-
PurposeCas9-GFP, dual gRNA expression vector for targeted integration of repair templates into safe harbor 231 locus in chromosome 4
-
Depositing Labs
-
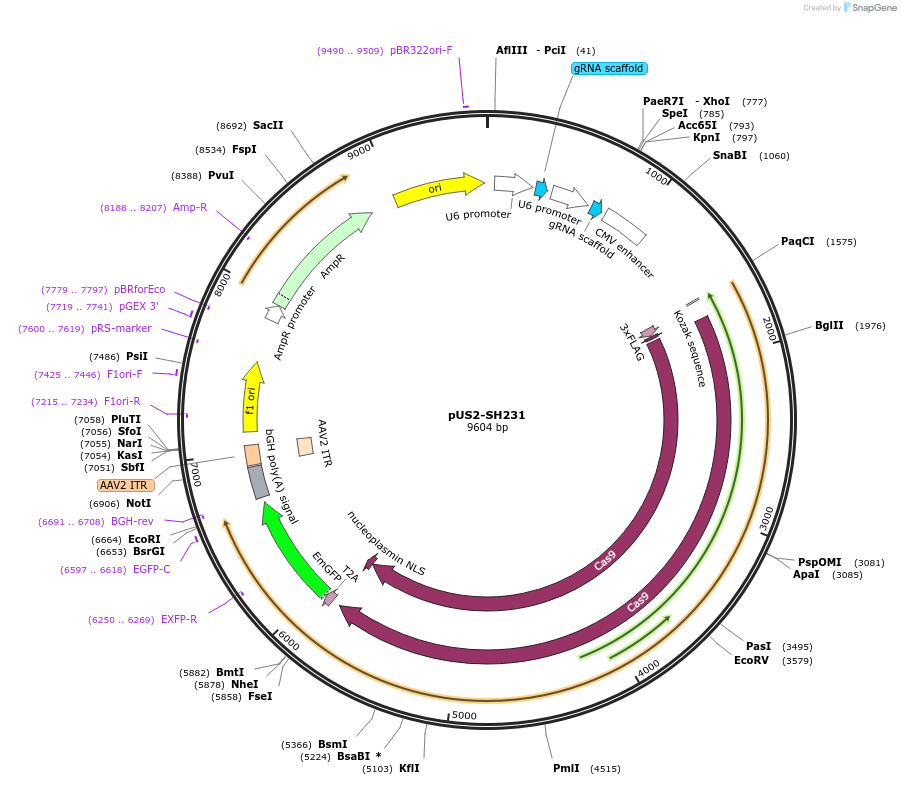
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 115150 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonecustom
- Backbone size w/o insert (bp) 2739
-
Vector typeMammalian Expression, CRISPR
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameDual SH231 gRNAs
-
Alt nameCas9-T2A-GFP
-
gRNA/shRNA sequencegcctcccccatagtaccat, Gatgtgctcactgagtctga
-
SpeciesStreptococcus pyogenes
-
Insert Size (bp)6866
- Promoter U6, CBh
-
Tags
/ Fusion Proteins
- 3xFLAG-SV40NLS (N terminal on backbone)
- GFP (C terminal on insert)
- Nucleoplasmin NLS (C terminal on backbone)
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer unknown
- (Common Sequencing Primers)
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Please visit https://www.biorxiv.org/content/early/2018/08/20/396390 for bioRxiv preprint.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pUS2-SH231 was a gift from Raymond Monnat & Michael Phelps (Addgene plasmid # 115150 ; http://n2t.net/addgene:115150 ; RRID:Addgene_115150) -
For your References section:
New Human Chromosomal Sites with "Safe Harbor" Potential for Targeted Transgene Insertion. Pellenz S, Phelps M, Tang W, Hovde BT, Sinit RB, Fu W, Li H, Chen E, Monnat RJ Jr. Hum Gene Ther. 2019 Jul;30(7):814-828. doi: 10.1089/hum.2018.169. Epub 2019 Mar 28. 10.1089/hum.2018.169 PubMed 30793977