-
Purpose(Empty Backbone)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 43831 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepRE107 (modified from pGP704)
-
Backbone manufacturerAddgene plasmid 43829
- Backbone size (bp) 5173
-
Modifications to backboneFrom pGP704: SacB1 from pUC58-sacB1 cloned into the EcoRI site. BamHI site in oriV converted to a ClaI site by partial digestion and Klenow treatment. From pRE107: Ampicillin resistance replaced with Tetracycline resistance (from pBR322) by blunted-BamHI digestion.
-
Vector typeBacterial Expression ; Bacterial allelic exchange vector with sacB1
-
Selectable markersSacB (sucrose sensitivity)
Growth in Bacteria
-
Bacterial Resistance(s)Tetracycline, 10 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)SM10 λpir
-
Growth instructionsContains lambda pir-dependent R6K replication origin; requires lambda pir-containing bacteria strain
-
Copy numberLow Copy
Cloning Information
- Cloning method Restriction Enzyme
- 5′ sequencing primer Tet-F (GGCAGGTAGATGACGACCAT)
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made bySM10 λpir bacterial strain and pGP704 plasmid backbone from John Mekalanos, Harvard Medical School, Boston, MA. SacB1 gene from plasmid pUC58-sacBI from Dennis Ohman, VCU Medical Center, Richmond, VA. TcR gene from pBR322, originally from Herbert Boyer, USCF, San Francisco, California.
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
The plasmids deposited here comprise a set of SacB1-dependent allelic exchange vectors improved from a previously described suicide vector, pGP704 (Miller and Mekalanos, 1988), by including a system to select for plasmid loss by recombination.
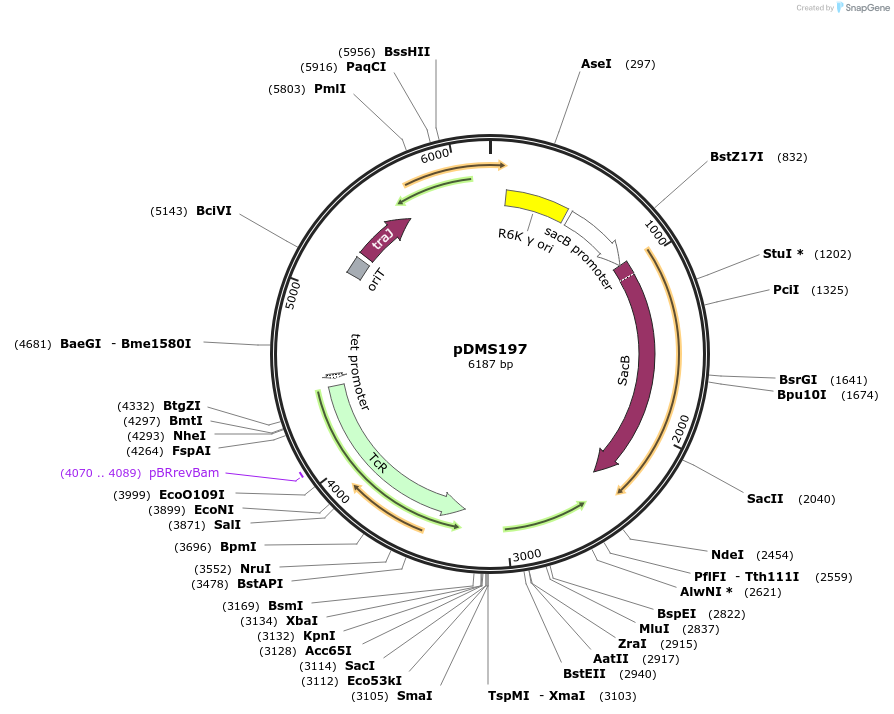
Plasmid pRE107 (Addgene plasmid #43829) was constructed by cloning the appropriate EcoRI fragment from pUC58-sacB1 (McIver et al., 1995) into pGP704 (see associated schematic image). The sacB1 allele is a modified variation of sacB, where unique restriction sites were removed by site directed mutagenesis (McIver et al., 1995). Additionally, the BamHI site in the R6K ori of the resulting plasmid was removed by partial BamHI digestion, blunting the resulting overhangs with PolIk (resulting in the formation of a ClaI site) and screening for its loss by restriction analysis.
To use this plasmid with bla fusions and increase the functionality of this system, the depositing laboratory replaced the ApR gene as follows.
The ApR gene flanked by the remaining BamHI sites was removed and replaced with either CmR, TcR or KmR resistance markers (see associated schematic image).
The TcR gene was cloned on a blunted EcoRI–StyI fragment from pBR322 into the blunted BamHI sites to give pDMS197. The BamHI site near the MCS was regenerated after cloning; the BamHI site near the 5' end of TcR/TetR was not.
The resulting plasmid together with others in this deposited series contain the conditional R6K ori, the origin of transfer (oriT) which allows conjugative transfer from permissive hosts, the sacB1 gene to provide negative selection and a MCS, along with a range of different AbR markers. Moreover, the use of Tc also allows an alternative negative selection using fusaric acid and chlortetracycline as described previously (Maloy and Nunn, 1981).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pDMS197 was a gift from Dieter Schifferli (Addgene plasmid # 43831 ; http://n2t.net/addgene:43831 ; RRID:Addgene_43831) -
For your References section:
Improved allelic exchange vectors and their use to analyze 987P fimbria gene expression. Edwards RA, Keller LH, Schifferli DM. Gene. 1998 Jan 30;207(2):149-57. 10.1016/S0378-1119(97)00619-7 PubMed 9511756