-
Depositing Lab
-
Publication
-
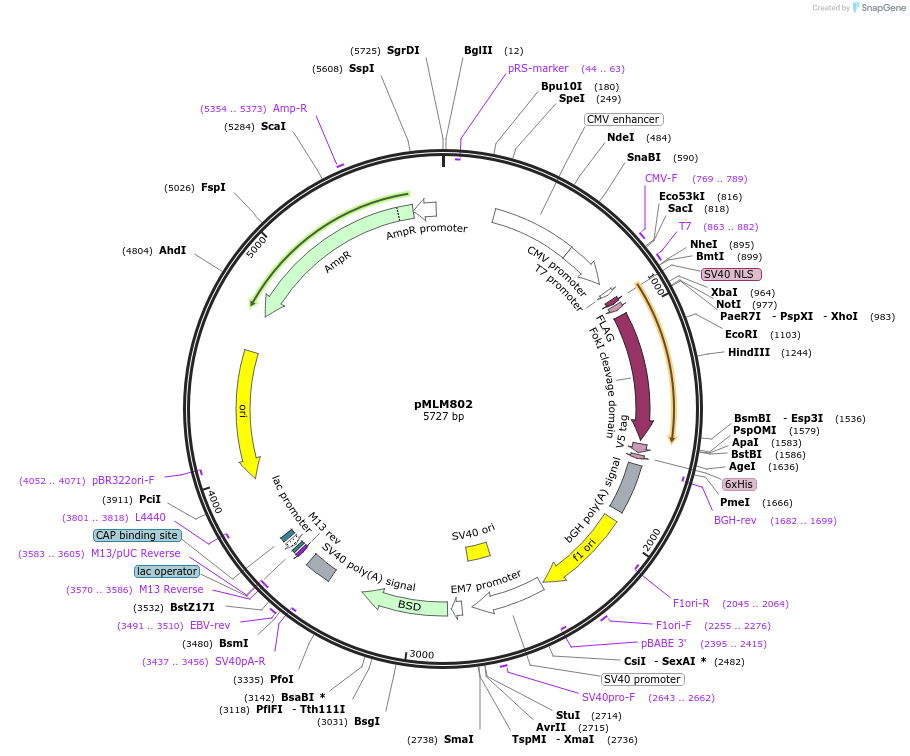
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 27203 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepMLM802
- Backbone size w/o insert (bp) 5727
-
Vector typeMammalian Expression ; zebrafish expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)XL1 Blue
-
Growth instructionsGrow in XL1 Blue
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameFokI endonuclease
-
Tag
/ Fusion Protein
- FLAG (N terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XbaI (not destroyed)
- 3′ cloning site NotI (not destroyed)
- 5′ sequencing primer CMV-F
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
For expression of a "-" heterodimeric FokI domain. (Miller et al., Nat Biotech 2007) in mammalian cells or zebrafish for ZFN targets with a 7bp spacer (Handel et al 2008, Mol Ther). To clone a zinc finger array into this plasmid, perform the following steps: (1) amplify the zinc finger array coding sequence from a B2H expression vector by PCR using primers OMM429 (5’-GATGACAAATCTAGACCCGGGGAGCG-3’) and OMM430 (5’-CTGCGGCCGCACCGGGTCCTGTGTGGGTTTTTAGGTG-3’), (2) digest the resulting PCR fragment with XbaI and NotI, and (3) clone the digested fragment into MLM802 vector backbone that has been digested with XbaI and NotI. The resulting plasmid will encode a ZFN that can be paired with a ZFN cloned into expression vector MLM800 to target a ZFN target site with a 7 bp spacer (Handel et al. 2008, Mol Ther.).
The difference between pMLM800/802 and pMLM290/292 is the length of the “spacer” sequence in the full ZFN target site. If the spacer is 7 bp, scientists should use pMLM800/802. If the spacer is 5 or 6 bps, scientists should use pMLM290/292.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pMLM802 was a gift from Keith Joung (Addgene plasmid # 27203 ; http://n2t.net/addgene:27203 ; RRID:Addgene_27203) -
For your References section:
An improved zinc-finger nuclease architecture for highly specific genome editing. Miller JC, Holmes MC, Wang J, Guschin DY, Lee YL, Rupniewski I, Beausejour CM, Waite AJ, Wang NS, Kim KA, Gregory PD, Pabo CO, Rebar EJ. Nat Biotechnol. 2007 Jul . 25(7):778-85. 10.1038/nbt1319 PubMed 17603475