-
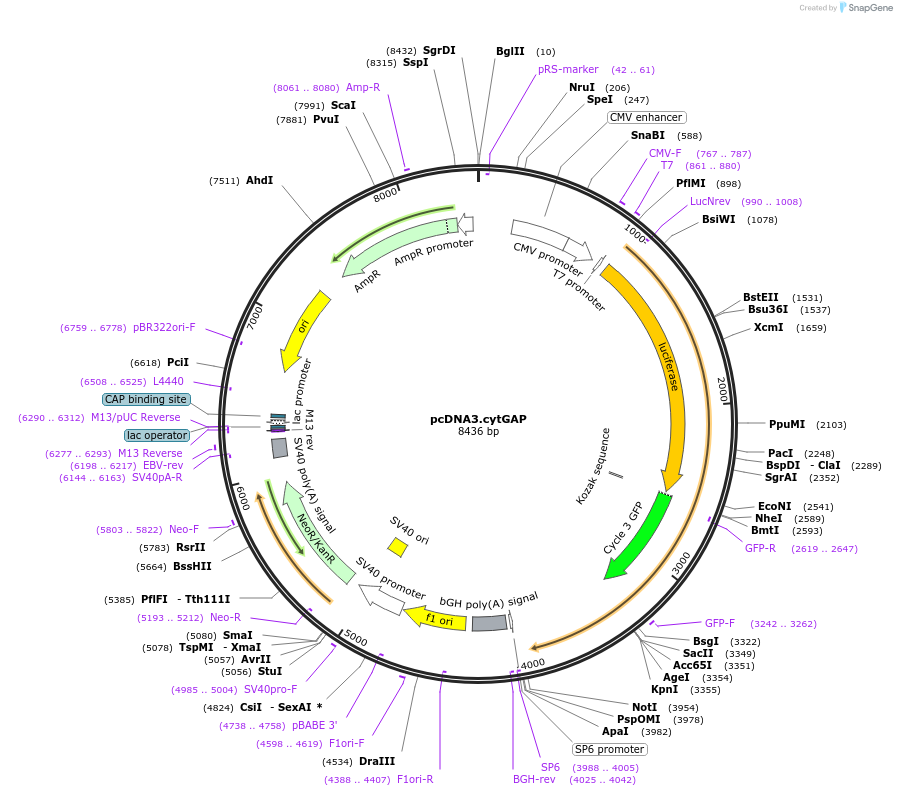
PurposeExpression of Ca2+ sensor GAP in the cytosol
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 78734 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepcDNA3
-
Vector typeMammalian Expression
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namecytGAP
-
SpeciesSynthetic
- Promoter CMV
-
Tag
/ Fusion Protein
- luciferase (N terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site unknown (unknown if destroyed)
- 3′ cloning site unknown (unknown if destroyed)
- 5′ sequencing primer CMV-F
- 3′ sequencing primer BGH-rev
- (Common Sequencing Primers)
Resource Information
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
GAP is a fusion of two Aequorea victoria proteins, GFP and apo-aequorin, with a linker in between. The excitation spectrum of GAP maintained the two peaks of wtGFP, at 403 and 470 nm, and it was sensitive to Ca2+.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pcDNA3.cytGAP was a gift from Teresa Alonso (Addgene plasmid # 78734 ; http://n2t.net/addgene:78734 ; RRID:Addgene_78734) -
For your References section:
GAP, an aequorin-based fluorescent indicator for imaging Ca2+ in organelles. Rodriguez-Garcia A, Rojo-Ruiz J, Navas-Navarro P, Aulestia FJ, Gallego-Sandin S, Garcia-Sancho J, Alonso MT. Proc Natl Acad Sci U S A. 2014 Feb 18;111(7):2584-9. doi: 10.1073/pnas.1316539111. Epub 2014 Feb 5. 10.1073/pnas.1316539111 PubMed 24501126