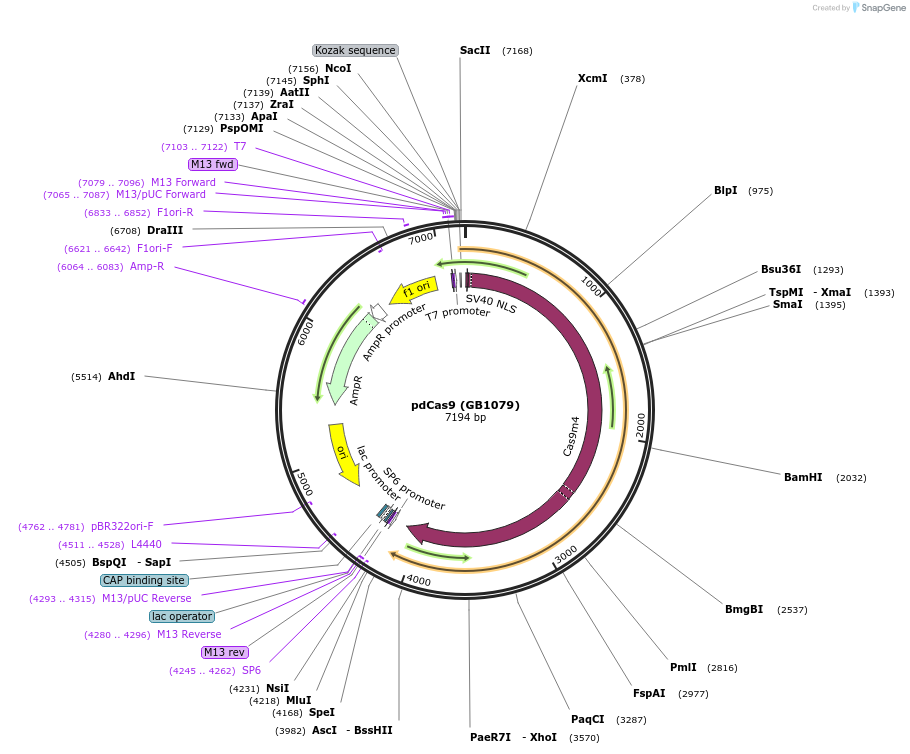
pdCas9 (GB1079)
(Plasmid
#75399)
-
PurposeProvides the human codon optimized CDS of Cas9 protein with mutated (D10A, H840A) and inactivated catalytic domains as a level 0 GoldenBraid part for C-terminal fusions
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 75399 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepUPD
-
Backbone manufacturerself-made; derived from pGEMT-Easy manufactured by Promega
- Backbone size w/o insert (bp) 3055
- Total vector size (bp) 7192
-
Vector typePlant Expression, CRISPR, Synthetic Biology
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameCas9 coding region with mutated (D10A, H840A) and inactivated catalytic domains (human codon optimised)
-
SpeciesStreptococcus pyogenes
-
Insert Size (bp)4137
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site BsmBI (destroyed during cloning)
- 3′ cloning site BsmBI (destroyed during cloning)
- 5′ sequencing primer T7
- 3′ sequencing primer SP6
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pdCas9 (GB1079) was a gift from Diego Orzaez (Addgene plasmid # 75399 ; http://n2t.net/addgene:75399 ; RRID:Addgene_75399) -
For your References section:
A modular toolbox for gRNA-Cas9 genome engineering in plants based on the GoldenBraid standard. Vazquez-Vilar M, Bernabe-Orts JM, Fernandez-Del-Carmen A, Ziarsolo P, Blanca J, Granell A, Orzaez D. Plant Methods. 2016 Feb 1;12:10. doi: 10.1186/s13007-016-0101-2. eCollection 2016. 101 [pii] PubMed 26839579