-
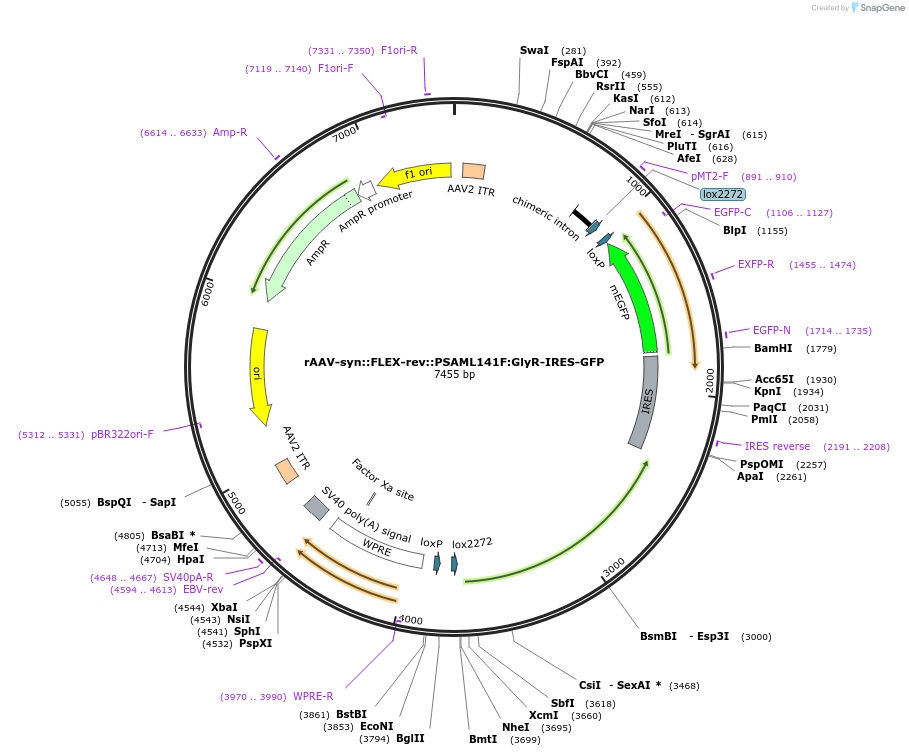
PurposeCre-dependent expression of PSAM-GlyR (L141F) neuronal inhibitor, with IRES-EGFP marker
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 32479 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backboneAAV2
- Backbone size w/o insert (bp) 5100
-
Modifications to backboneWPRE (Woodchuck hepatitis virus posttranscriptional regulatory element) before polyA sequence
-
Vector typeAAV, Cre/Lox ; Adeno-associated virus, FLEX switch
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)Stbl2
-
Copy numberLow Copy
Gene/Insert
-
Gene/Insert namePSAML141F-GlyR-IRES-GFP
-
SpeciesH. sapiens (human)
-
Insert Size (bp)2643
- Promoter Synapsin
-
Tag
/ Fusion Protein
- IRES-EGFP (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site SpeI blunt (destroyed during cloning)
- 3′ cloning site SpeI blunt (destroyed during cloning)
- 5′ sequencing primer gtaagtatcaaggttacaag
- 3′ sequencing primer ctgacaacgggccacaactc
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Note on the difference between PSAMY115F,L141F:GlyR and PSAML141F:GlyR silencers:
In Magnus et al, the PSAMY115F,L141F-GlyR channel was emphasized for silencing because this channel has the lowest acetylcholine responsiveness. However, PSAML141F-GlyR constructs, lacking the Y115F mutation, already have low ACh potency. The Y115F mutation also slightly increases the EC50 for the activator ligand, PSEM89S. Since PSAML141F-GlyR constructs already have very low acetylcholine potency, in many cases it may not be worth the slight reduction in PSEM potency. We have made both constructs available for researchers who may find that they have different requirements with respect to acetylcholine responsiveness.
Note on handling these rAAV constructs:
These rAAV vectors are prone to recombination. This is a well known issue with these rAAV vectors and is due to the inverted terminal repeats (ITRs) required for rAAV production. To minimize recombination, we propagate these plasmids in Stbl2 cells from Invitrogen. Also, to minimize recombination, cells should be cultured at 30 C.
Note that these cultures will grow slowly (20 h for minipreps). Better yields and culture times are obtained with 2xYT as the media. This is strongly recommended.
Because recombination may still happen, we do a panel of restriction digestions to assess whether the ITRs are in tact. Separate digestions with Sma1 and Msc1 should be performed. The expected patterns can be calculated from the attached sequence.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
rAAV-syn::FLEX-rev::PSAML141F:GlyR-IRES-GFP was a gift from Scott Sternson (Addgene plasmid # 32479 ; http://n2t.net/addgene:32479 ; RRID:Addgene_32479) -
For your References section:
Chemical and genetic engineering of selective ion channel-ligand interactions. Magnus CJ, Lee PH, Atasoy D, Su HH, Looger LL, Sternson SM. Science. 2011 Sep 2;333(6047):1292-6. 10.1126/science.1206606 PubMed 21885782