pcDNA-ZapCY2
(Plasmid
#36320)
-
Depositing Lab
-
Sequence Information
Full plasmid sequence is not available for this item.
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 36320 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepcDNA3.1(+)
-
Backbone manufacturerInvitrogen
- Backbone size w/o insert (bp) 5500
- Total vector size (bp) 7100
-
Vector typeMammalian Expression
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH10B
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameZapCY2 high affinity Zn(II) sensor
-
SpeciesSynthetic
-
Insert Size (bp)1626
-
Mutationsee comment
-
GenBank ID
- Promoter CMV
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BamHI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer T7 Forward
- 3′ sequencing primer BGH Reverse
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
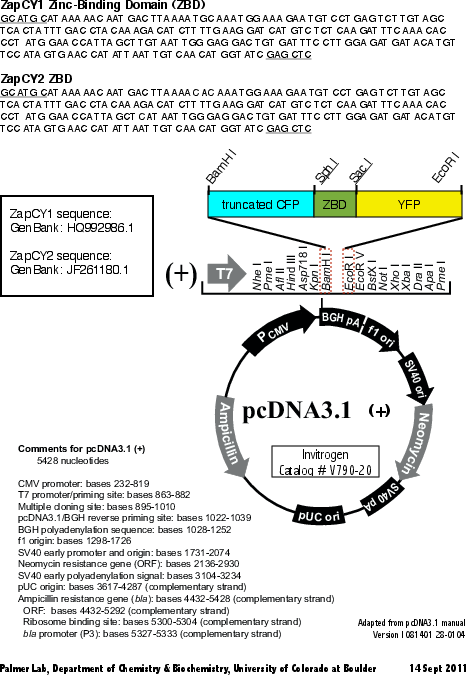
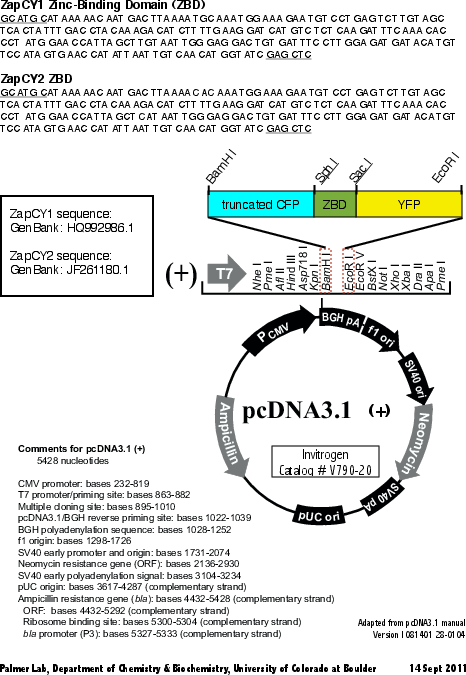
Second generation genetically encoded, ratiometric, fluorescent biosensor. Zn(II)-binding domain derived from the first 2 zinc fingers of Zap 1 from Saccharomyces cerevisiae. Contains truncated CFP and Citrine fluorescent proteins.
ZapCY2 is derived from ZapCY1. The first cysteine in each zinc finger is mutated to histidine.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pcDNA-ZapCY2 was a gift from Amy Palmer (Addgene plasmid # 36320 ; http://n2t.net/addgene:36320 ; RRID:Addgene_36320) -
For your References section:
Measuring steady-state and dynamic endoplasmic reticulum and Golgi Zn2+ with genetically encoded sensors. Qin Y, Dittmer PJ, Park JG, Jansen KB, Palmer AE. Proc Natl Acad Sci U S A. 2011 May 3;108(18):7351-6. Epub 2011 Apr 18. 10.1073/pnas.1015686108 PubMed 21502528
Map uploaded by the depositor.