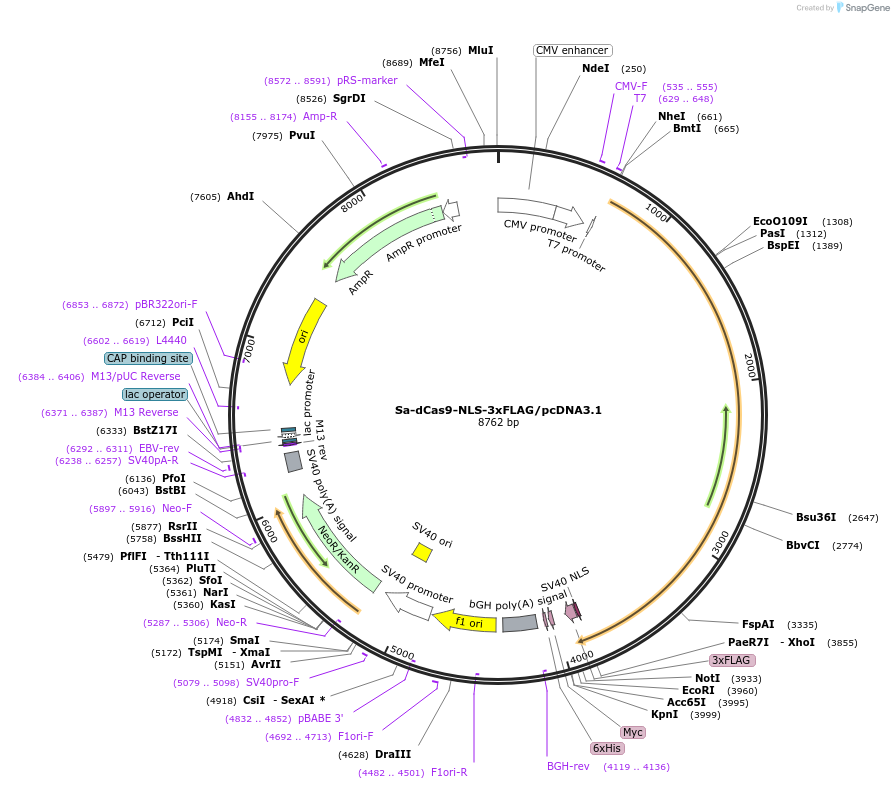
Sa-dCas9-NLS-3xFLAG/pcDNA3.1
(Plasmid
#98041)
-
PurposeExpresses Sa-dCas9-NLS-3xFLAG in mammalian cells for enChIP analysis to purify specific genomic regions of interest
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 98041 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepcDNA3.1
-
Backbone manufacturerInvitrogen
- Backbone size w/o insert (bp) 5500
- Total vector size (bp) 8767
-
Vector typeMammalian Expression, CRISPR
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)NEB Stable
-
Growth instructionsHigh frequency of recombination. Culture at 30°C.
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameSa-dCas9-NLS-3xFLAG
-
SpeciesSynthetic; Staphylococcus aureus
-
Insert Size (bp)3264
-
MutationD10A + H557A
- Promoter CMV
-
Tags
/ Fusion Proteins
- NLS (nuclear localization signal) (C terminal on insert)
- 3xFLAG tag (C terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site Nhe I (not destroyed)
- 3′ cloning site Not I (not destroyed)
- 5′ sequencing primer T7
- 3′ sequencing primer BGH Reverse
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byA portion of this plasmid was derived from a plasmid made by Keith Joung (Addgene plasmid 70703)
-
Article Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This plasmid can be used to isolate specific genomic regions of interest using a catalytically inactive Cas9 fused with a tag(s).
Purify: locus-specific chromatin immunoprecipitation (enChIP)
This system is compatible with BPK2660 (www.addgene.org/70709) from the Joung lab.
Additional information and protocols can be found at:
http://www.med.hirosaki-u.ac.jp/~bgb/iChIP_protocols/index_e.html
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
Sa-dCas9-NLS-3xFLAG/pcDNA3.1 was a gift from Hodaka Fujii (Addgene plasmid # 98041 ; http://n2t.net/addgene:98041 ; RRID:Addgene_98041) -
For your References section:
enChIP systems using different CRISPR orthologues and epitope tags. Fujita T, Yuno M, Fujii H. BMC Res Notes. 2018 Feb 27;11(1):154. doi: 10.1186/s13104-018-3262-4. 10.1186/s13104-018-3262-4 PubMed 29482606