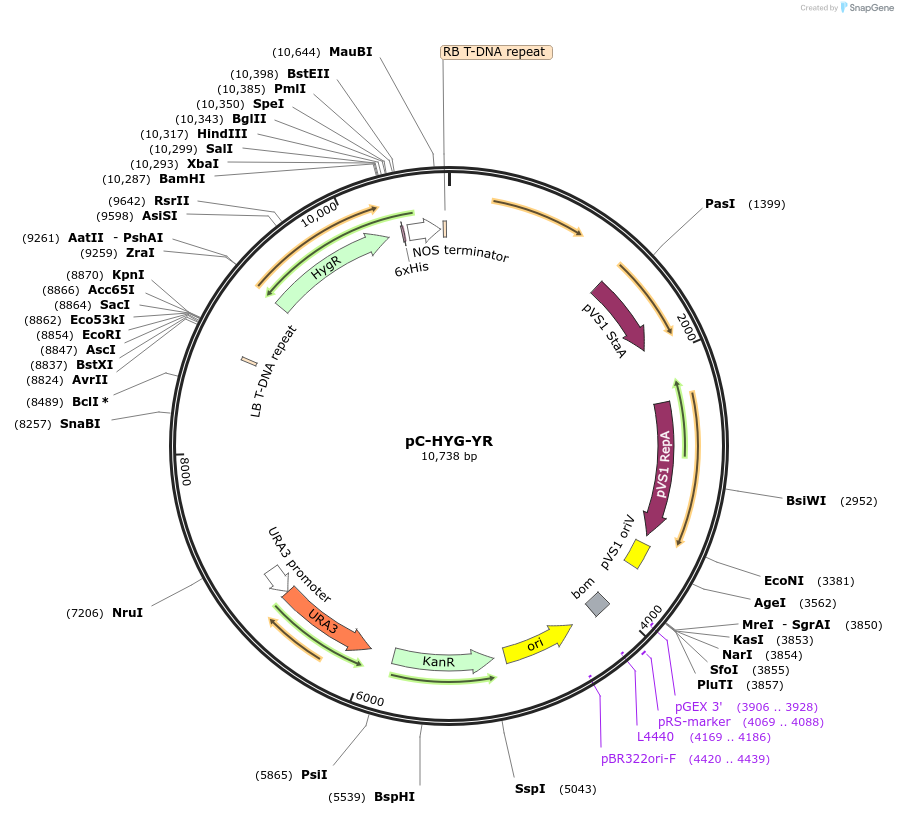
pC-HYG-YR
(Plasmid
#61765)
-
PurposeYeast recombinational cloning compatible Agrobacterium tumefaciens ternary vector containing hygromycin selectable marker on transfer DNA (TDNA).
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 61765 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepCAMBIA-0380
-
Backbone manufacturerCAMBIA
- Backbone size w/o insert (bp) 6812
- Total vector size (bp) 10737
-
Modifications to backboneThe Agrobacterium tumfaciens binary vector pCAMBIA-0380 was modified to construct pCHYG (Motteram et al, 2009) by inserting the hygromycin phosphotransferase (hph) gene under the control of Aspergillus nidulans trpC promoter on the TDNA region. A 2µ origin of replication and URA3 gene, from the yeast episomal plasmid YEp24 (Genbank L09156.1), was then inserted at the SacII restriction site in pCHYG to construct yeast recombinational cloning compatible ternary vector pC-HYG-YR.
-
Vector typeTernary vector for Agrobacterium mediated transformation of fungi
-
Selectable markersHygromycin, URA3
Growth in Bacteria
-
Bacterial Resistance(s)Kanamycin, 50 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert 1
-
Gene/Insert nameHygromycin
-
SpeciesSynthetic
-
GenBank ID
- Promoter trpC promoter
Cloning Information for Gene/Insert 1
- Cloning method Restriction Enzyme
- 5′ cloning site EcoRI (not destroyed)
- 3′ cloning site HindIII (not destroyed)
- 5′ sequencing primer LB-F1 (5'-gtggtgtaaacaaattgacgc-3')
- 3′ sequencing primer RB-F1 (5'-ggataaaccttttcacgccc-3')
- (Common Sequencing Primers)
Gene/Insert 2
-
Gene/Insert name2 micron origin or replication and URA3 gene for S. cerevisiae
-
SpeciesS. cerevisiae (budding yeast)
-
Insert Size (bp)2487
-
GenBank IDKC562906.1
Cloning Information for Gene/Insert 2
- Cloning method Restriction Enzyme
- 5′ cloning site SacII (not destroyed)
- 3′ cloning site SacII (not destroyed)
- 5′ sequencing primer ccgcggTTAGTTTTGCTGGCCGCATC
- 3′ sequencing primer ccgcggCGCTTCCACAAACATTGCTC
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe 2 micron origin of replication and URA3 gene were cloned from YEP24 (Genbank L09156.1)
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Jason Rudd (Rothamsted research UK) provided Agrobacterium tumfaciens vector pCHYG. For more information, see: Motteram et al, 2009. Molecular Plant-Microbe Interactions, 22, 790-799. https://www.ncbi.nlm.nih.gov/pubmed/19522561.
Restriction sites EcoRI and HindIII can be used to linearise the vector inside left (LB) and right (RB) borders of the TDNA. Note that the trpC promoter and hygromycin sequences are present within the EcoRI and HindII cloning sites.
See Supplementary Fig. S1. of associated publication (Sidhu et al., 2015) for schematic of single step assembly of gene deletion ternary vectors using yeast recombinational cloning.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pC-HYG-YR was a gift from Ken Haynes (Addgene plasmid # 61765 ; http://n2t.net/addgene:61765 ; RRID:Addgene_61765) -
For your References section:
Exploitation of sulfonylurea resistance marker and non-homologous end joining mutants for functional analysis in Zymoseptoria tritici. Sidhu YS, Cairns TC, Chaudhari YK, Usher J, Talbot NJ, Studholme DJ, Csukai M, Haynes K. Fungal Genet Biol. 2015 Jun;79:102-9. doi: 10.1016/j.fgb.2015.04.015. 10.1016/j.fgb.2015.04.015 PubMed 26092796