We narrowed to 1,843 results for: FAST
-
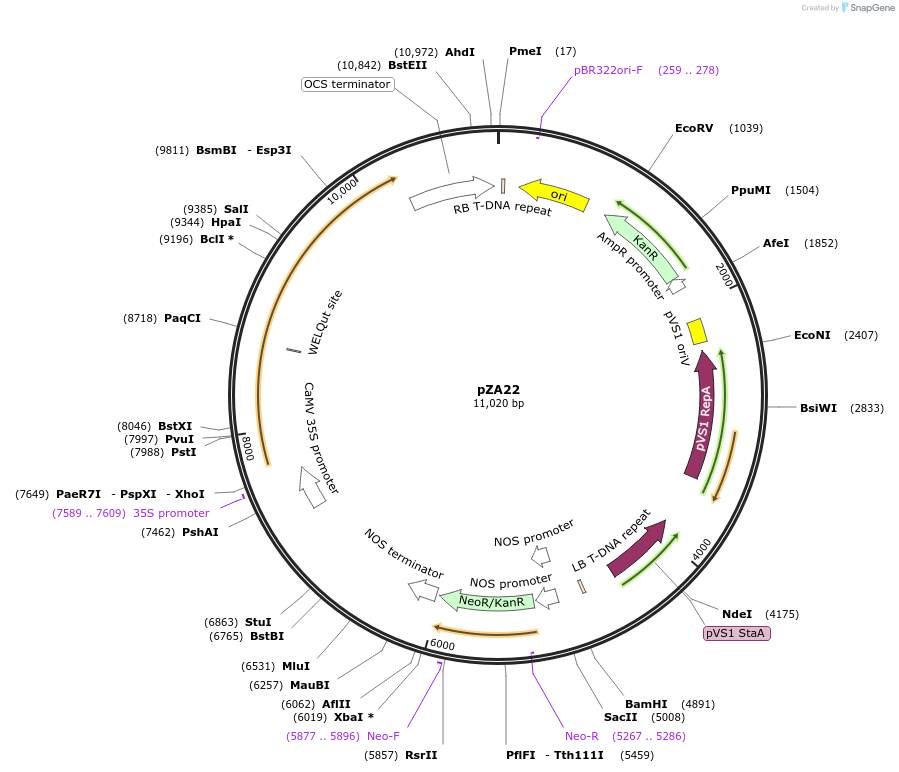
Plasmid#158500PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantMutationD481VAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
pZA45
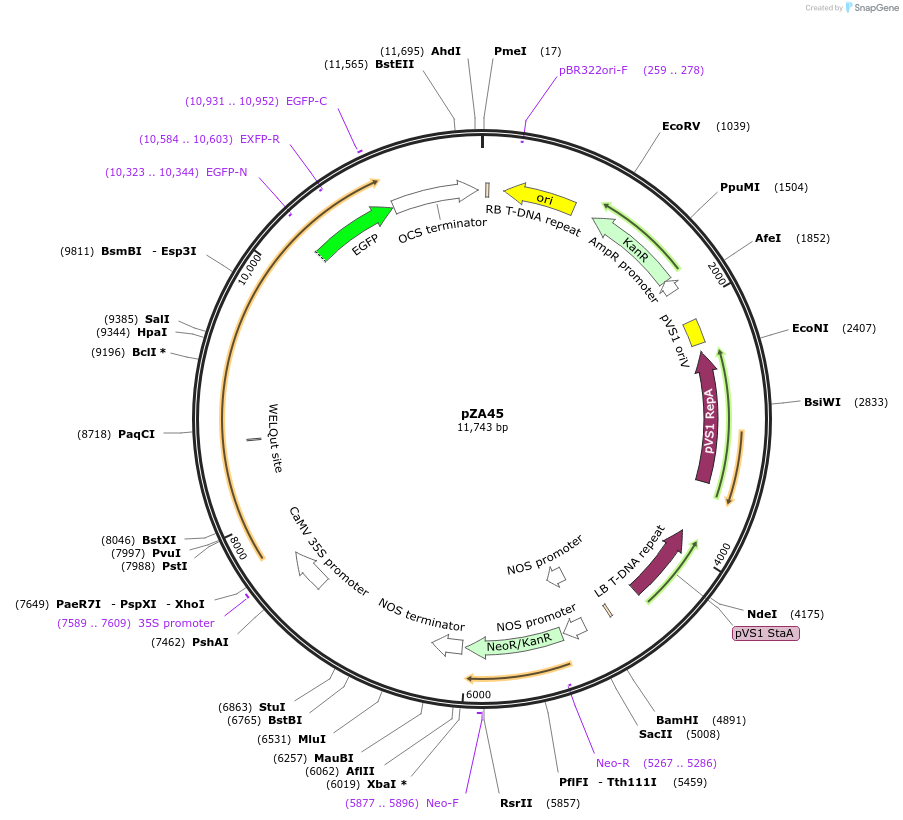
Plasmid#158523PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-eGFP. Km plant selection marker.DepositorInsertZAR1-eGFP
TagseGFPExpressionPlantAvailable SinceSept. 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
pZA37
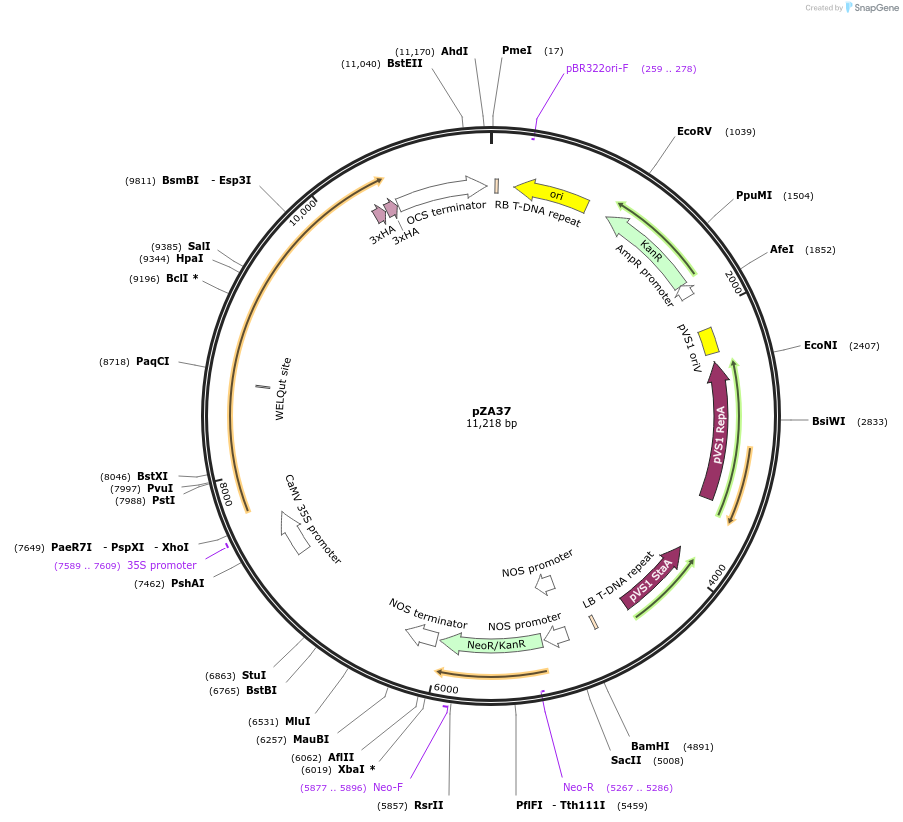
Plasmid#158515PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-6xHA. Km plant selection marker.DepositorInsertZAR1-6xHA
Tags6xHAExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
pZA39
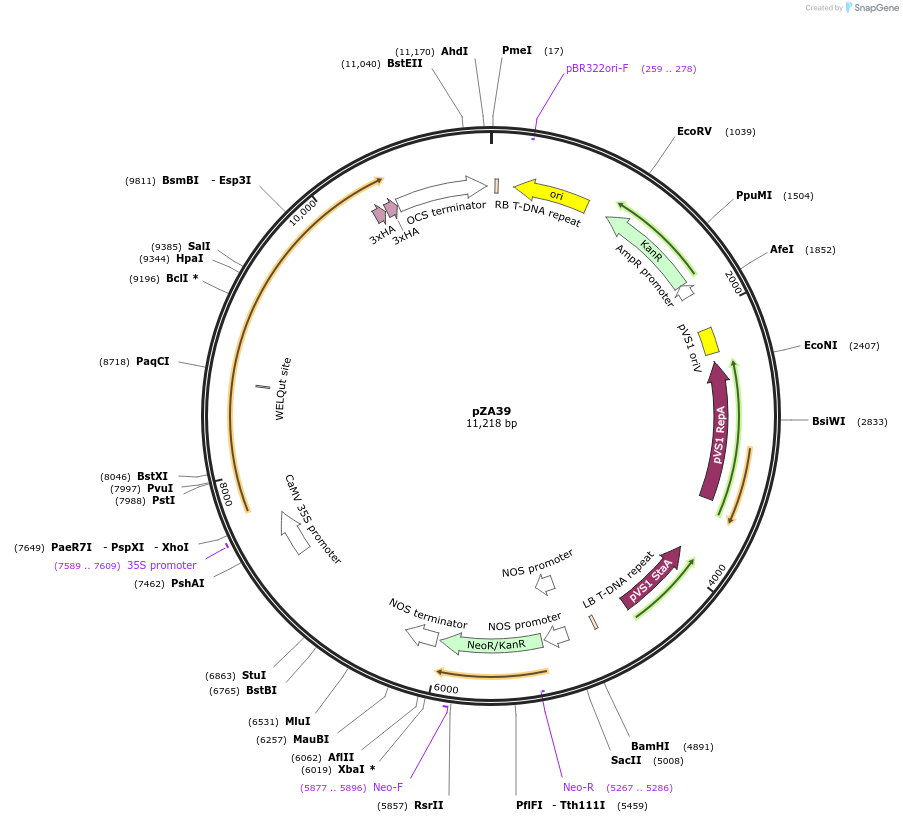
Plasmid#158517PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 L17E-6xHA. Km plant selection marker.DepositorInsertZAR1-6xHA
Tags6xHAExpressionPlantMutationL17EAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
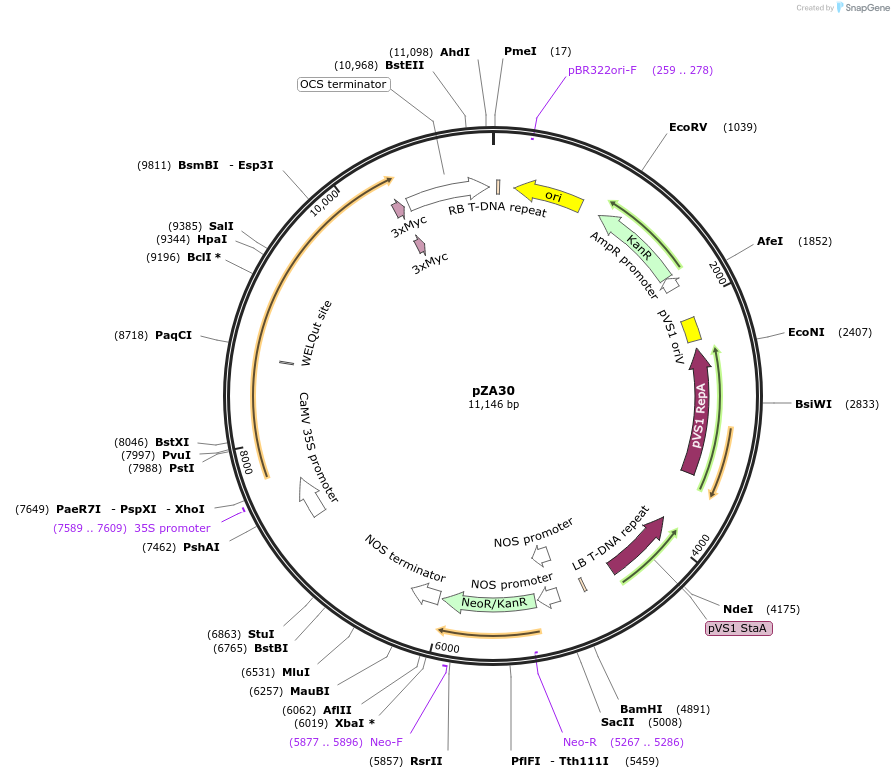
pZA30
Plasmid#158508PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V-4xMyc. Km plant selection marker.DepositorInsertZAR1-4xMyc
Tags4xMycExpressionPlantMutationD481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
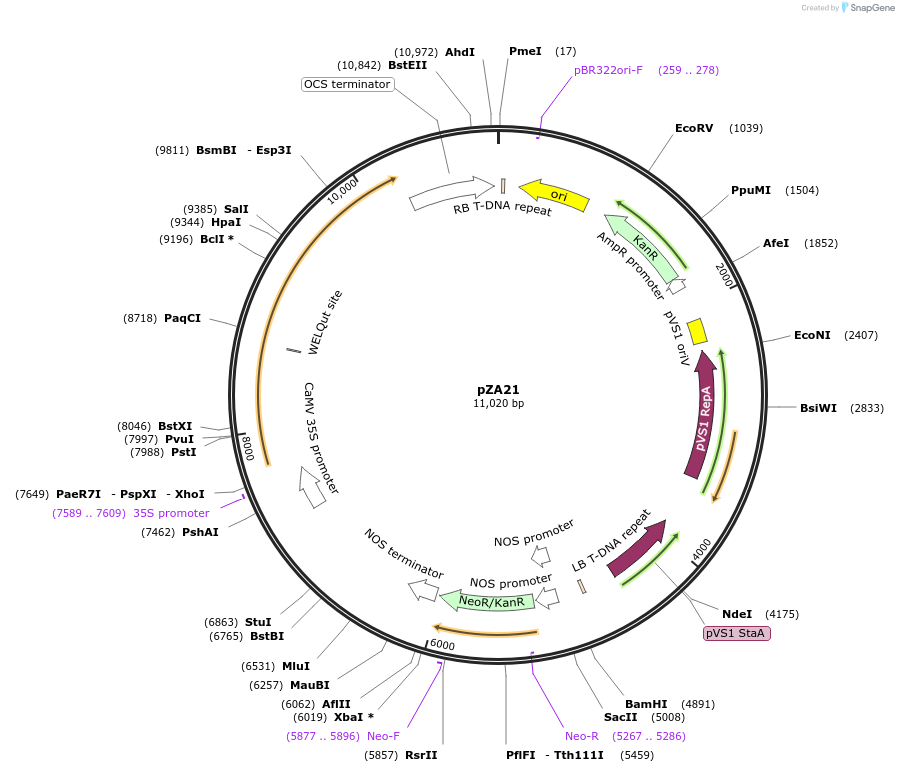
pZA21
Plasmid#158499PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
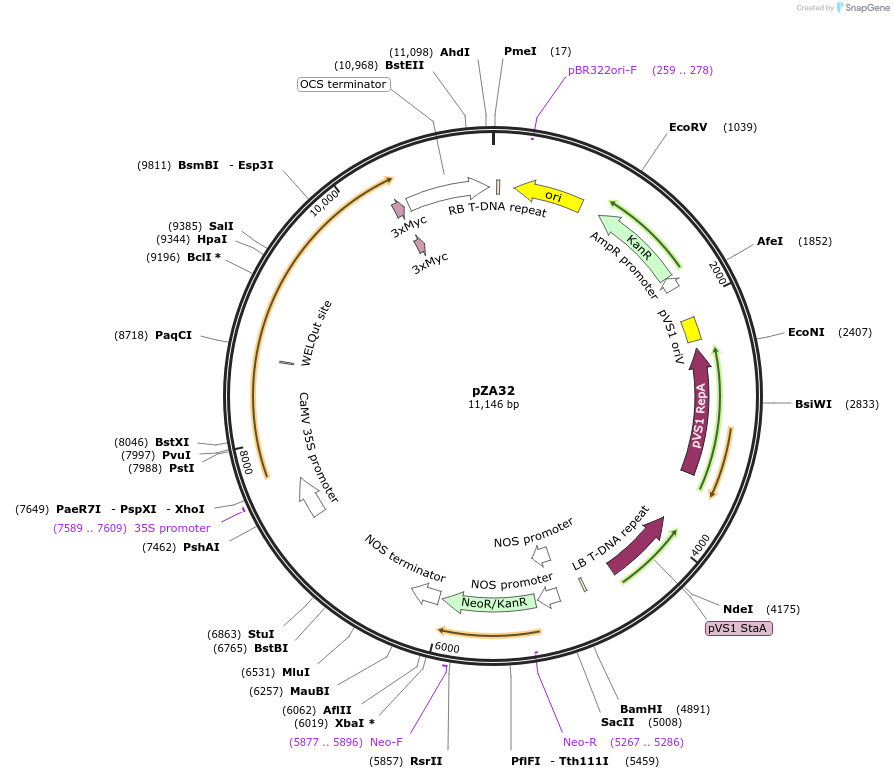
pZA32
Plasmid#158510PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V/L17E-4xMyc. Km plant selection marker.DepositorInsertZAR1-4xMyc
Tags4xMycExpressionPlantMutationL17E, D481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
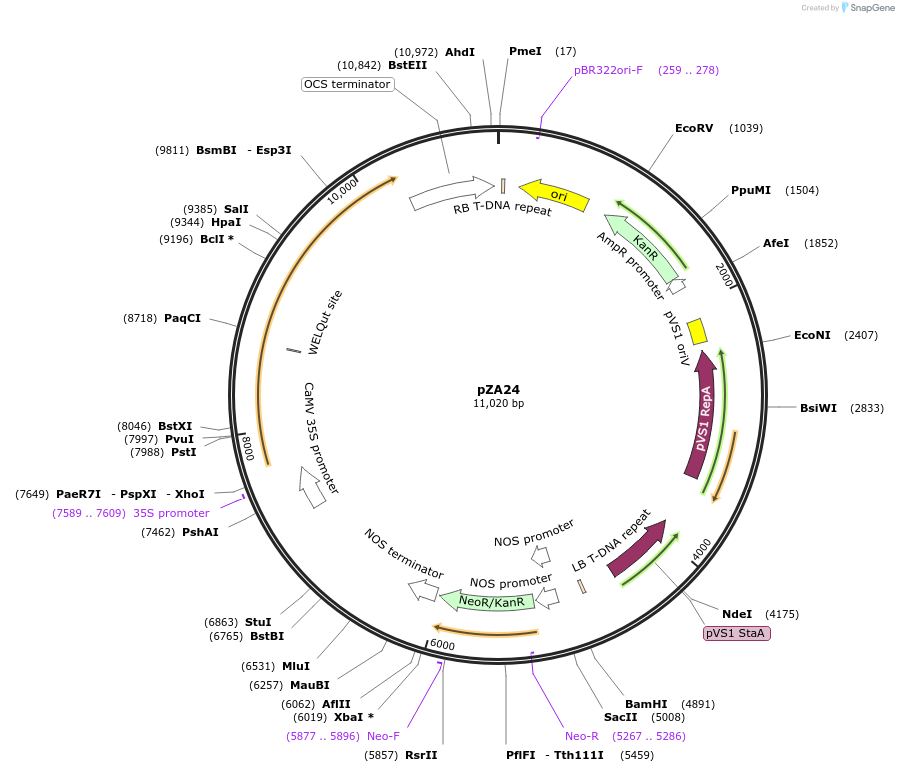
pZA24
Plasmid#158502PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V/L17E. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantMutationL17E, D481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
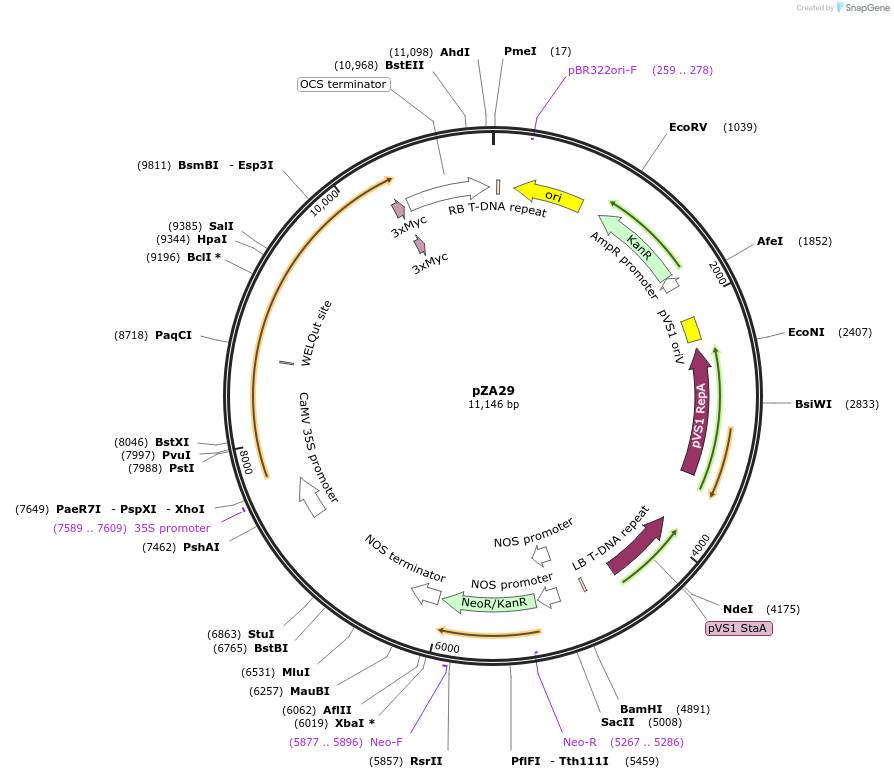
pZA29
Plasmid#158507PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1-4xMyc. Km plant selection marker.DepositorInsertZAR1-4xMyc
Tags4xMycExpressionPlantAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
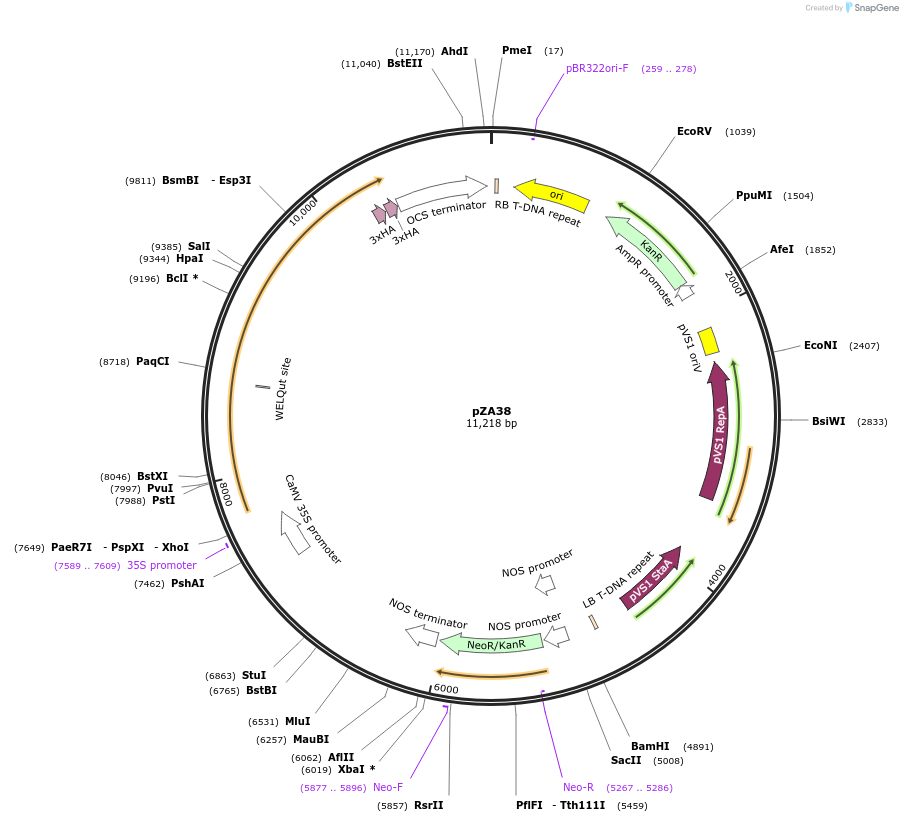
pZA38
Plasmid#158516PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V-6xHA. Km plant selection marker.DepositorInsertZAR1-6xHA
Tags6xHAExpressionPlantMutationD481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
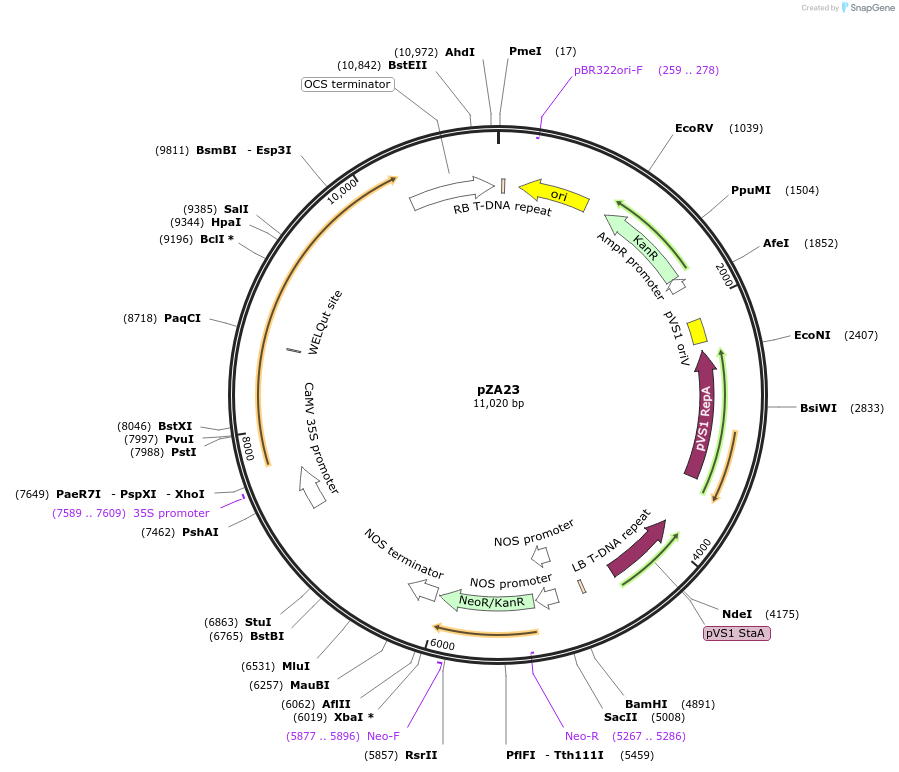
pZA23
Plasmid#158501PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 L17E. Km plant selection marker.DepositorInsertZAR1
ExpressionPlantMutationL17EAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
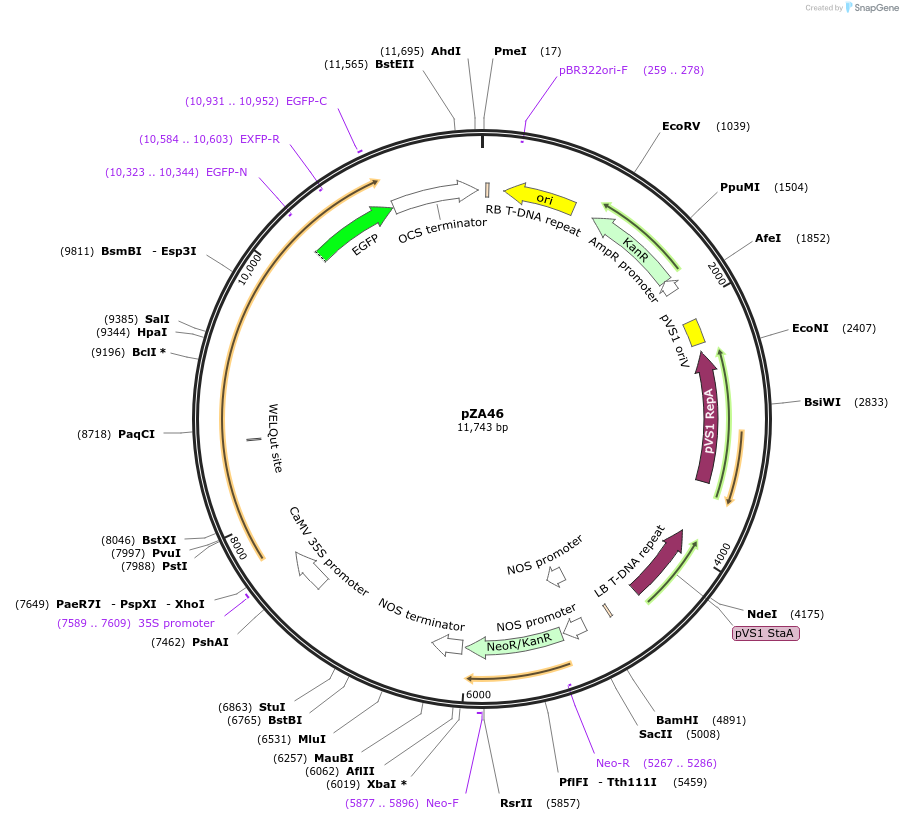
pZA46
Plasmid#158524PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V-eGFP. Km plant selection marker.DepositorInsertZAR1-eGFP
TagseGFPExpressionPlantMutationD481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
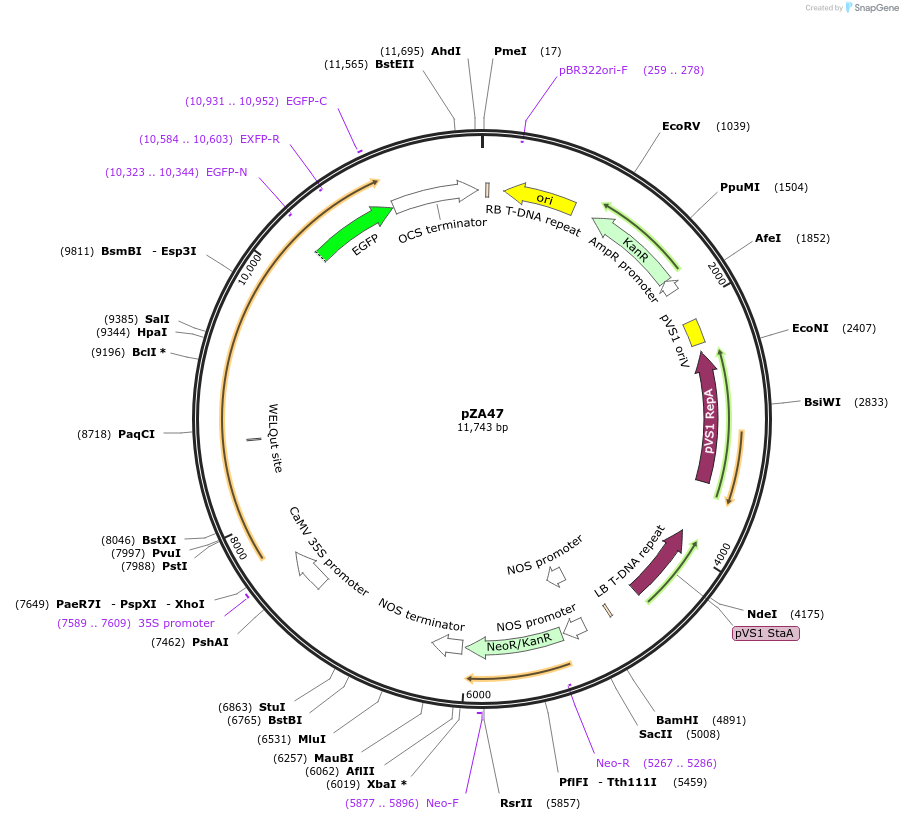
pZA47
Plasmid#158525PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 L17E-eGFP. Km plant selection marker.DepositorInsertZAR1-eGFP
TagseGFPExpressionPlantMutationL17EAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
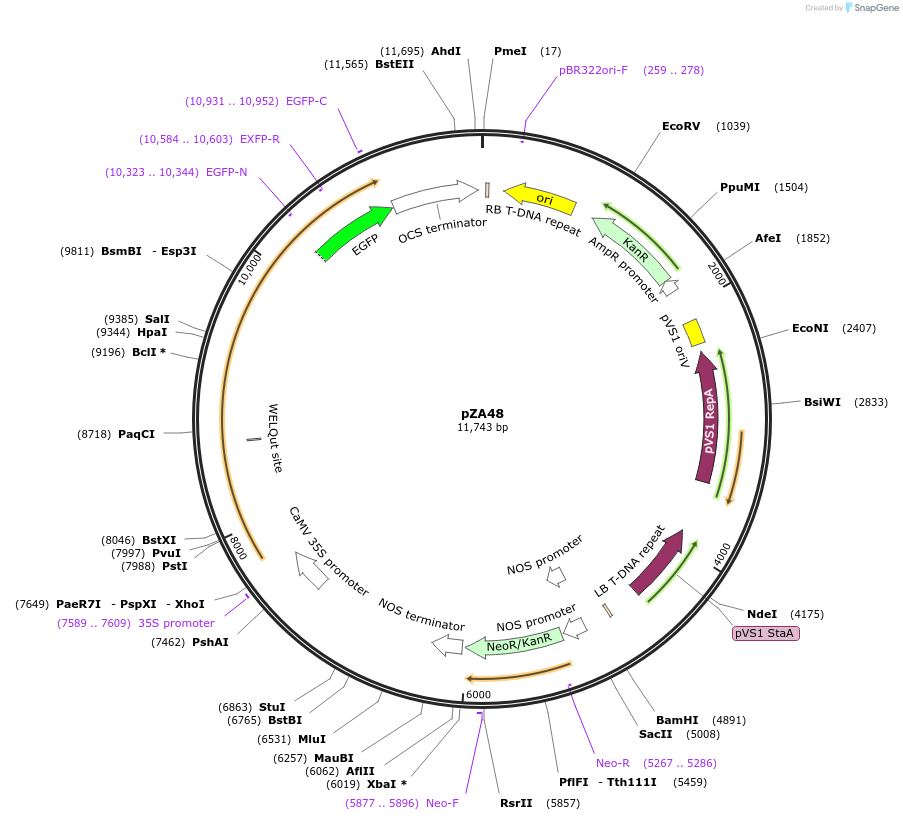
pZA48
Plasmid#158526PurposeIn planta gene expression of Golden gate compatible N. benthamiana ZAR1 D481V/L17E-eGFP. Km plant selection marker.DepositorInsertZAR1-eGFP
TagseGFPExpressionPlantMutationL17E, D481VAvailable SinceAug. 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
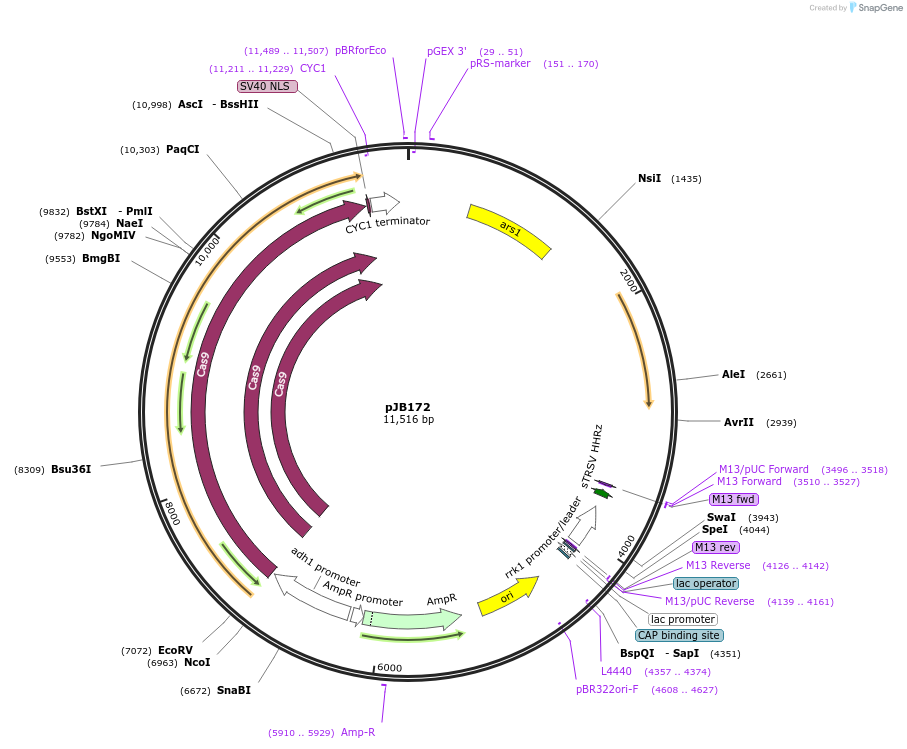
pJB172
Plasmid#86992PurposeCRISPR/Cas9 in fission yeast using fluoride selection and targetting pil1DepositorInsertgRNA targeting pil1
ExpressionYeastAvailable SinceFeb. 22, 2017AvailabilityAcademic Institutions and Nonprofits only -
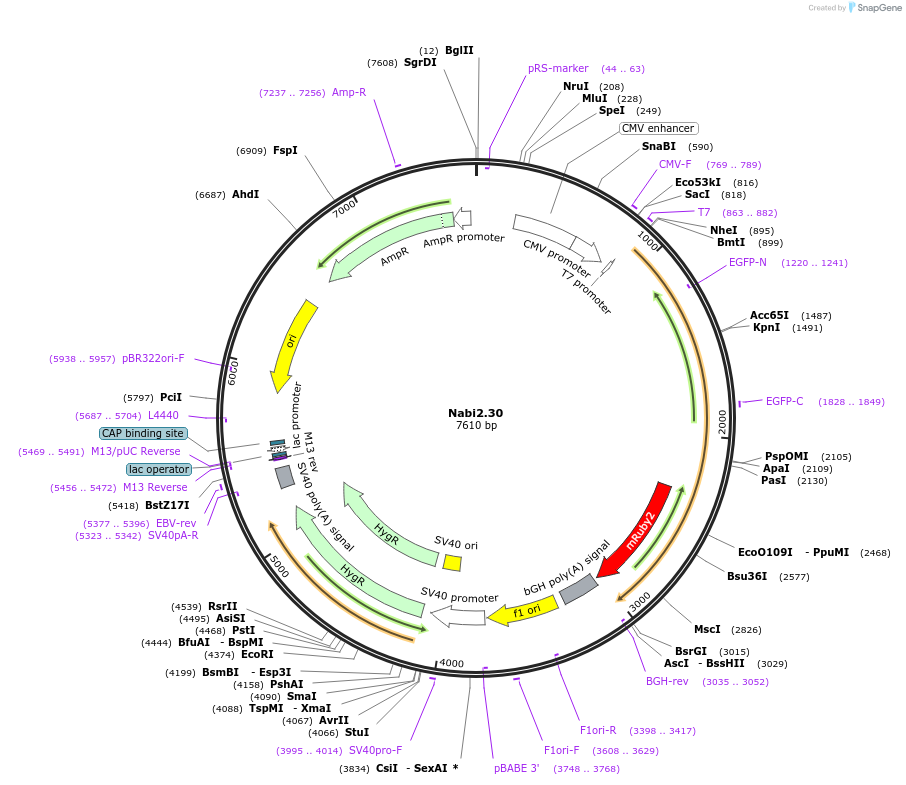
Nabi2.30
Plasmid#73798PurposeNabi2.30 is a FRET (Förster Resonance Energy Transfer)-based protein voltage sensor. This probe contains Clover and mRuby2 inserted at different locations in the Ciona voltage sensitive domain.DepositorInsertNabi2.30
ExpressionMammalianAvailable SinceApril 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
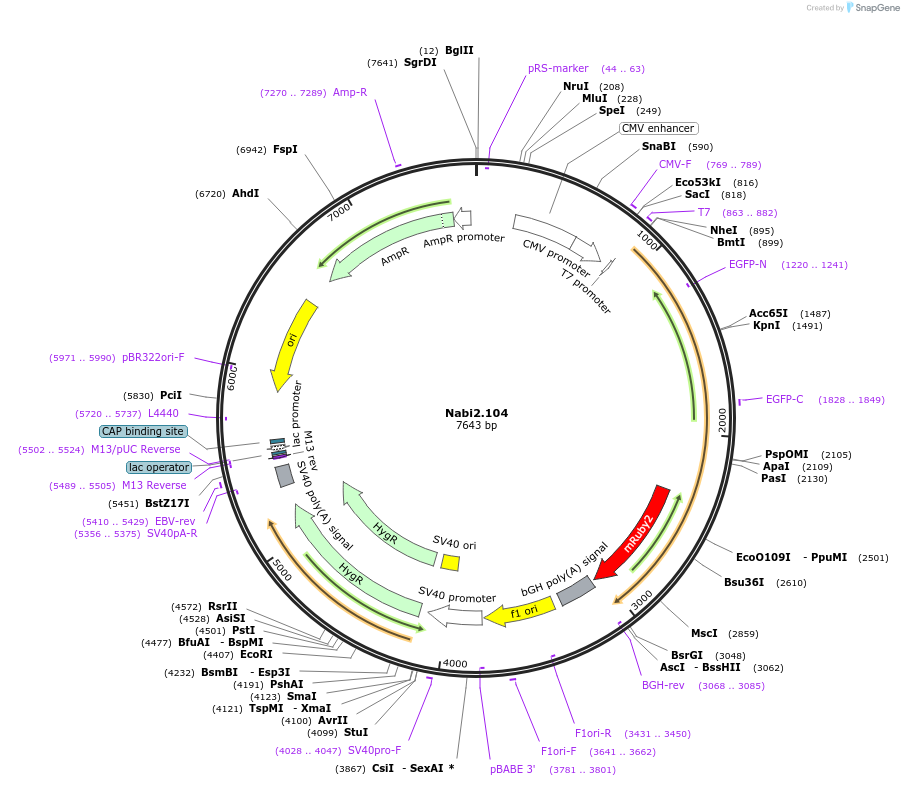
Nabi2.104
Plasmid#73799PurposeNabi2.104 is a FRET (Förster Resonance Energy Transfer)-based protein voltage sensor. This probe contains Clover and mRuby2 inserted at different locations in the Ciona voltage sensitive domain.DepositorInsertNabi2.104
ExpressionMammalianAvailable SinceApril 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
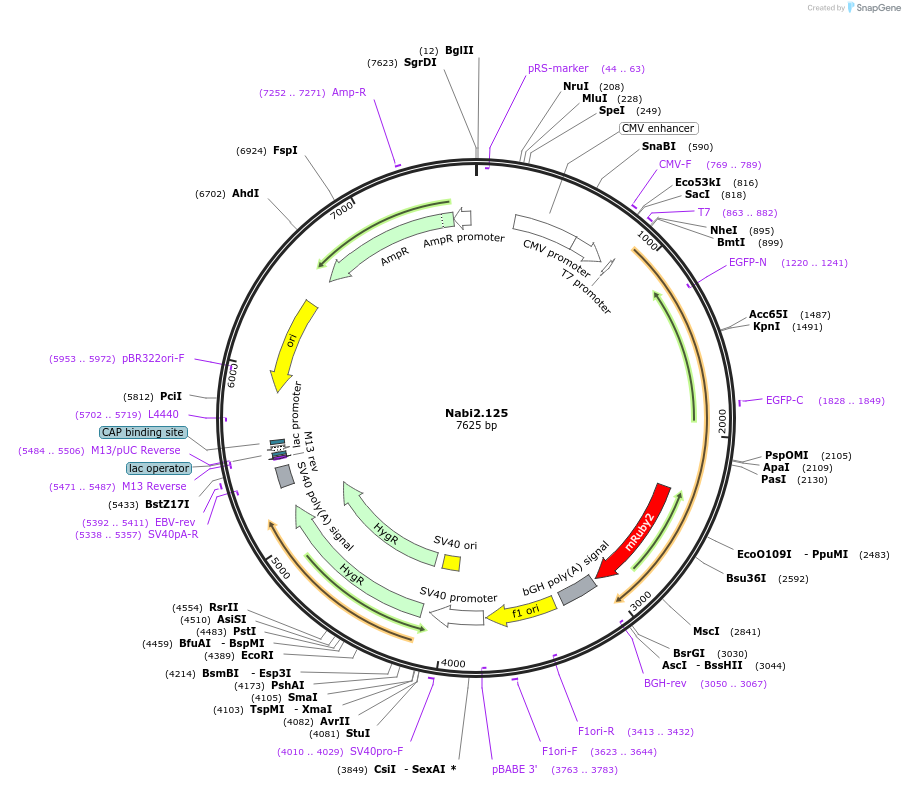
Nabi2.125
Plasmid#73800PurposeNabi2.125 is a FRET (Förster Resonance Energy Transfer)-based protein voltage sensor. This probe contains Clover and mRuby2 inserted at different locations in the Ciona voltage sensitive domain.DepositorInsertNabi2.125
ExpressionMammalianAvailable SinceApril 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
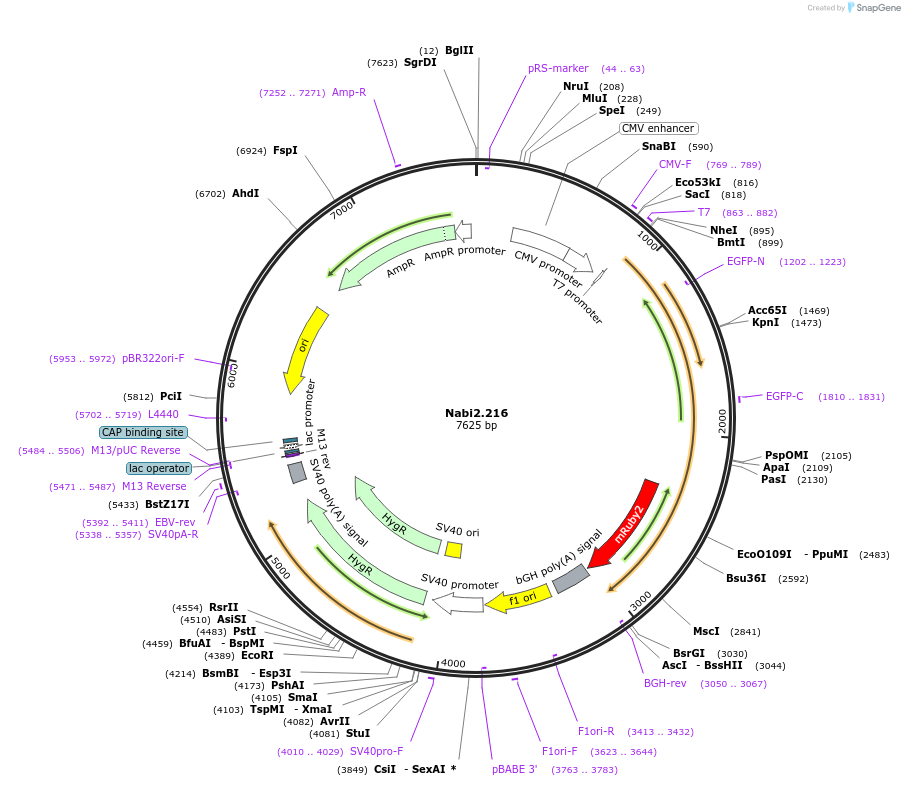
Nabi2.216
Plasmid#73801PurposeNabi2.216 is a FRET (Förster Resonance Energy Transfer)-based protein voltage sensor. This probe contains Clover and mRuby2 inserted at different locations in the Ciona voltage sensitive domain.DepositorInsertNabi2.216
ExpressionMammalianAvailable SinceApril 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
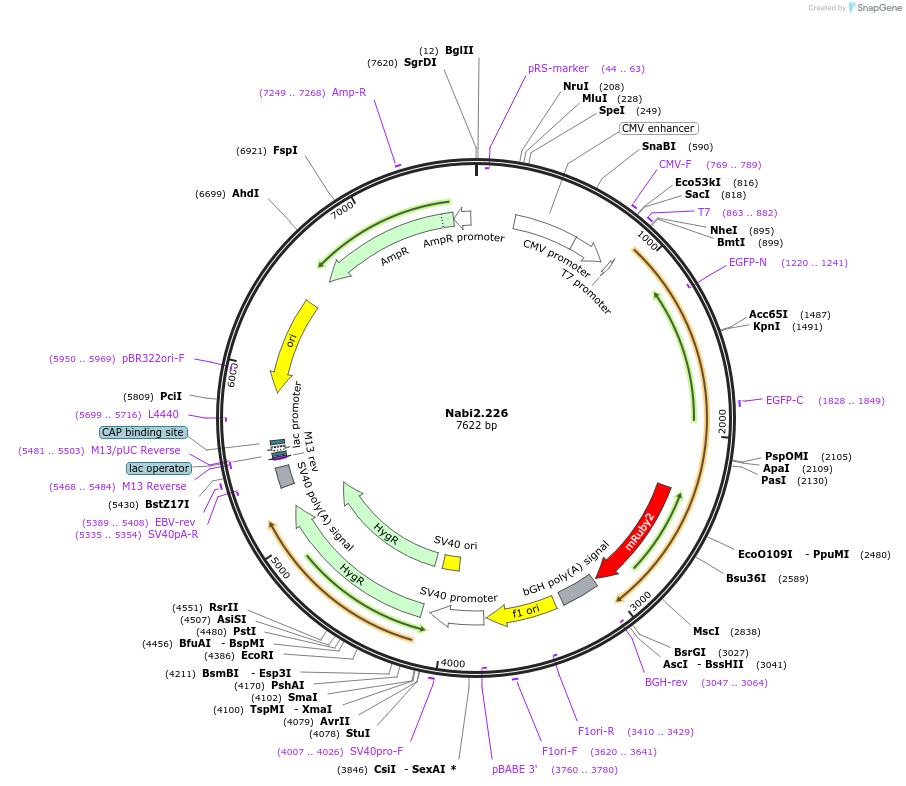
Nabi2.226
Plasmid#73802PurposeNabi2.226 is a FRET (Förster Resonance Energy Transfer)-based protein voltage sensor. This probe contains Clover and mRuby2 inserted at different locations in the Ciona voltage sensitive domain.DepositorInsertNabi2.226
ExpressionMammalianAvailable SinceApril 7, 2016AvailabilityAcademic Institutions and Nonprofits only