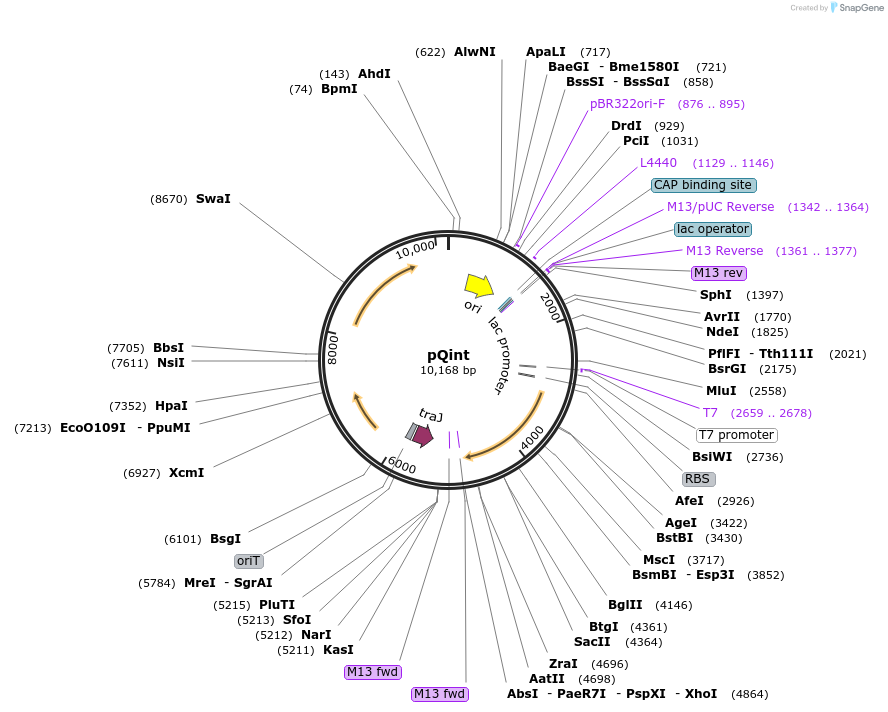
pQint
(Plasmid
#25819)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 25819 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepAT19
- Backbone size w/o insert (bp) 6559
-
Vector typebacterial gene inactivation
-
Selectable markerserythromycin
Growth in Bacteria
-
Bacterial Resistance(s)Erythromycin, 200 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Growth instructionsreplicates in Clostridium phytofermentans and E.coli with 200ug/ml erm
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameqQint
-
Insert Size (bp)3609
-
Mutationplasmid to make targeted insertions in Clostridium phytofermentans chromosome using group II intron
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site SmaI (destroyed during cloning)
- 3′ cloning site SmaI (destroyed during cloning)
- 5′ sequencing primer n/a
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Between Addgene sequence and author sequence, there is a 12 nucleotide mismatched region. It is in the intron sequence, which needs to be replaced in order to use the plasmid and should not be of concern.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pQint was a gift from George Church (Addgene plasmid # 25819 ; http://n2t.net/addgene:25819 ; RRID:Addgene_25819) -
For your References section:
Targeted gene inactivation in Clostridium phytofermentans shows that cellulose degradation requires the family 9 hydrolase Cphy3367. Tolonen AC, Chilaka AC, Church GM. Mol Microbiol. 2009 Dec . 74(6):1300-13. 10.1111/j.1365-2958.2009.06890.x PubMed 19775243