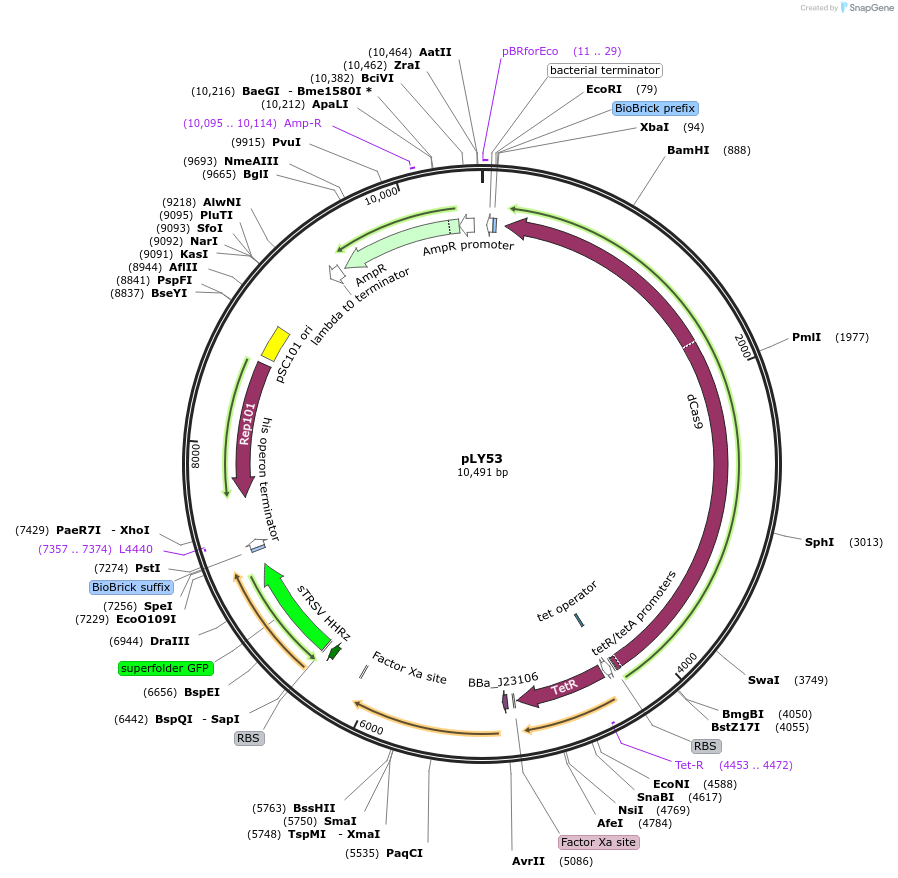
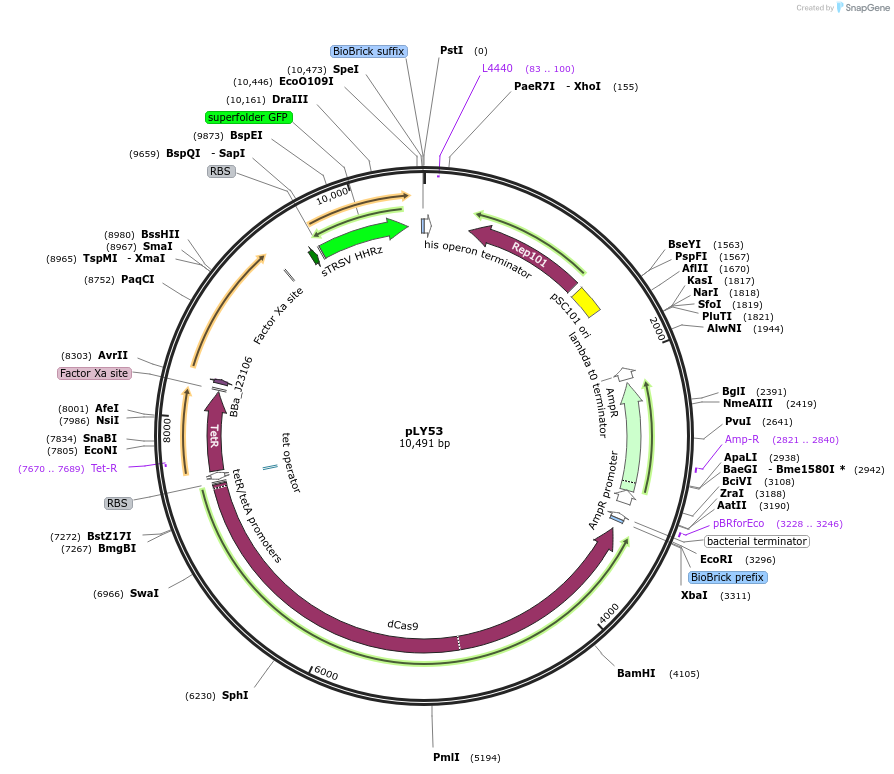
pLY53
(Plasmid
#130911)
-
PurposeA CRISPR activation device with the necessary genes (dcas9 controlled by Ptet, pspFΔHTH::λN22plus controlled by promoter J23106), and the reporter part (PpspA-LEA1B1 with sfgfp::ASV).
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 130911 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepSB4A3mut
-
Backbone manufacturerOriginal version designed by: Reshma Shetty; Mutations introduced by: unknown
- Backbone size w/o insert (bp) 3337
- Total vector size (bp) 10491
-
Vector typeBacterial Expression, Synthetic Biology
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Growth instructionsCulture in the Lennox’s Luria-Bertani (LB) medium (10 g/L peptone, 5 g/L yeast extract, 5 g/L NaCl)
-
Copy numberLow Copy
Gene/Insert 1
-
Gene/Insert namedcas9
-
SpeciesStreptococcus pyogenes
-
Insert Size (bp)4170
-
MutationMutations D10A and H840A on WT cas9. Synonymous mutation at R63 of dcas9 (CGT>CGC); Synonymous mutation at R447 of dcas9 (CGA>CGT)
- Promoter Ptet
Cloning Information for Gene/Insert 1
- Cloning method Unknown
- 5′ sequencing primer TGCCACCTGACGTCTAAGAA
- 3′ sequencing primer TGCGCTGTTAATCACTTTACTTTTATCTAATCTTG
- (Common Sequencing Primers)
Gene/Insert 2
-
Gene/Insert nametetR
-
SpeciesE. coli
-
Insert Size (bp)790
-
MutationSynonymous mutation at S2 of tetR (TCT>TCA)
- Promoter Ptet
Cloning Information for Gene/Insert 2
- Cloning method Unknown
- 5′ sequencing primer GAGCTGTCCGTTTGAGGCGAGTCGCTTCCGCTG
- 3′ sequencing primer CACGAATATTTCCCGGCCAACGATAATTCAGC
- (Common Sequencing Primers)
Gene/Insert 3
-
Gene/Insert namepspFΔHTH::λN22plus
-
SpeciesSynthetic; E. coli
-
Insert Size (bp)1084
-
MutationThe HTH domain of the pspF gene was deleted (297-326). Mutations D2N and Q4R in the λN22 domain.
- Promoter Anderson promoter: J23106
-
Tag
/ Fusion Protein
- pspFΔHTH::λN22plus (C terminal on insert)
Cloning Information for Gene/Insert 3
- Cloning method Restriction Enzyme
- 5′ cloning site XbaI (destroyed during cloning)
- 3′ cloning site SpeI (destroyed during cloning)
- 5′ sequencing primer AGCATCAAGTCGCTAAAGAAGAAAGGGAAACACCTACTACTG
- 3′ sequencing primer CTCGCCACGCACGGAAAACTTATGACCGTTGAC
- (Common Sequencing Primers)
Gene/Insert 4
-
Gene/Insert namesfgfp
-
SpeciesSynthetic
-
Insert Size (bp)1110
- Promoter PpspA-LEA1B1
-
Tag
/ Fusion Protein
- ASV tag (C terminal on insert)
Cloning Information for Gene/Insert 4
- Cloning method Restriction Enzyme
- 5′ cloning site XbaI (destroyed during cloning)
- 3′ cloning site SpeI, PstI (not destroyed)
- 5′ sequencing primer CTGCAACTCAGTTTGCAAATGAATGCAC
- 3′ sequencing primer ATTACCGCCTTTGAGTGAGC
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The Top10 bacterial strain should be used for downstream applications
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pLY53 was a gift from Baojun Wang (Addgene plasmid # 130911 ; http://n2t.net/addgene:130911 ; RRID:Addgene_130911) -
For your References section:
Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria. Liu Y, Wan X, Wang B. Nat Commun. 2019 Aug 26;10(1):3693. doi: 10.1038/s41467-019-11479-0. 10.1038/s41467-019-11479-0 PubMed 31451697