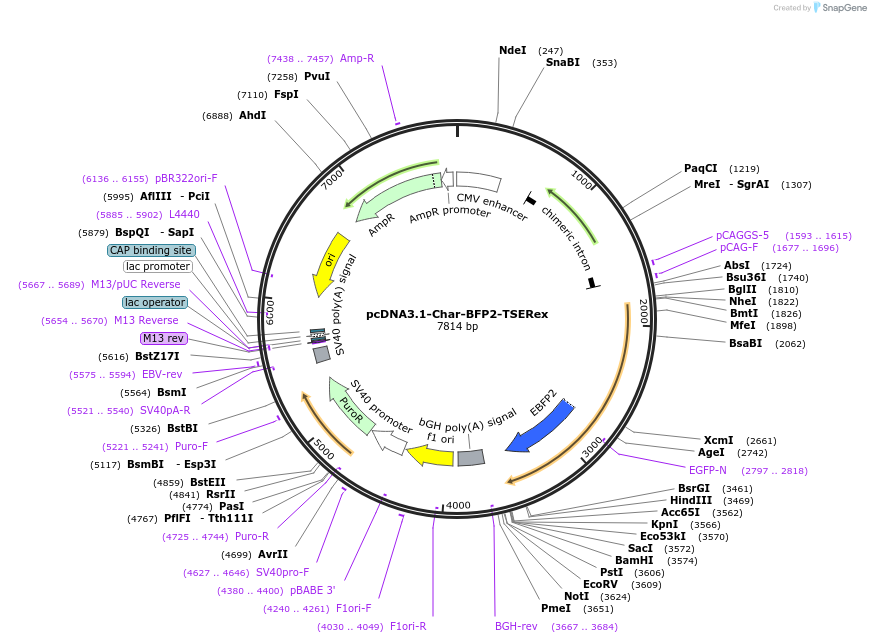
pcDNA3.1-Char-BFP2-TSERex
(Plasmid
#124792)
-
PurposeExpresses the proton pump rhodopsin in mammalian cells under CAG promoter
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 124792 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepcDNA3.1/puro-CAG
-
Backbone manufacturerMichael Lin lab (Addgene plasmid 52519)
- Backbone size w/o insert (bp) 6105
- Total vector size (bp) 7779
-
Vector typeMammalian Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameChar-BFP2
-
SpeciesH. sapiens (human)
-
Insert Size (bp)1680
- Promoter CAG
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer pCAG-F
- (Common Sequencing Primers)
Resource Information
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Addgene NGS found a single nucleotide deletion in the Kozak sequence upstream of Char-BFP2 compared to a reference sequence provided by the depositing lab
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pcDNA3.1-Char-BFP2-TSERex was a gift from Yulong Li (Addgene plasmid # 124792 ; http://n2t.net/addgene:124792 ; RRID:Addgene_124792) -
For your References section:
PARIS, an optogenetic method for functionally mapping gap junctions. Wu L, Dong A, Dong L, Wang SQ, Li Y. Elife. 2019 Jan 14;8. pii: 43366. doi: 10.7554/eLife.43366. 10.7554/eLife.43366 PubMed 30638447