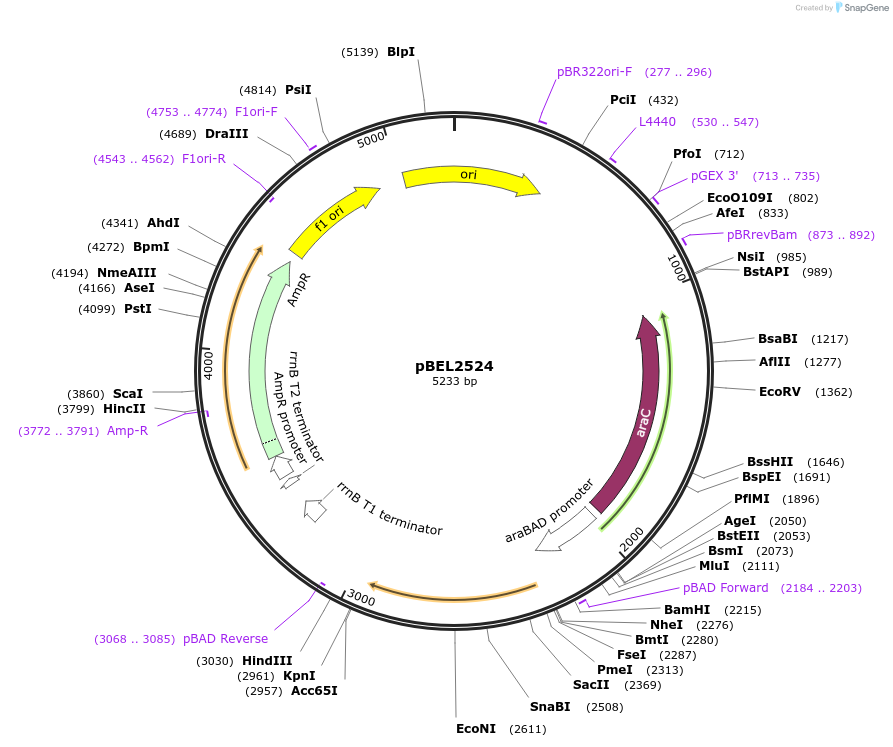
pBEL2524
Plasmid#195646PurposeComplementation of MlaC(A168R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A168R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
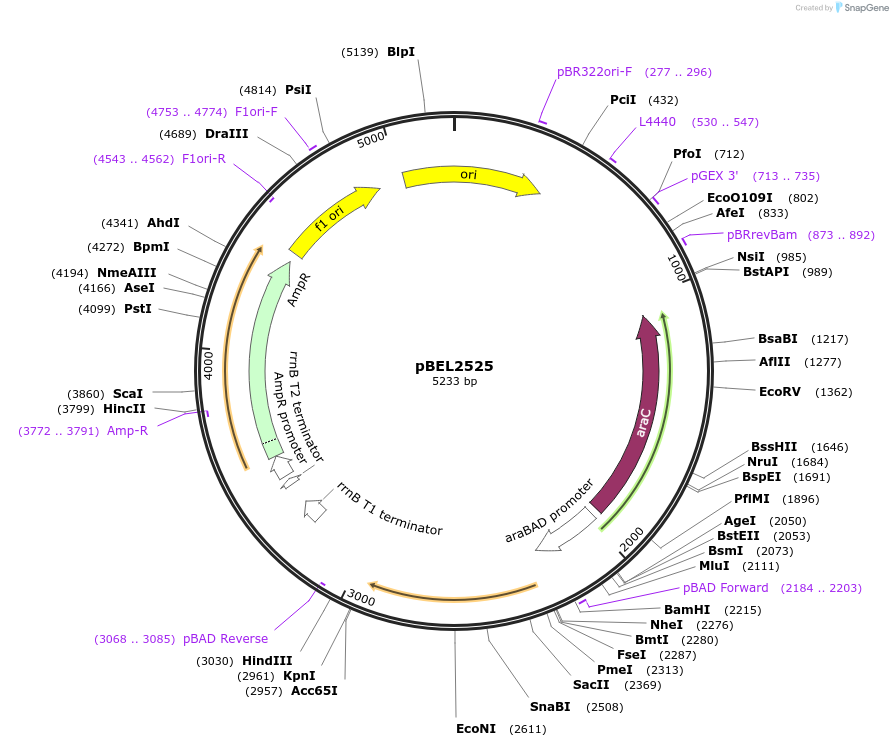
pBEL2525
Plasmid#195647PurposeComplementation of MlaC(D165I)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(D165I)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
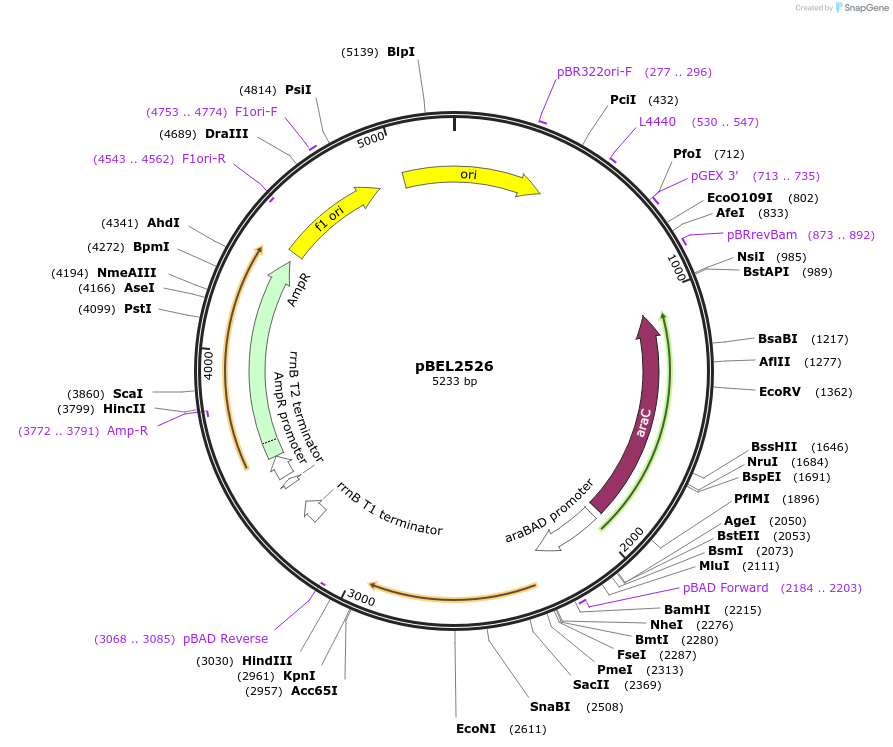
pBEL2526
Plasmid#195648PurposeComplementation of MlaC(E169R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(E169R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2527
Plasmid#195649PurposeComplementation of MlaC(G170K)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(G170K)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
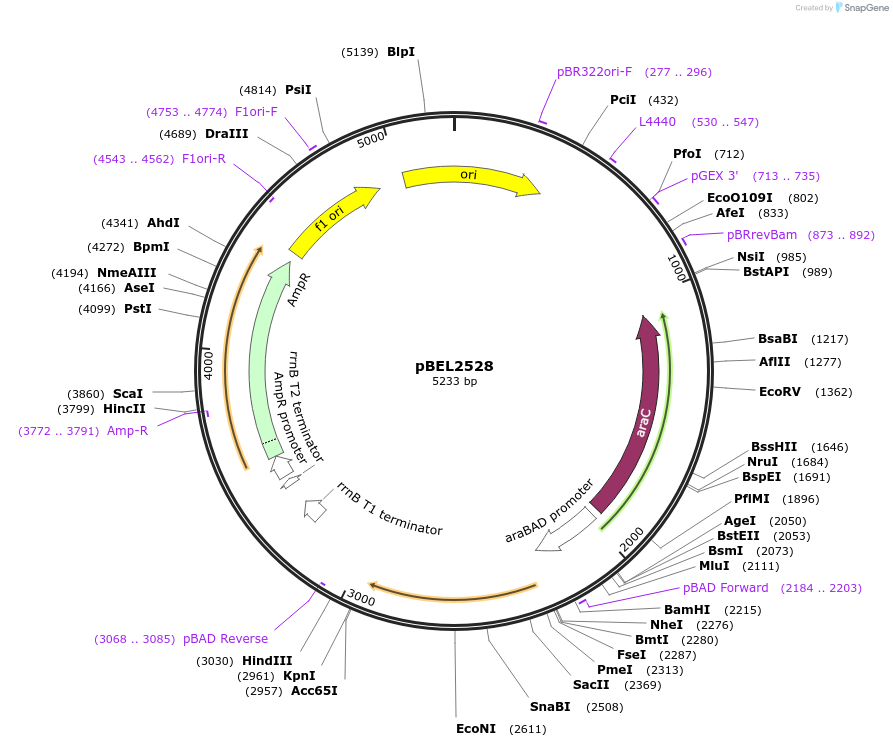
pBEL2528
Plasmid#195650PurposeComplementation of MlaC(I130D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(I130D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
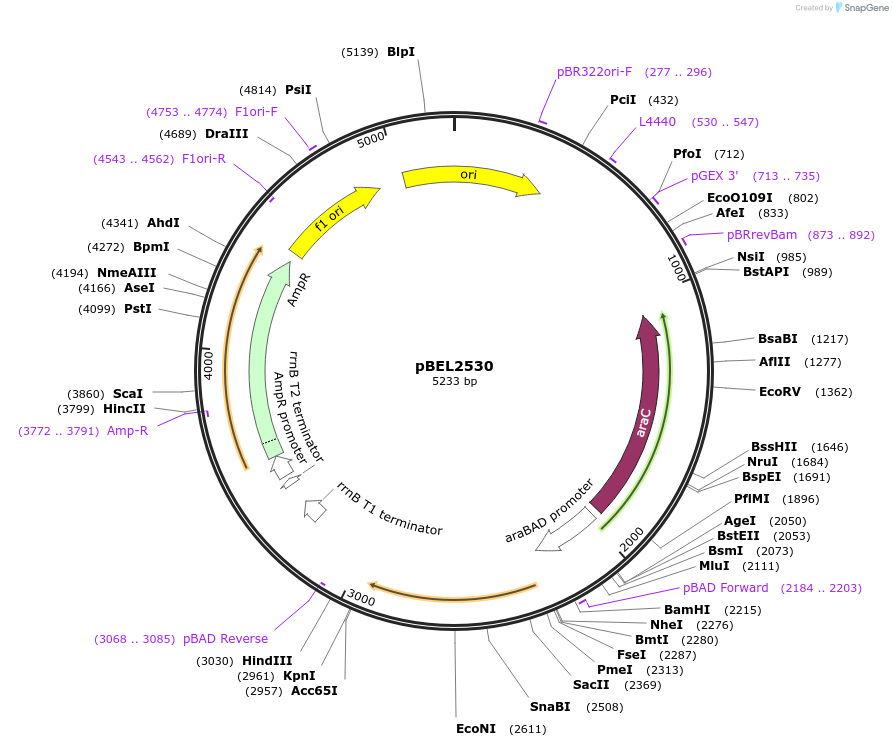
pBEL2530
Plasmid#195651PurposeComplementation of MlaC(I137D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(I137D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
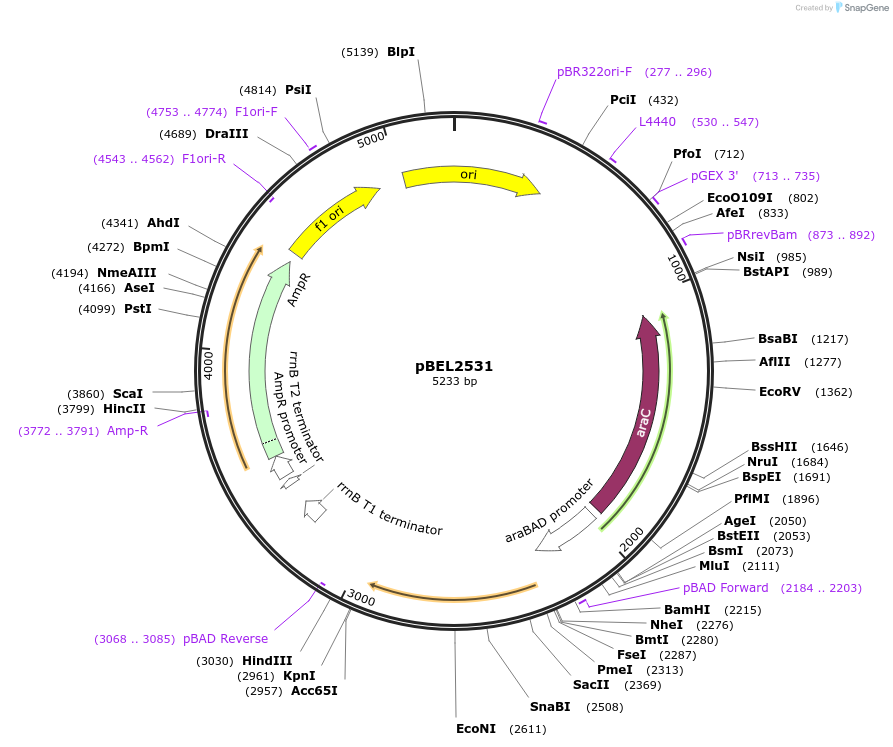
pBEL2531
Plasmid#195652PurposeComplementation of MlaC(L76D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(L76D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
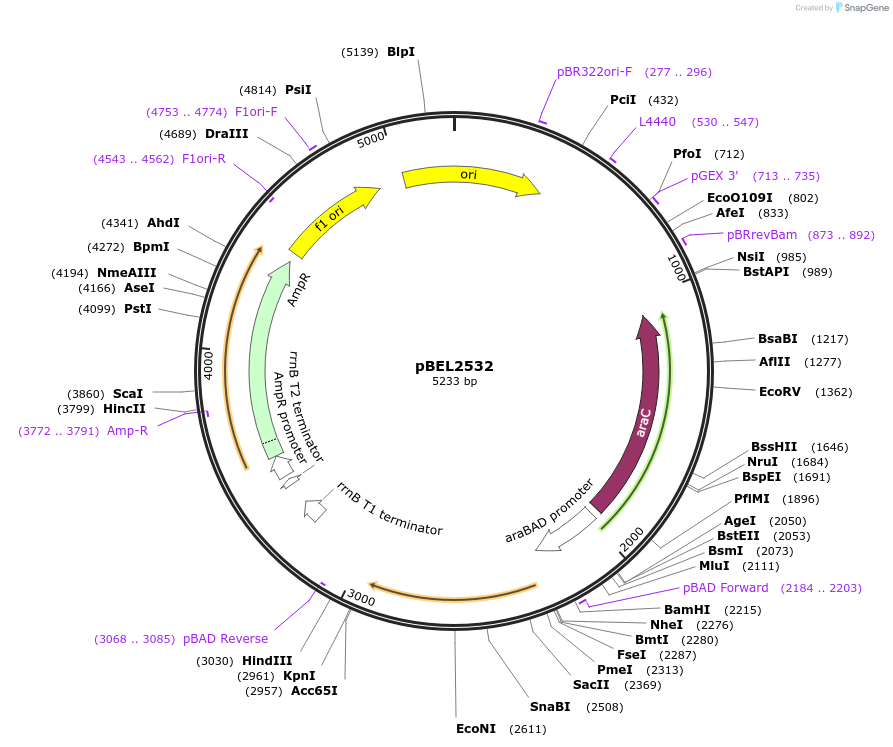
pBEL2532
Plasmid#195653PurposeComplementation of MlaC(N155A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(N155A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
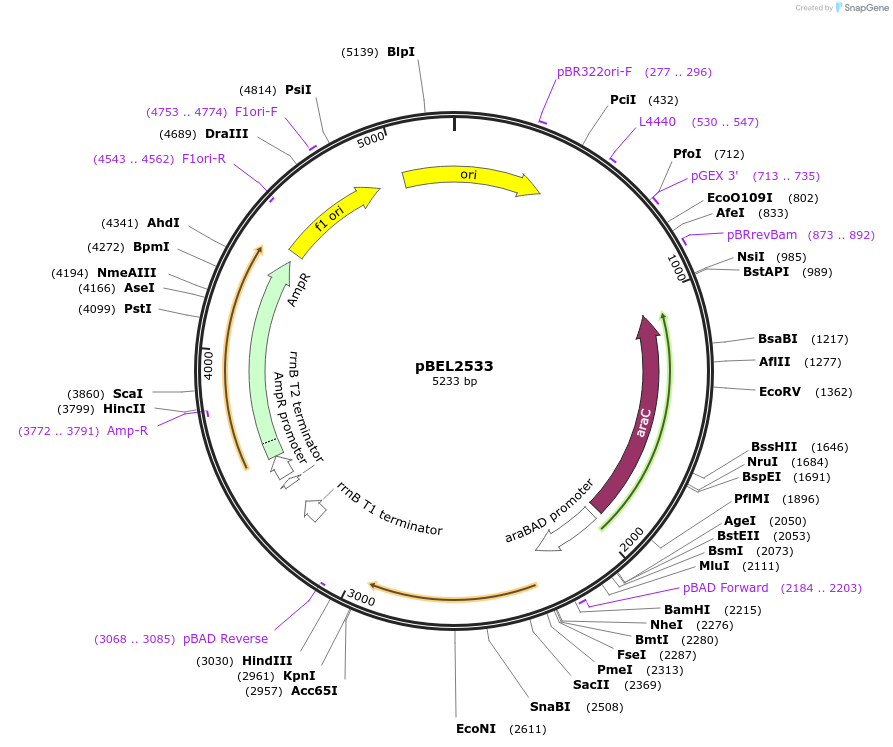
pBEL2533
Plasmid#195654PurposeComplementation of MlaC(N179E)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(N179E)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
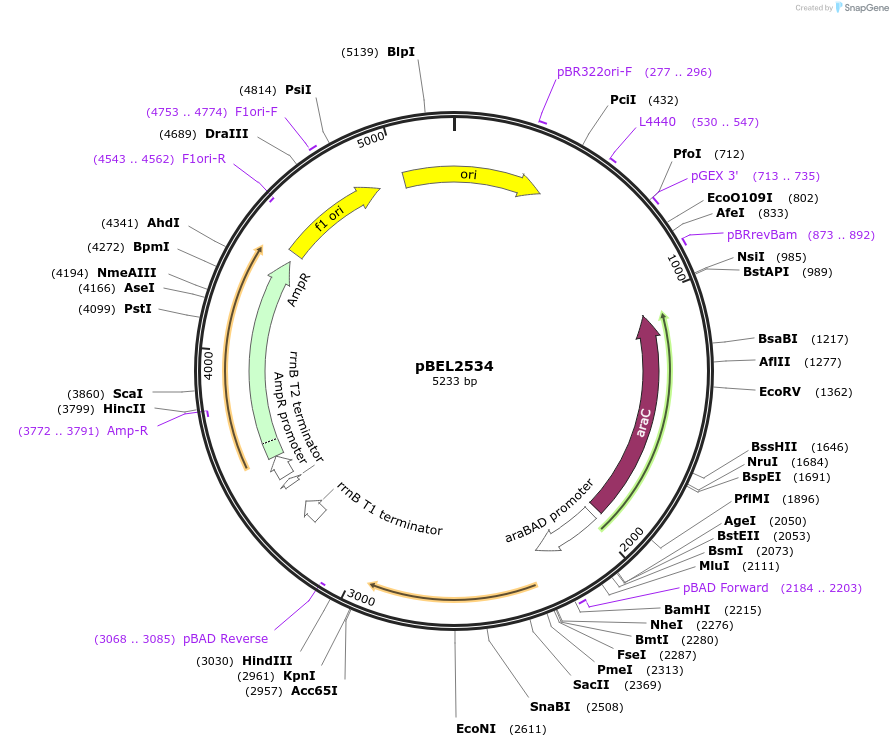
pBEL2534
Plasmid#195655PurposeComplementation of MlaC(Q115A)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(Q115A)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
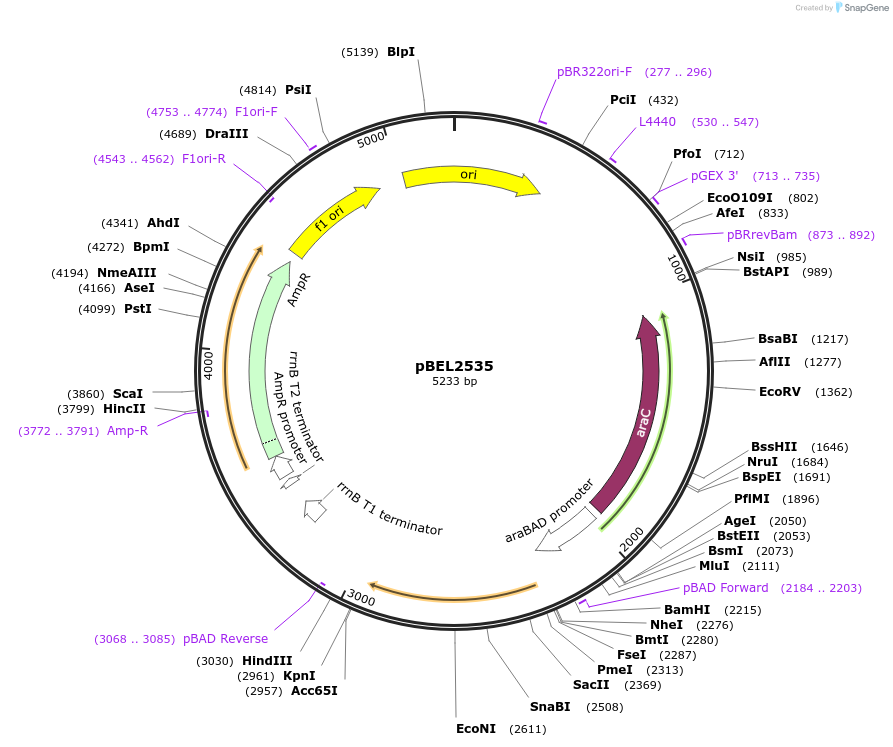
pBEL2535
Plasmid#195656PurposeComplementation of MlaC(R134E)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R134E)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
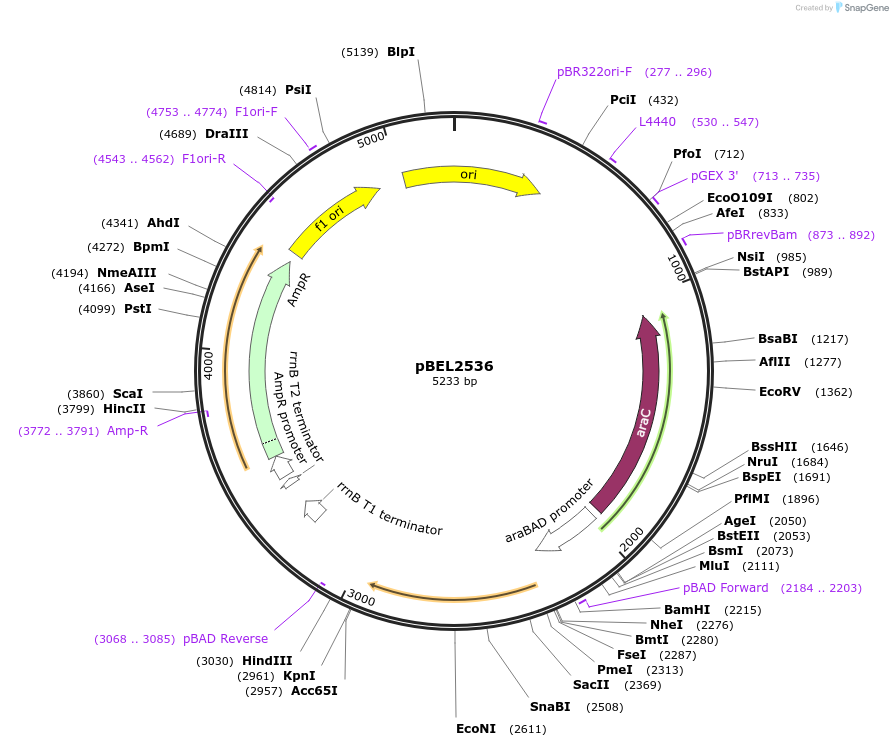
pBEL2536
Plasmid#195657PurposeComplementation of MlaC(R147D)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(R147D)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
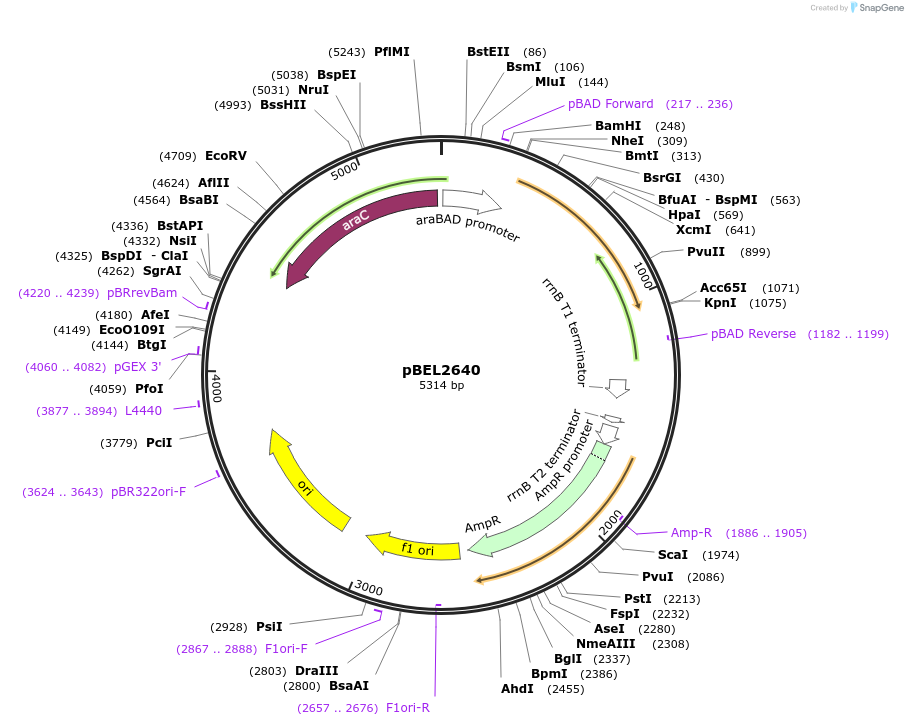
pBEL2640
Plasmid#195629PurposeComplementation of MlaA(L245N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(L245N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
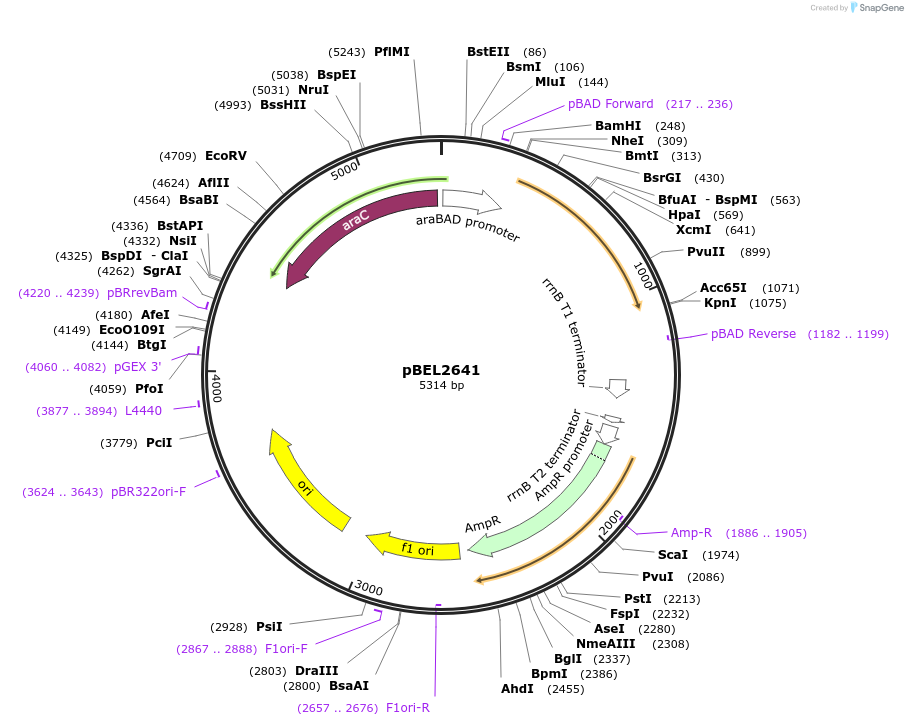
pBEL2641
Plasmid#195630PurposeComplementation of MlaA(I248N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(I248N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
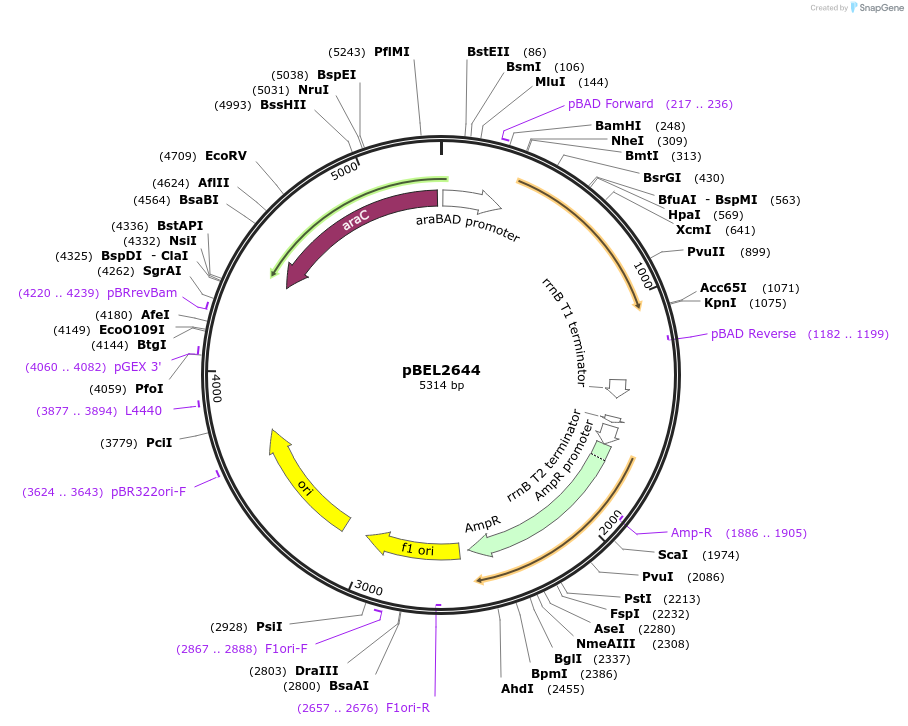
pBEL2644
Plasmid#195631PurposeComplementation of MlaA(D198N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(D198N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
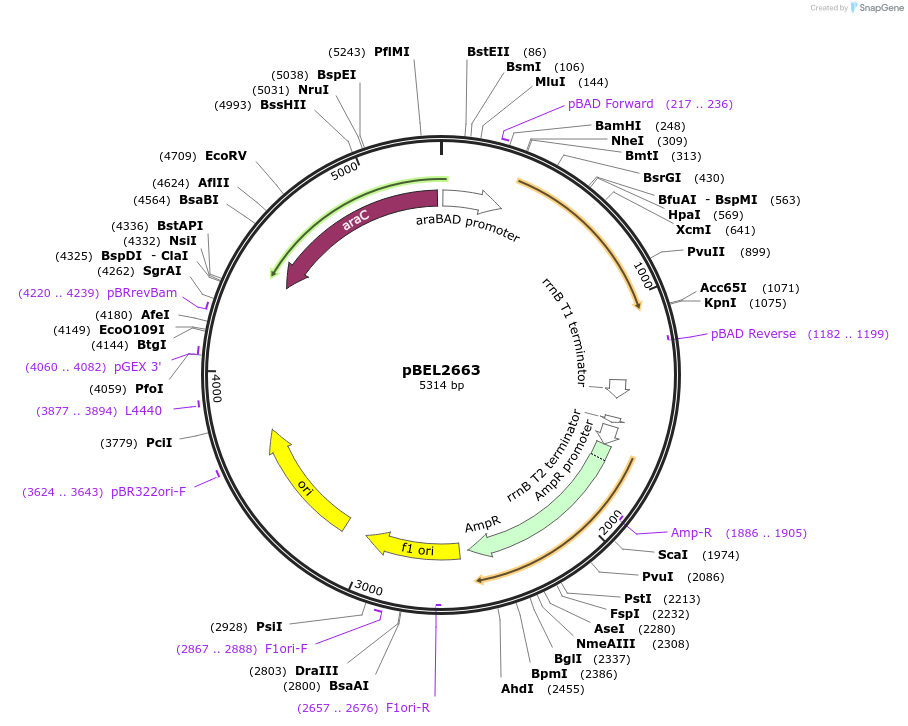
pBEL2663
Plasmid#195632PurposeComplementation of MlaA(F223N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(F223N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
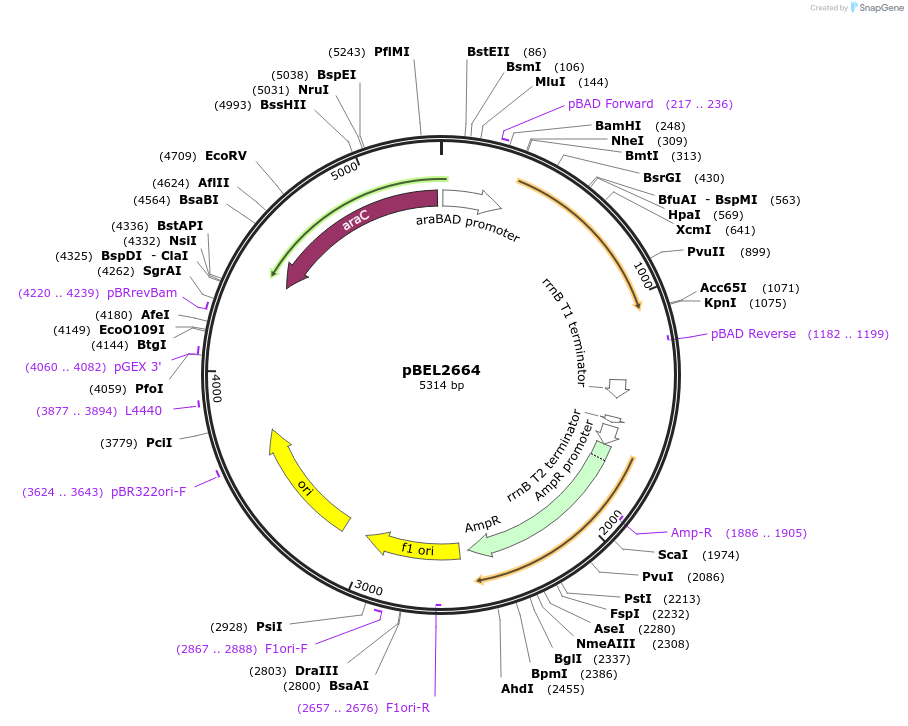
pBEL2664
Plasmid#195633PurposeComplementation of MlaA(L230N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(L230N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2665
Plasmid#195634PurposeComplementation of MlaA(delta244-251)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(delta244-251)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
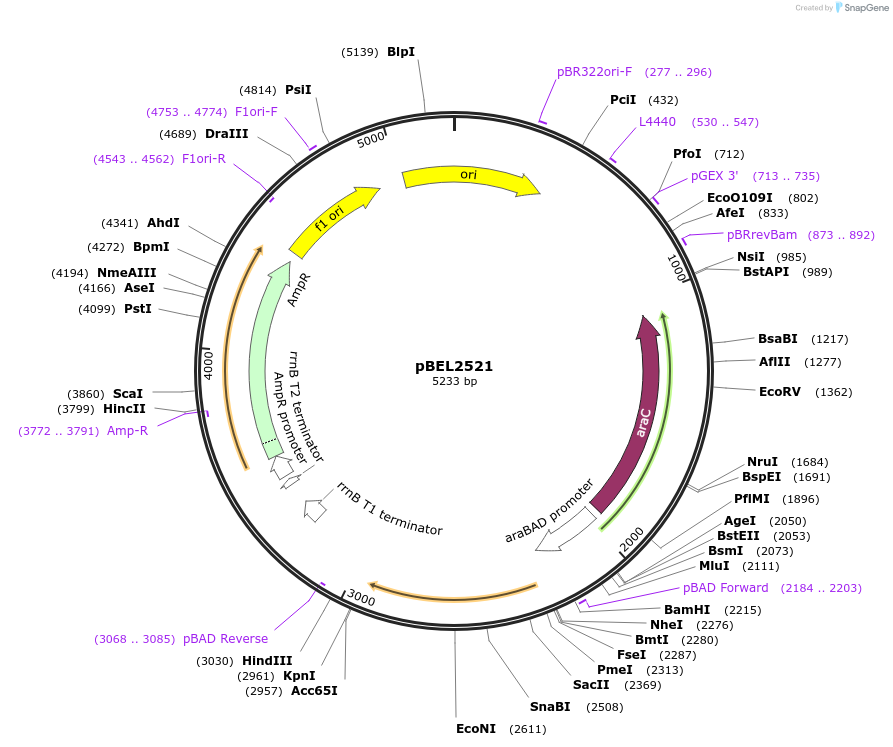
pBEL2521
Plasmid#195643PurposeComplementation of MlaC(A34R)DepositorInsertMlaC
ExpressionBacterialMutationMlaC(A34R)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
pBEL2512
Plasmid#195626PurposeComplementation of MlaA(delta238-251)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(delta238-251)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
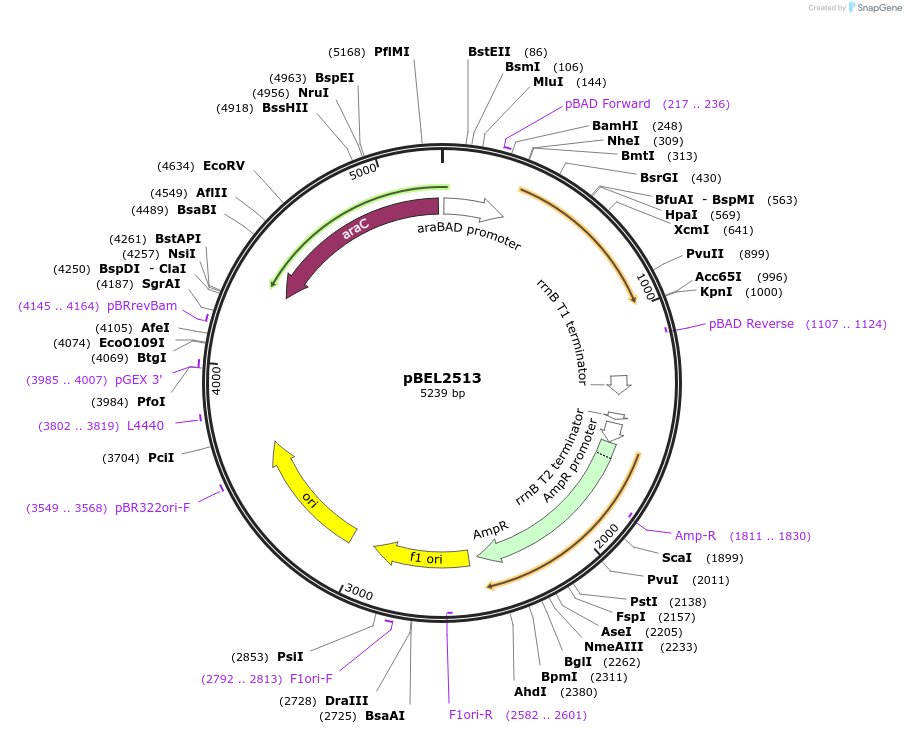
pBEL2513
Plasmid#195627PurposeComplementation of MlaA(delta227-251)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(delta227-251)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
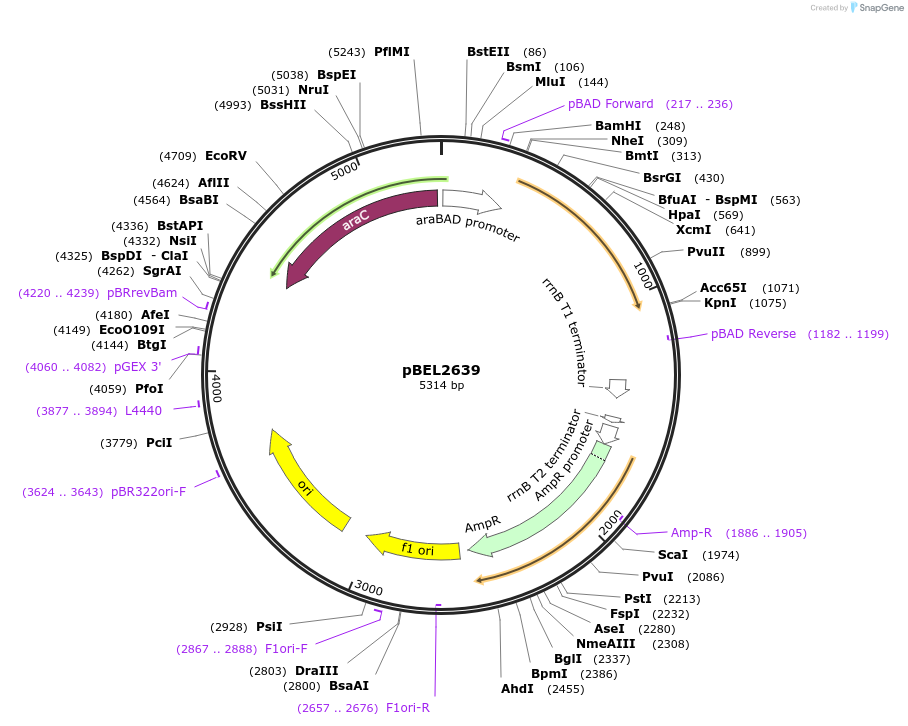
pBEL2639
Plasmid#195628PurposeComplementation of MlaA(I241N)DepositorInsertMlaA
ExpressionBacterialMutationMlaA(I241N)Available SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
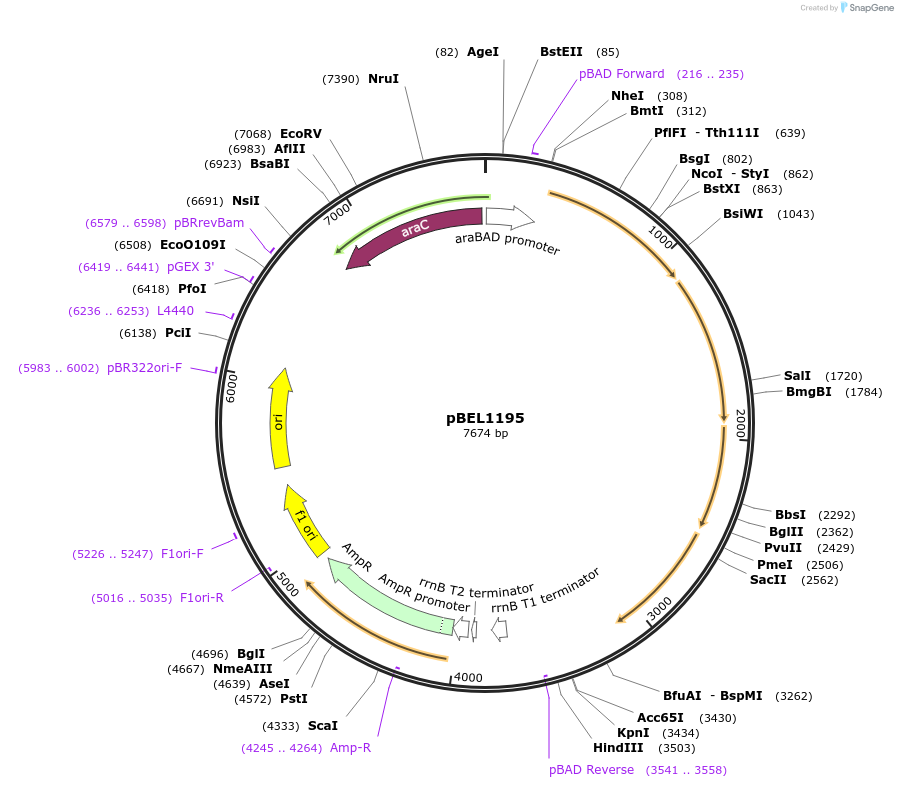
pBEL1195
Plasmid#196027PurposeComplementation of MlaF-MlaE-MlaD-MlaC-MlaBDepositorInsertMlaFEDCB
ExpressionBacterialAvailable SinceMay 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
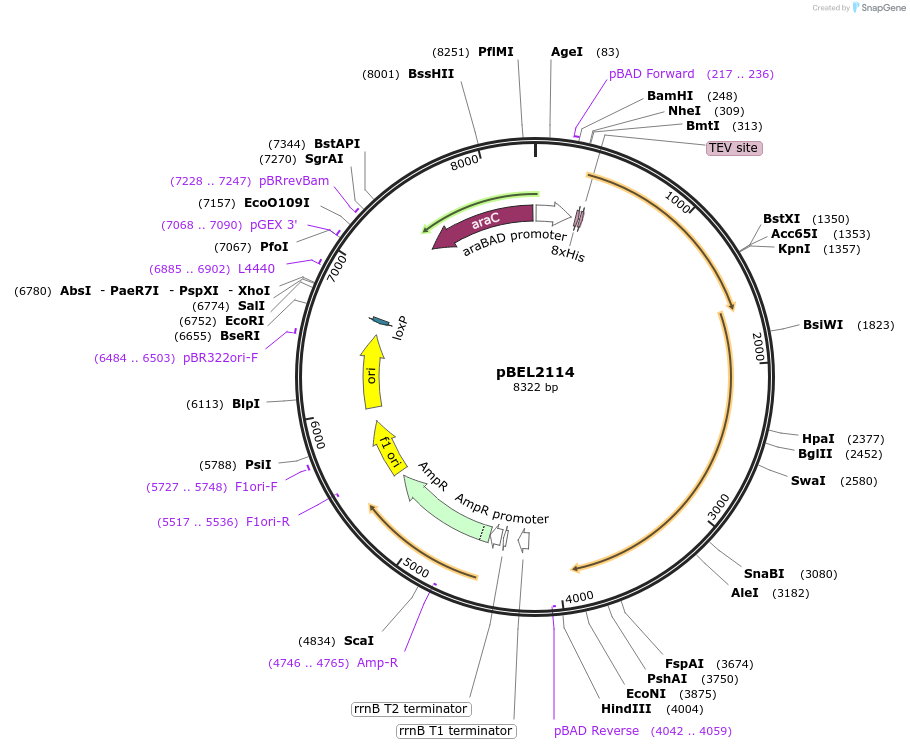
pBEL2114
Plasmid#175961PurposeFor expression of LetA-[ΔRing6 LetB] operon from E. coli. N-terminal his tag on LetA, araBAD promoterDepositorInsertsLetA
LetB
Tags8xHis 2xQH and TEV cleavage site on LetAExpressionBacterialMutationΔ634-746PromoteraraBADAvailable SinceOct. 3, 2022AvailabilityAcademic Institutions and Nonprofits only -
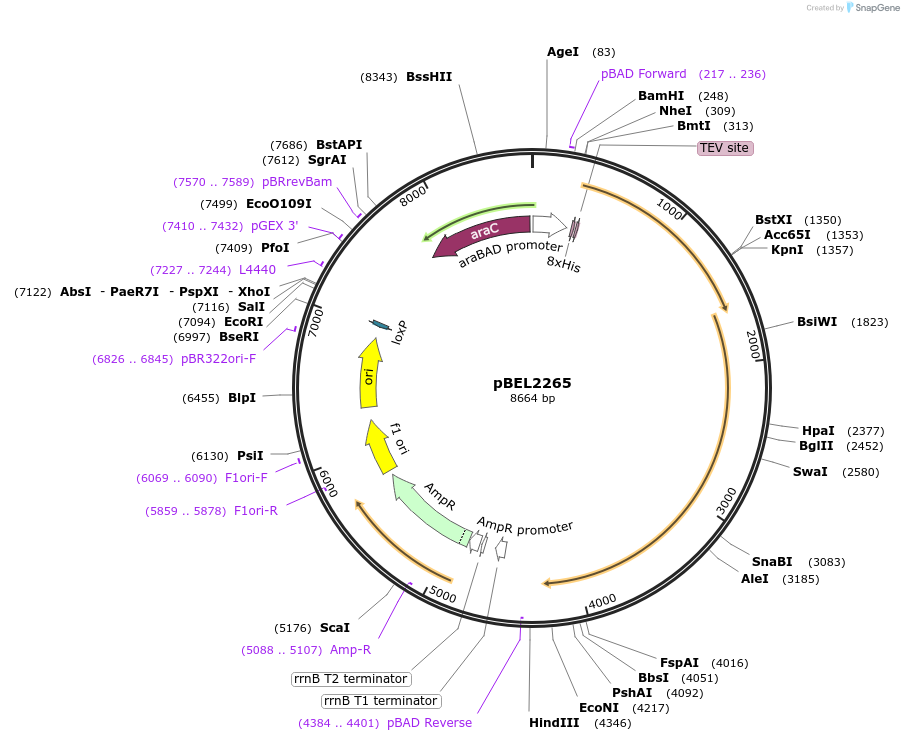
pBEL2265
Plasmid#175962PurposeFor expresson of LetA-[LetB PLL3->6] operon from E. coli. N-terminal his tag on LetA, araBAD promoterDepositorInsertsLetA
LetB
Tags8xHis 2xQH and TEV cleavage site on LetAExpressionBacterialMutationΔ347-361, replace with 702-716PromoteraraBADAvailable SinceOct. 3, 2022AvailabilityAcademic Institutions and Nonprofits only -
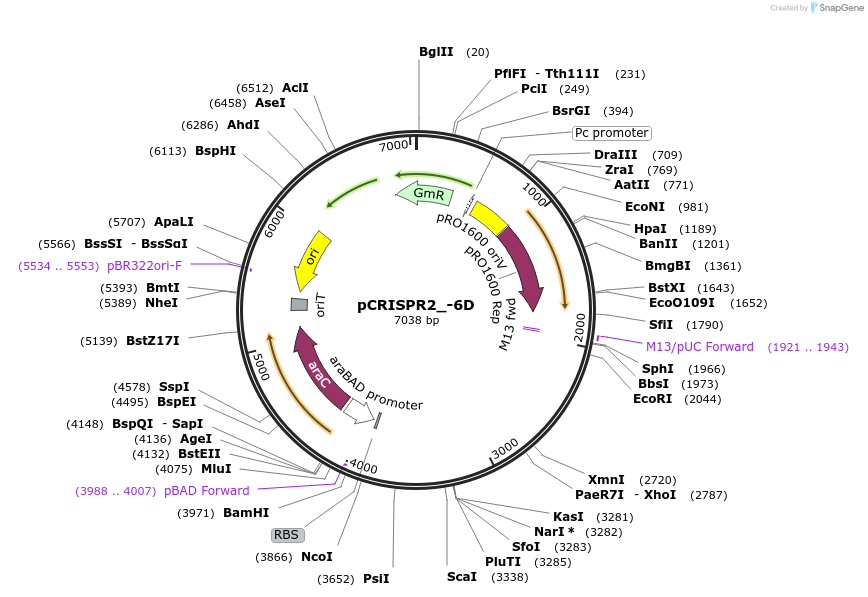
pCRISPR2_-6D
Plasmid#149387PurposeP. aeruginosa PA14 CRISPR2 locus, with 6bp deletion downstream of IHFprox siteDepositorInsertPseudomonas aeruginosa PA14 CRISPR2 locus
ExpressionBacterialMutation6bp deleted downstream of the IHFprox sitePromoterpBADAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
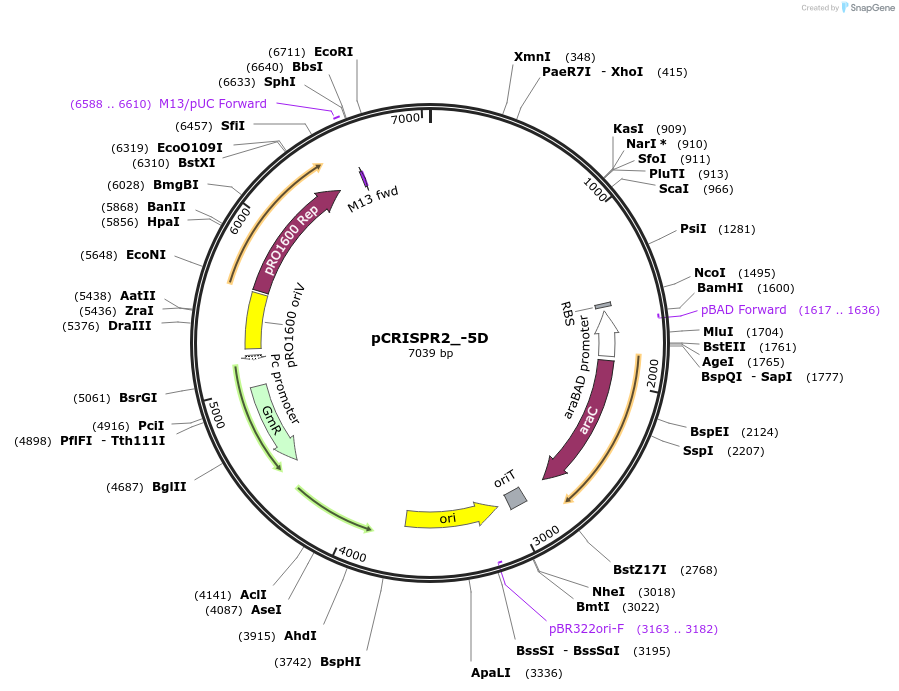
pCRISPR2_-5D
Plasmid#149388PurposeP. aeruginosa PA14 CRISPR2 locus, with 5bp deletion downstream of IHFprox siteDepositorInsertPseudomonas aeruginosa PA14 CRISPR2 locus
ExpressionBacterialMutation5bp deleted downstream of the IHFprox sitePromoterpBADAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
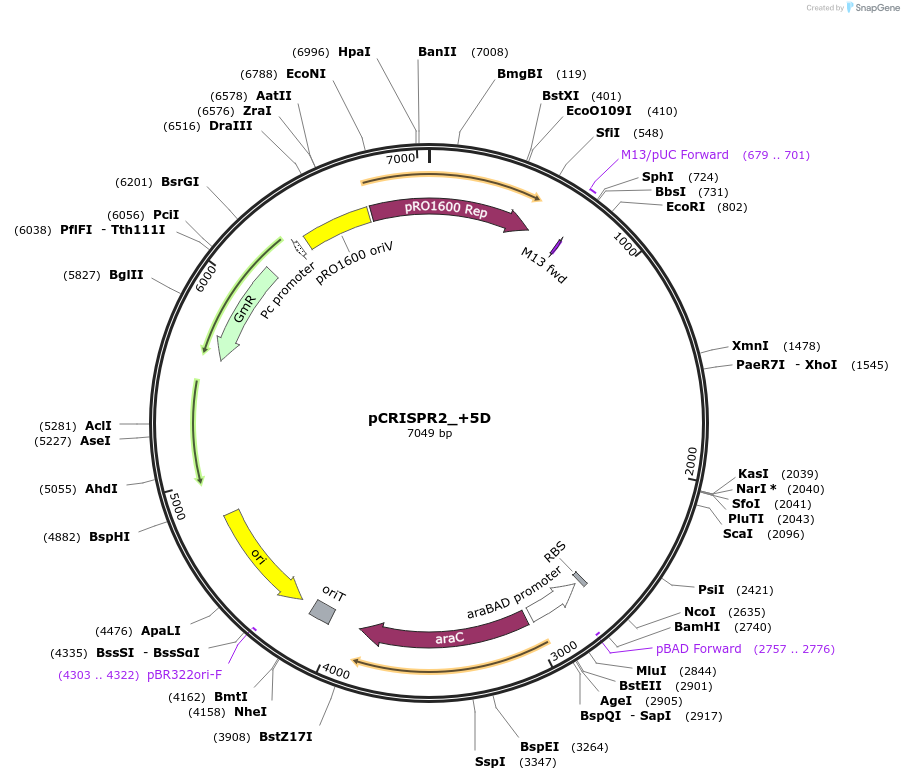
pCRISPR2_+5D
Plasmid#149390PurposeP. aeruginosa PA14 CRISPR2 locus, with 5bp addition downstream of IHFprox siteDepositorInsertPseudomonas aeruginosa PA14 CRISPR2 locus
ExpressionBacterialMutation5bp inserted downstream of the IHFprox sitePromoterpBADAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
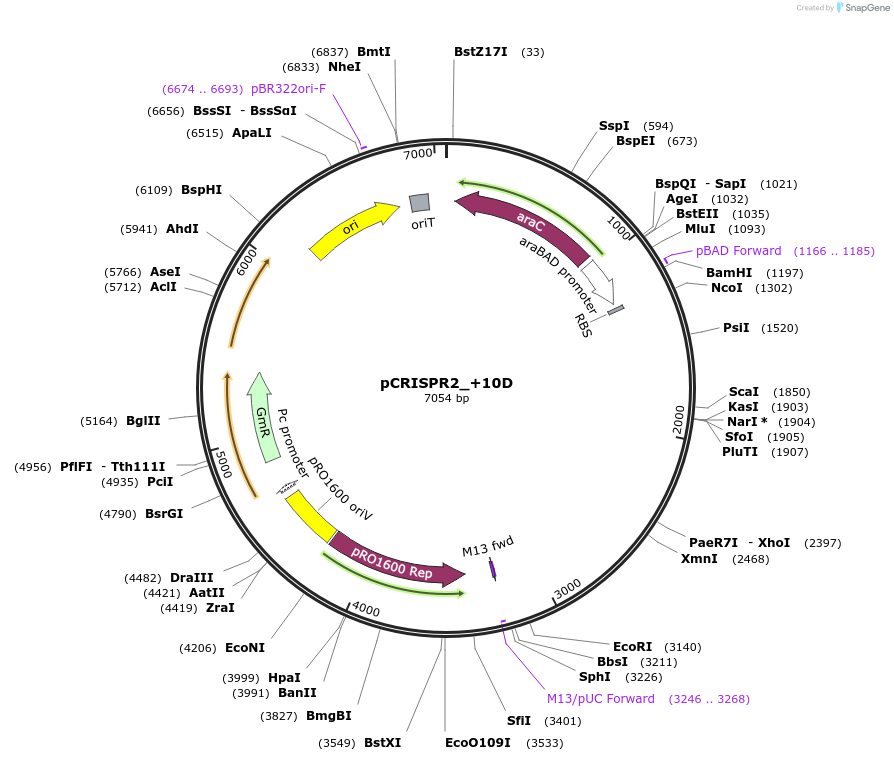
pCRISPR2_+10D
Plasmid#149391PurposeP. aeruginosa PA14 CRISPR2 locus, with 10bp addition downstream of IHFprox siteDepositorInsertPseudomonas aeruginosa PA14 CRISPR2 locus
ExpressionBacterialMutation10bp inserted downstream of the IHFprox sitePromoterpBADAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
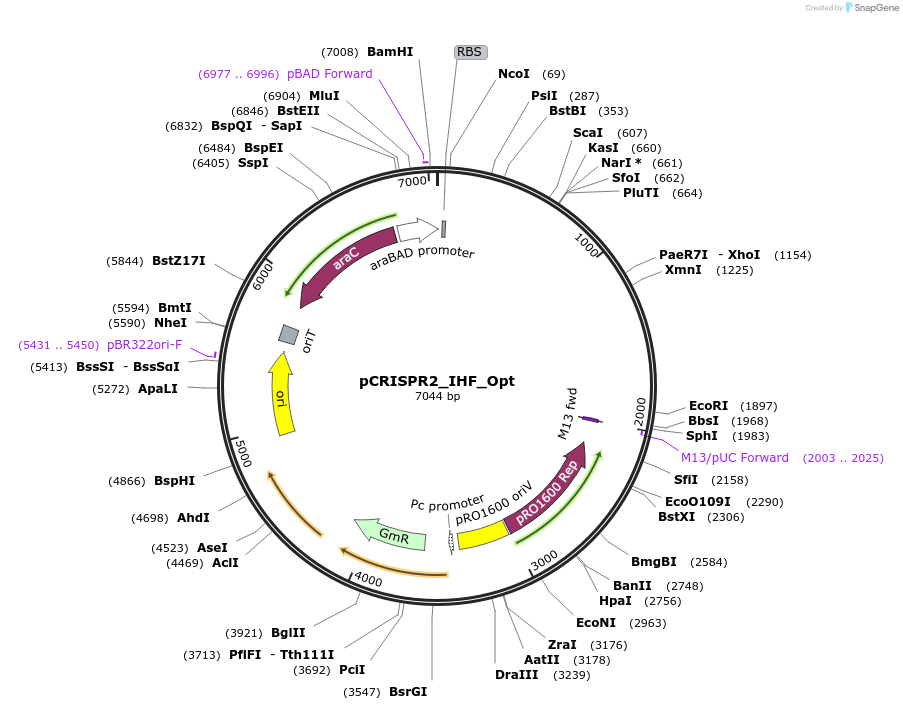
pCRISPR2_IHF_Opt
Plasmid#149395PurposeP. aeruginosa PA14 CRISPR2 locus, with "optimized" IHFprox siteDepositorInsertPseudomonas aeruginosa PA14 CRISPR2 locus
ExpressionBacterialMutationIHFprox site is optimized at key positions needed…PromoterpBADAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
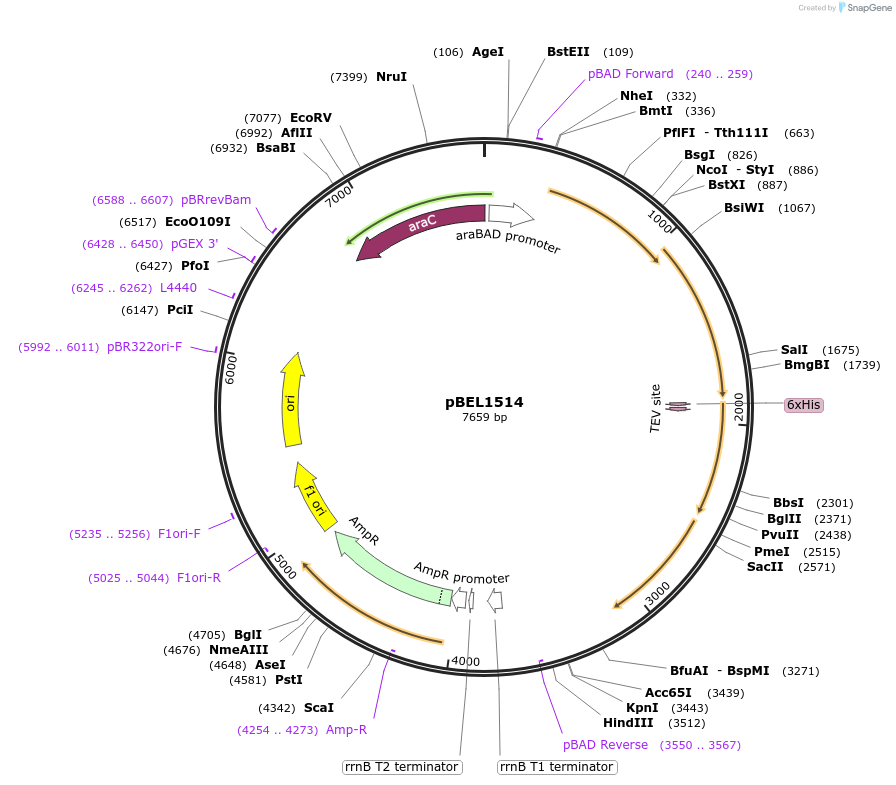
pBEL1514
Plasmid#155147PurposeExpresses MlaFEDCB operon with MlaF C-terminus deleted [1-246], His-tag on MlaDDepositorInsertmlaF(1-246)-mlaE-[6xHis-2xQH-TEV-mlaD]-mlaCB operon
PromoteraraBADAvailable SinceDec. 1, 2020AvailabilityAcademic Institutions and Nonprofits only -
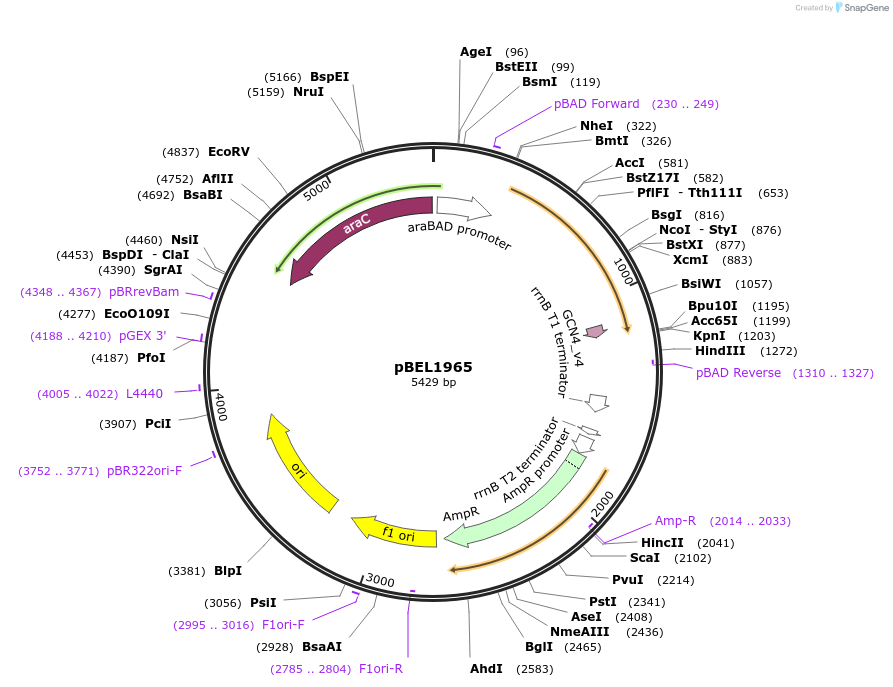
pBEL1965
Plasmid#155152PurposeExpresses MlaF(1-246)-(6 aa linker)-GCN4DepositorInsertmlaF(1-246)-GCN4_Dimerization_Domain
Mutationdeleted amino acids 247-269 in MlaF; added dimeri…PromoteraraBADAvailable SinceDec. 1, 2020AvailabilityAcademic Institutions and Nonprofits only -
pBEL1966
Plasmid#155153PurposeExpresses MlaF(1-246)-(10 aa linker)-GCN4DepositorInsertmlaF(1-246)-GCN4_Dimerization_Domain
Mutationdeleted amino acids 247-269 in MlaF; added dimeri…PromoteraraBADAvailable SinceDec. 1, 2020AvailabilityAcademic Institutions and Nonprofits only