-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 31222 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepAPIC-HIH
- Backbone size w/o insert (bp) 10193
-
Vector typeInsect Expression
Growth in Bacteria
-
Bacterial Resistance(s)Chloramphenicol and Ampicillin, 25 & 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)ccdB Survival
-
Growth instructionsccdB Survival™ 2 T1R Cells @ 37C.
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameCD4-tdTom
-
Alt nameCD4-tdTomato
-
SpeciesH. sapiens (human), D. melanogaster (fly); Aequorea victoria
-
Insert Size (bp)2271
-
MutationFusion of signal peptide of Drosophila Akh gene, human CD4 transmembrane domain, tdTomato, and Kir2.1 ER exit signal.
-
Entrez GeneCD4 (a.k.a. CD4mut, IMD79, Leu-3, OKT4D, T4)
-
Tags
/ Fusion Proteins
- CD4 (N terminal on insert)
- tdTomato (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XhoI (not destroyed)
- 3′ cloning site XbaI (not destroyed)
- 5′ sequencing primer ACTGCAACTACTGAAATCTGCC
- 3′ sequencing primer GGCGCACAGAAATGATTACAAC
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Gateway destination vector for cloning any enhancer to drive expression of membrane marker CD4-tdTom
The depositing laboratory recommends growing bacteria either on LB plates with 80 ug/ml Carbenicillin or in LB liquid media with 60 ug/ml Carbenicillin (a semi-synthetic ampicillin analog)
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
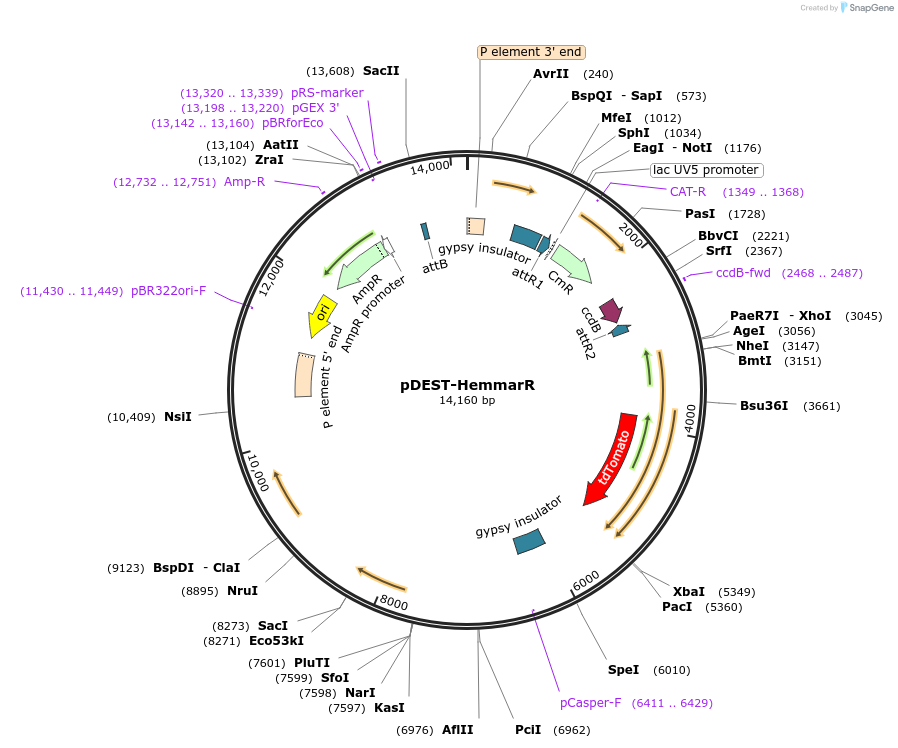
pDEST-HemmarR was a gift from Yuh-Nung Jan (Addgene plasmid # 31222 ; http://n2t.net/addgene:31222 ; RRID:Addgene_31222) -
For your References section:
Enhancer-driven membrane markers for analysis of nonautonomous mechanisms reveal neuron-glia interactions in Drosophila. Han C, Jan LY, Jan YN. Proc Natl Acad Sci U S A. 2011 Jun 7;108(23):9673-8. doi: 10.1073/pnas.1106386108. Epub 2011 May 23. 10.1073/pnas.1106386108 PubMed 21606367