-
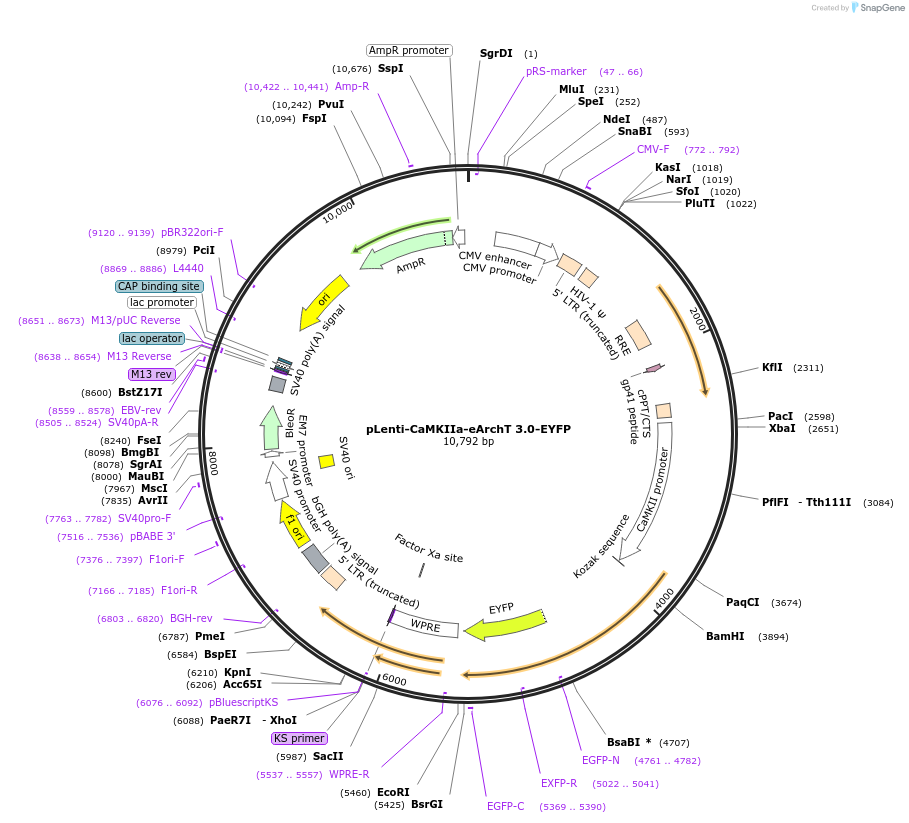
PurposeLentiviral expression of CaMKIIa-driven ArchT 3.0 for optical inhibition.
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 35513 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepLenti
- Backbone size w/o insert (bp) 9245
-
Modifications to backboneAddition of a CaMKIIa promoter and a WPRE
-
Vector typeMammalian Expression, Lentiviral
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)Stbl3
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameeArchT 3.0-EYFP
-
SpeciesHalorubrum sp. TP009
-
Insert Size (bp)1548
-
MutationEnhanced trafficking signal and ER export signal added
- Promoter CaMKIIa
-
Tag
/ Fusion Protein
- EYFP (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer AGTCCTGCAGTATTGTGTAT
- 3′ sequencing primer GCAATAGCATGATACAAAGG
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byArchT as reported by the Boyden Lab was synthesized by DNA 2.0 and fused with EYFP
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The enhanced trafficking signals allow for better membrane targeting in mammalian cells resulting in 3-5 fold increase in currents.
Please note that there may be several sequence discrepancies between depositor's reference sequence and Addgene's quality control sequence. These changes are in vector regions only and should not affect plasmid function.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pLenti-CaMKIIa-eArchT 3.0-EYFP was a gift from Karl Deisseroth (Addgene plasmid # 35513 ; http://n2t.net/addgene:35513 ; RRID:Addgene_35513) -
For your References section:
Principles for applying optogenetic tools derived from direct comparative analysis of microbial opsins. Mattis J, Tye KM, Ferenczi EA, Ramakrishnan C, O'Shea DJ, Prakash R, Gunaydin LA, Hyun M, Fenno LE, Gradinaru V, Yizhar O, Deisseroth K. Nat Methods. 2011 Dec 18;9(2):159-72. doi: 10.1038/nmeth.1808. 10.1038/nmeth.1808 PubMed 22179551