-
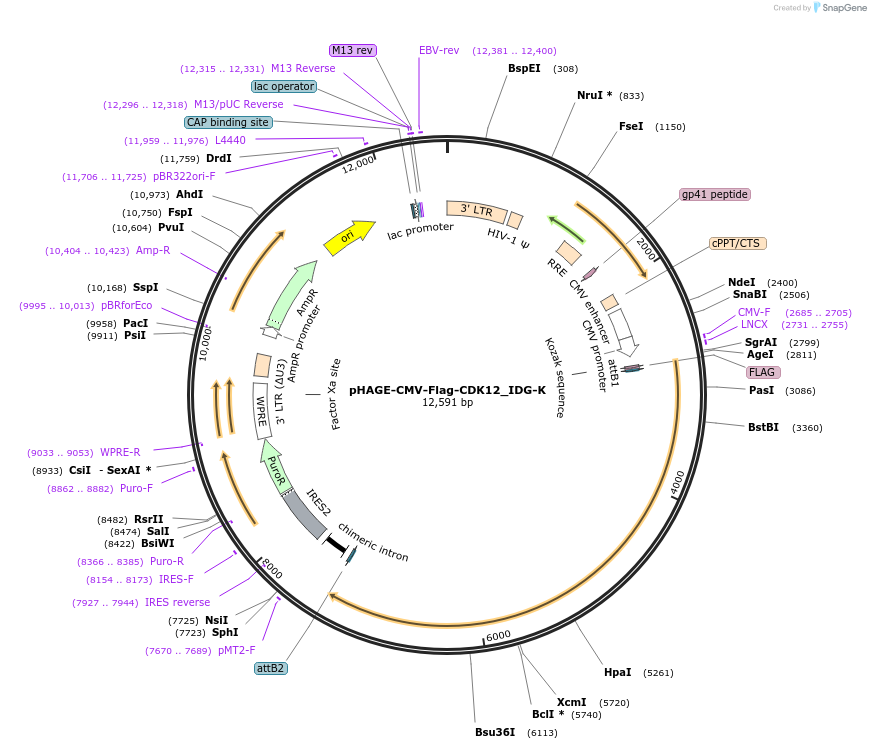
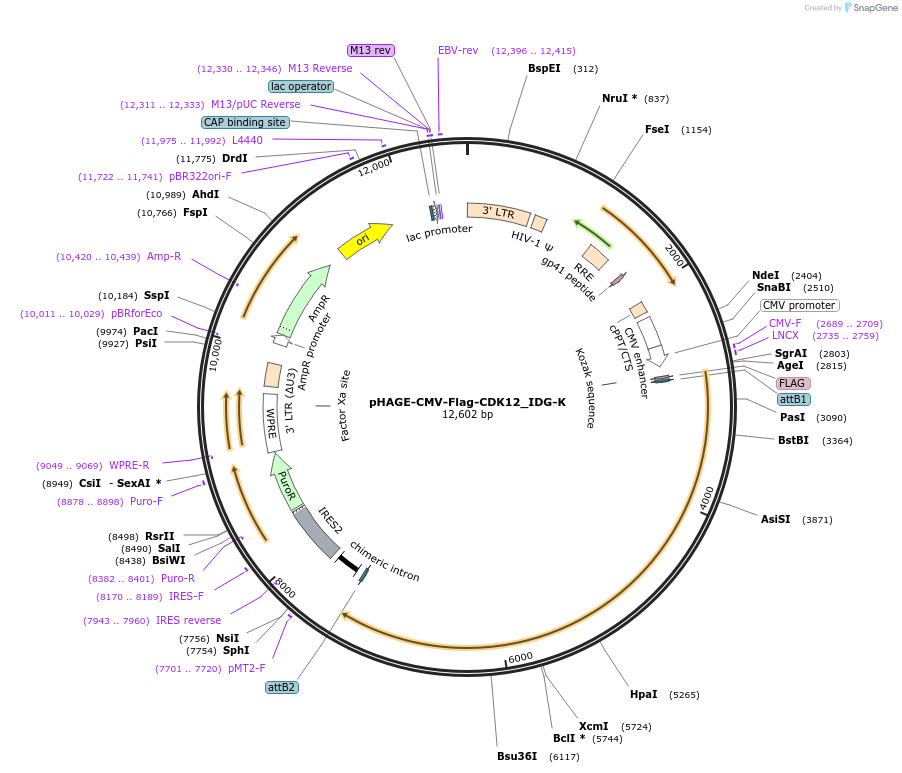
PurposeGateway destination clone of CDK12 (human) tagged with N-terminal Flag for generating protein-protein networks using fusion tag affinity-based proteomics
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 135281 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepHAGE
- Backbone size w/o insert (bp) 9763
-
Vector typeMammalian Expression, Lentiviral ; Gateway Destination
-
Selectable markersPuromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameFlag-CDK12
-
Alt nameCDK12
-
SpeciesH. sapiens (human)
-
Entrez GeneCDK12 (a.k.a. CRK7, CRKR, CRKRS)
- Promoter CMV
-
Tag
/ Fusion Protein
- Flag (N terminal on insert)
Cloning Information
- Cloning method Gateway Cloning
- 5′ sequencing primer CGCAAATGGGCGGTAGGCGTG
- 3′ sequencing primer CAATCTTAGCGCAGAAGTCATG
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
These plasmids were generated as part of the Illuminating the Druggable Genome (IDG) program sponsored by the NIH Common Fund. The goal of this program is to identify, gather, and distribute information and resources for proteins that currently are not well-studied yet belong to commonly drug-targeted protein families: protein kinases, non-olfactory G-protein coupled receptors (GPCRs), and ion channels. The IDG program is designed to develop fundamental research tools for understudied proteins, elucidate their function, and disseminate the IDG-related resources and data to the greater scientific community. These lentiviral gateway destination plasmids were generated, as part of the Kinase-Data and Resource Generating Center (DRGC), using the single fragment or multisite gateway cloning technology. They were used for generating proteomics-based protein-interaction and protein-proximity networks that can be accessed through the kinase-DRGC website https://darkkinome.org/. Each plasmid has either an N- or C-terminal fusion protein (Flag/ V5/V5-miniTurbo/V5-TurboID/miniTurbo-V5/TurboID-V5) tagged understudied kinase driven under either a CMV or Ubiquitin (UBC) promoter. All inserts in the pENTR plasmids used to generate the deposited destination plasmids were end-sequenced and insert size validated prior to multi-site gateway cloning. The combined insert size of the single fragment or three fragment destination plasmids were confirmed by restriction digestion (BsrGI, and EcoRV for pDest667; BsrGI, EcoRV, and BstZ17I for pDest663; and BsrGI for pHAGE) and all plasmids were partially sequenced to ensure in-frame ORFs and fusion tags.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pHAGE-CMV-Flag-CDK12_IDG-K was a gift from Ben Major (Addgene plasmid # 135281 ; http://n2t.net/addgene:135281 ; RRID:Addgene_135281)