-
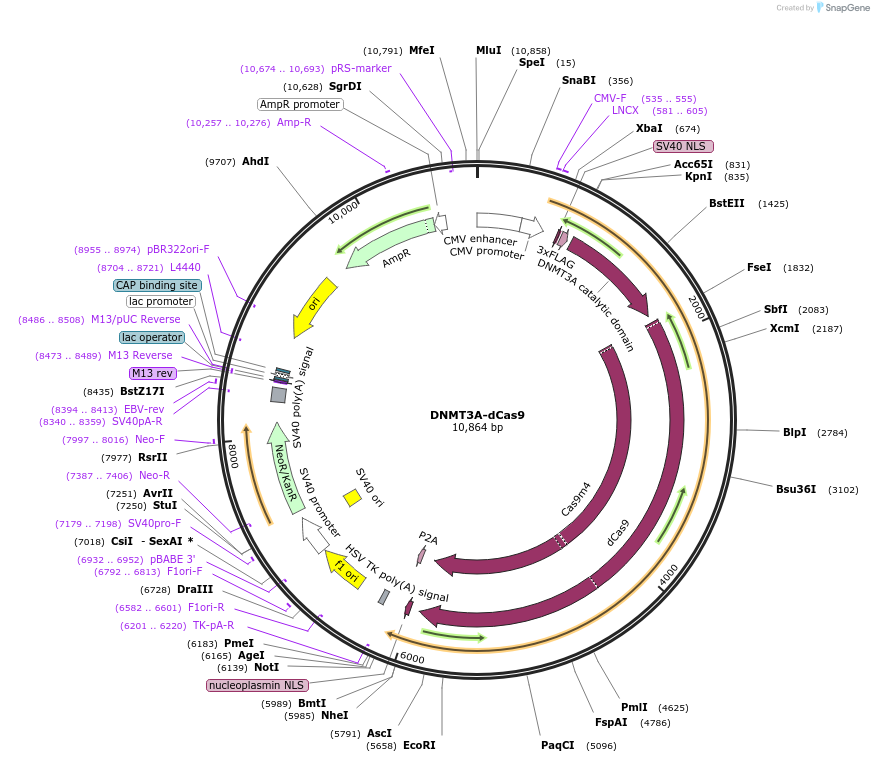
PurposeDNMT3A fused to N-terminus of dCas9; pCDNA3 vector backbone, mammalian expression
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 100090 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepcDNA3.3-TOPO
-
Backbone manufacturerThermoFisher
- Backbone size w/o insert (bp) 5407
- Total vector size (bp) 10864
-
Vector typeMammalian Expression, CRISPR
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameDNMT3A
-
SpeciesH. sapiens (human)
-
Mutationincludes aa 602-912
-
GenBank IDNP_072046.2
-
Entrez GeneDNMT3A (a.k.a. DNMT3A2, HESJAS, M.HsaIIIA, TBRS)
- Promoter CMV
-
Tag
/ Fusion Protein
- 3XFLag-NLS-DNMT3A-dCas9-NLS
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer CMV-F
- 3′ sequencing primer polyA TK-R
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
DNMT3A-dCas9 was a gift from David Segal (Addgene plasmid # 100090 ; http://n2t.net/addgene:100090 ; RRID:Addgene_100090) -
For your References section:
dCas9-based epigenome editing suggests acquisition of histone methylation is not sufficient for target gene repression. O'Geen H, Ren C, Nicolet CM, Perez AA, Halmai J, Le VM, Mackay JP, Farnham PJ, Segal DJ. Nucleic Acids Res. 2017 Sep 29;45(17):9901-9916. doi: 10.1093/nar/gkx578. 10.1093/nar/gkx578 PubMed 28973434