Optogenetics Guide
The field of optogenetics combines optics and genetic engineering to manipulate cells and biomolecular processes with light. Light-activated tools generally fall into two main types: tools based on light-gated ion channels known as opsins (including chimeric receptors such as OptoXRs) or optical switches. Naturally-occuring opsins open and close when exposed to light and change the membrane potential, a behavior that can be harnessed for precise control of electrical activity in a target cell type. OptoXRs are chimeras of a G-protein coupled receptor (GPCR) and an opsin engineered to activate a specific signaling pathway when exposed to light. Optical switches harness light-responsive conformational changes in natural light-sensing domains to modulate a specific protein or process. Optogenetic tools are particularly useful for controlling neuronal activity, but they can be deployed to interrogate and control a wide variety of processes across cell types.
Check out our blog for practical tips, explore Addgene's Optogenetics Plasmid Collection, or read on to learn more about these tools.
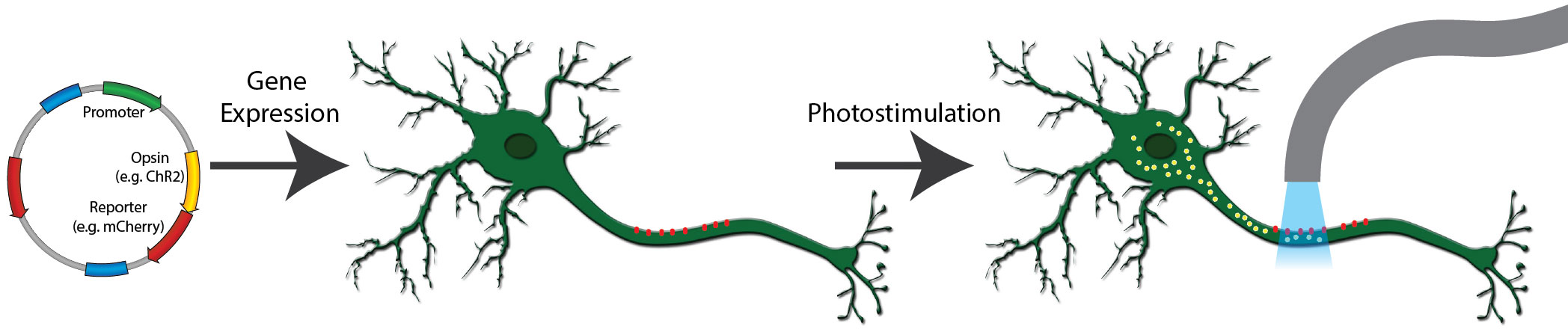
Figure 1: A typical optogenetics experiment. A channelrhodopsin fused to mCherry is expressed in neurons (red dots). When exposed to light of the correct wavelength, the channelrhodopsin pore opens and cations flow into the cell (yellow dots), activating the neuron.
Opsins
Opsins are light-gated ion channels or pumps that absorb light at specific wavelengths. Upon activation by light, these channels and pumps respond by opening or closing, which conducts the flow of ions into or out of the cell. Scientists have identified a variety of naturally occurring microbial opsins that respond to different wavelengths of light, like blue or yellow light. These various opsins also initiate different electrochemical responses, such as nonspecific cation influx vs. proton efflux. Researchers have used genetic engineering to improve these natural opsins by inducing point mutations to alter the absorption spectrum or adding trafficking signals to localize opsins to the cell membrane.
Microbial opsins, such as those described below, can be targeted and expressed in specific subsets of neurons, allowing precise spatiotemporal control of these neurons by turning on and off the light source. Optogenetics has been broadly applied to study the physiology of the brain and nervous system to better map and understand neuronal circuits. Optogenetic tools have also been used to trigger and study certain behavioral responses in model organisms like mice, zebrafish, and Drosophila. These tools have been instrumental in neurological disorder research, helping scientists to better understand Parkinson's disease, depression, drug addiction, and more.
Next, let's explore some commonly used microbial opsins, including a brief overview of the wild-type and some examples of relevant variants.
Channelrhodopsins
Channelrhodopsins are a foundational optogenetics tool. They typically allow the fast depolarization of neurons upon exposure to light through direct stimulation of ion channels. Naturally occurring channelrhodopsins were discovered in the green algae Chlamydomonas reinhardtii. Channelrhodopsin-1 (ChR1) is excited by blue light and permits nonspecific cation influx into the cell when stimulated. Channelrhodopsin-2 (ChR2), the first widely adopted optogenetic tool, is also a blue light activated cation channel. ChR2 is preferred over ChR1 because ChR2 has higher conductance at physiological pH and traffics well to the membrane.
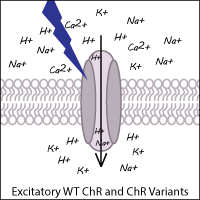
The optogenetic toolbox has been expanded by scientists through both the identification of novel ChRs from other algal species and the development of synthetic variants to enhance the functionality of ChR. Examples of ChRs from other species include CoChR (from Chloromonas oogama) and SdChR (from Scherffelia dubia). Synthetic variants have been created via genetic point mutations, codon optimization, and chimeric fusion of domains from two different ChRs. These ChR variants still function as light-gated cation channels resulting in excitation (depolarization) of the neuron.
Inhibitory (hyperpolarizing) ChR variants
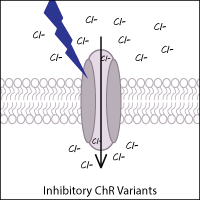
Anion-conducting ChR variants that inhibit neurons have been created and identified in other species. By acting as light-gated chloride channels, these variants result in the hyperpolarization of neurons. Examples of anion channel variants from other species include GtACR1 and GtACR2 (from the cryptophyte Guillardia theta).
Halorhodopsins
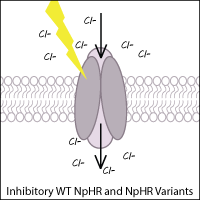
Halorhodopsins are light-gated, inward-directed chloride pumps isolated from halobacteria. Wild-type halorhodopsin, known as NpHR (from Natronomonas pharaoni), causes hyperpolarization (inhibition) of the cell when triggered with yellow light, thus inhibiting function of the neuron.
Archaerhodopsins
Archaerhodopsin-3 (Arch) from Halorubrum sodomense and its variants are commonly used to inhibit neurons in optogenetic experiments. Arch is a light-activated outward proton pump that hyperpolarizes (inhibits) the cell when triggered by green-yellow light.
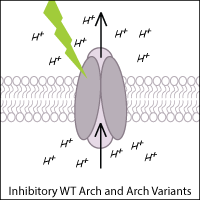
Leptosphaeria Rhodopsin
Leptosphaeria rhodopsin (Mac) is a blue-green light-activated proton pump derived from the fungus Leptosphaeria maculans. Mac and its variants allow for inhibition of neurons using blue-green light.
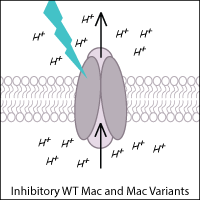
Kalium Channelrhodopsins
Potassium channelrhodopsins are versatile and robust optogenetic inhibitors that act by hyperpolarizing cells via high K+ efflux. Synthetic light-regulated K+ channels include BLINK1 and BLINK2, while natural KCRs have been identified in Hyphochytrium catenoides (HcKCRs) and Wobblia lunata (WiChR).
Other Opsin-based Tools
OptoXRs
GPCRs activate signaling cascades driving a variety of physiological processes in response to specific stimuli. In fact, the animal GPCRs responsible for vision found in rod and cone photoreceptor cells are light-sensitive rhodopsins. Expressing these natural proteins in non-native cell types enables optogenetic activation of the major heterotrimeric G-protein signaling pathways in animal cells, but they can be difficult to control precisely due to the complexity of these signaling networks and other limitations like bleaching. A more selective way to stimulate the activity of a particular GPCR is to engineer a chimeric protein consisting of an opsin and the GPCR of interest (referred to as an optoGPCR or OptoXR). Combining the extracellular domain of an endogenous light-sensitive receptor, like mammalian rhodopsin, with the intracellular domain of an endogenous GPCR, such as adrenergic receptor, makes a receptor that activates the native signaling cascade upon light exposure. OptoXRs and OptoGPCRs enable a broad range of pathways to be activated and explored, especially in non-excitable or non-neuronal cell types.
Luminopsins
Luminopsins (LMOs), or luminescent opsins, are opto-chemogenetic tools consisting of a light-emitting luciferase fused to a light-sensing element such as an opsin. When the luciferase substrate is added, the luciferase enzyme generates light that activates the opsin. While LMOs can also be activated by standard optogenetics workflows (such as laser or LED illumination), systemic injection of a small molecule is less invasive than implantation of fiber optics, making LMOs a versatile option.
Learn more about LMOs and other chemogenetic systems in our Chemogenetics Guide.
Glossary of Common Microbial Opsins and Variants
- Channelrhodopsins from C. reinhardtii
- Other Channelrhodopsins
- Anion Channels
- Halorhodopsins
- Archaerhodopsins
- Leptosphaeria Rhodopsins
These lists are not exhaustive — check out our Optogenetics Plasmid Collection to find plasmids for these and other tools for your experiments.
Channelrhodopsins from Chlamydomonas reinhardtii
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| ChR2 | Widely used light-gated cation channel from Chlamydomonas reinhardtii (CrChR2) | 470 |
| ChR2/H134R | Widely used variant with larger photocurrent compared to CrChR2 | 450 |
| ChETA | E123T mutation; faster kinetics but reduced photocurrent amplitude | 490 |
| ChR/T159C | T159C mutation; displays increased photocurrents | 470 |
| SFO/SSFO | Step Function Opsins and Stabilized Step Function Opsins; mutations at C128 and D156 of CrChR delay the closing of the channel | 470 (activation)
590 (inactivation) |
| ReaChR | Red-activatable variant of CrChR2 | 590 |
Other Channelrhodopsins
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| VChR1 | Red-shifted channelrhodopsin from Volvox carteri | 570 |
| Chronos | High-speed, red-shifted channel from Stigeoclonium helveticum | 500 |
| Chrimson, ChrimsonR | Red-shifted channel from Chlamydomonas noctigama; ChrimsonR carries K176R mutation with increased on/off kinetics | 590 |
| ChRmine | Red-shifted channelrhodopsin from Tiarina fusus; variants include rsChRmine (further red-shifted), hsChRmine (high speed), and frChRmine (combined high speed and more red-shifted); good response with 585 nm or 650 nm stimulation | ~520 (ChRmine, hsChRmine)
~585 (rsChRmine, frChRmine) |
| PsChR2 | High-efficiency blue-shifted channelrhodopsin from Platymonas subcordiformis | 445 |
| CoChR | Channelrhodopsin from Chloromonas oogama; high-efficacy soma-targeted variant soCoChR | 470 |
| CheRiff | Blue-shifted channelrhodopsin from Scherffelia dubia | 460 |
| C1C2 | ChR1-ChR2 chimera | 470 |
| ChIEF, ChEF, ChD | Engineered chimeras of ChR1 and ChR2; increased photocurrent amplitude | 450–470 |
| C1V1(t/t) | ChR1-VChR1 chimera with E122T and E162T mutations; increased photocurrent amplitude, on/off kinetics, and red-shifted peak action spectra | 540 |
| HcKCR | Kalium channelrhodopsins, light-gated K+ channels from Hyphochytrium catenoides | 540 (HcKCR1)
490 (HcKCR2) |
| WiChR | Kalium channelrhodopsin, light-gated K+ channel from Wobblia lunata | 490 |
Anion Channels
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| GtACR | Chloride-conducting channel from Guillardia theta, used to silence neuronal activity | 515 (GtACR1)
470 (GtACR2) |
| iC++ | Artificial anion conducting channelrhodopsin, used to silence neuronal activity | 488 |
| iChloC | Chloride-conducting channel, CrChR2 with mutations E90R, D156N, T159C, CE83Q, and E101S; increased photocurrent amplitude | 465 |
| SwiChR | Step-waveform inhibitory ChR, chloride channel derived from C1C2; SwiChRCA includes mutation C128A with increased photocurrent amplitude | 475 |
| PsChR1 | Red-shifted chloride-conducting channel from Proteomonas sulcata | 540 |
| Phobos | Blue-shifted iC++ variant; increased photocurrent amplitude | 467 |
| Aurora | Red-shifted anion-selective ReaChR variant; increased photocurrent amplitude | 517 |
Halorhodopsins
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| Jaws | Red-shifted, light-driven inward chloride pump from Haloarcula (Halobacterium) salinarum (strain Shark), used to silence neuronal activity | 632 |
| Halo/NpHR | Light-driven inward chloride pump from Natronomonas pharaonis, used to silence neuronal activity; Halo is human codon-optimized | 589 |
| eNpHR, eNpHR2.0, eNpHR3.0 | eNpHR variants with increased photocurrent amplitude; eNpHR3.0 includes trafficking signal from Kir2.1 and ER export signal for improved membrane targeting | 589 |
Archaerhodopsins
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| Arch | Light-activated outward proton pump from Halorubrum sodomense | 566 |
| eArch 3.0 | Arch with addition of trafficking signal from Kir2.1; increased photocurrent amplitude | 566 |
| ArchT | Light-activated outward proton pump from Halorubrum strain TP009; improved light sensitivity over Arch | 566 |
| eArchT 3.0 | ArchT with addition of trafficking signal from Kir2.1; increased photocurrent amplitude | 566 |
Leptosphaeria Rhodopsins
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| Mac | Light-activated outward proton pump from Leptosphaeria maculans | 540 |
| eMac 3.0 | Mac with addition of trafficking signal from Kir2.1; increased photocurrent amplitude | 540 |
Optical Switches
Researchers have harnessed a variety of plant and bacterial photoreceptors to control protein systems with light. These "photoswitchable" proteins offer exquisite spatial and temporal control of protein activity. Phytochromes, cryptochromes, light oxygen voltage (LOV), and other photosensitive domains have been adapted to control many protein functions, including kinase activity, localization, gene editing, transcriptional activation, and more. In addition to systems based on these natural photosensing domains, a growing number of synthetic or engineered systems have been developed.
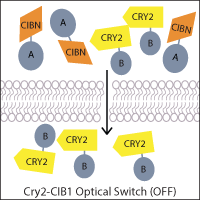
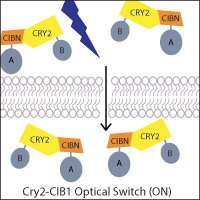
One of the most widely-used systems is the A. thaliana cryptochrome Cry2 and its binding partner cryptochrome-interacting basic helix-loop-helix CIB1. Blue light induces a conformational change in Cry2 that allows CIB1 binding, which can be utilized to control the localization of a protein of interest. For example, a genetically-encoded nuclear Cry2 fusion can recruit a CIB1-fused protein to the nucleus upon light activation. Alternatively, if one fragment of a protein of interest, such as Cre recombinase, is fused to Cry2 and the other fragment to CIB1, light-stimulated heterodimerization can reconstitute the protein. This principle has also been applied to generate two-part transcription factors, where Cry2 and CIB1 are fused to a transcriptional activation domain and DNA binding domain, allowing light to activate transcription.
Another popular strategy takes advantage of the LOV domains found in proteins used by many plants, microalgae, fungi, and bacteria to sense environmental conditions. LOV domains from a variety of species (Avena sativa, Botrytis cinerea, Rhodobacter sphaeroides, and others) have been fused to different effector domains of proteins to generate novel engineered light-controlled molecules. Phytochromes and LOV domains undergo light-induced changes in protein conformation, controlling activity of a fused protein through allosteric regulation. Alternatively, two-component systems like iLID or PhyB/PIF allow a user to reconstitute fragments of a protein of interest or localization signal to control protein activity or location. Researchers continue to engineer new tools and improve upon previous tools through engineering, design, and mutagenesis.
Glossary of Common Optical Switches
These lists are not exhaustive — check out our Optogenetics Plasmid Collection to find plasmids for these and other tools for your experiments.
UV Receptors
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| UVR8 | Plant photoreceptor protein that forms photolabile homodimers with slow reversal kinetics and a UV-B absorption profile | 282 |
BLUF Domains
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| bPAC | Light-activated adenylyl cyclase from Beggiatoa sp. for light-induced cAMP modulation, based on BLUF (blue light-utilizing FAD) domains | 441 |
| PixD-PixE | PixD and PixE associate into large multi-subunit complexes in the dark and reversibly dissociate upon blue light stimulation | 450 |
Cryptochromes
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| CRY2 and CIB1 | Blue-light–mediated induction of protein interactions based on A. thaliana Cryptochrome 2 and CIB1 | 450 |
| CRY2olig, CRY2clust | CRY2 variants that undergo homo-oligomerization; can be used to disrupt target protein activity or localization | 450 |
| LARIAT | Light-activated reversible inhibition by assembled trap; inhibits protein function by reversibly sequestering targets into large protein clusters through CRY2-CIB1 oligomerization | 450 |
| OPTOSTIM | PHR domain of CRY2 from A. thaliana is fused to truncated forms of cytosolic STIM1 to control calcium channels | ~488 |
LOV Domains
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| AsLOV2 | Light-activated conformational change in A. sativa LOV2 can be coupled to control a wide variety of target proteins | 450 |
| EL222 | Homodimerizes and binds to DNA upon blue light illumination, reverses in dark | 450–495 |
| iLID | Improved light-inducible dimers: SsrA peptide is embedded in the LOV2 domain, while its binding partner SspB is expressed separately; blue light activation allows SsrA to bind with SspB, each of which may be fused to a protein of interest | 450 |
| LAD | Light-activated dimerization, using proteins GIGANTEA (GI) and the LOV domain of FKF1 | ~450–495 |
| LITEZ | Light-inducible transcription using engineered zinc finger proteins, using proteins GIGANTEA (GI) and the LOV domain of FKF1 | ~450–495 |
| LINUS | Light-inducible nuclear localization signal based on the LOV2 domain of A. sativa phototropin 1 | ~450–495 |
| LOVTRAP | Zdk binds to the LOV domain in the dark; attaching one member of the Zdk/LOV2 pair to a target protein and the other to the membrane sequesters the target protein, to be released only upon illumination | 450 |
| miniSOG | Singlet oxygen generator; can be used to disrupt cellular function or ablate cells local generation of reactive oxygen species or to introduce contrast for electron microscopy | 450 |
| pDawn/pDusk | Use pDusk for blue-light-repressible or pDawn for light-activated gene expression in E. coli; YF1 (a synthetic kinase incorporating a LOV domain) phosphorylates the transcriptional activator FixJ, driving gene expression | 470 |
| TULIP | Tunable light-inducible dimerization tags; proteins fused to LOVpep reversibly dimerize with PDZ tags (cpPDZ, ePDZb, or ePDZb1) upon blue light excitation | 450 |
| VVD | Homodimerizes upon blue light, slowly reversible; VVDfast is also available | 450 |
Phytochromes
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| PhyB/PIF | Light-controlled, reversible dimerization of Phytochrome Interacting Factor 3 or 6 (PIF3 or PIF6) with PhyB | 660 |
| PhyA/FYH1 and PhyA/FHL | Light-controlled reversible dimerization of PhyA with FHY1 or FHL; PhyA and FHY1 also known as REDMAP | 660 |
| BphP1/PpsR2 | Bacterial phytochrome BphP1 and its natural partner PpsR2 from Rhodopseudomonas palustris bacteria | 740–780 |
| iLight | Near-infrared inducible dimerization based on the photosensory core module of phytochrome-activated diguanylyl cyclase from Idiomarina sp (IsPadC-PCM) | 660 |
Fluorescent Protein Domains
| Name | Description | Peak excitation λ (nm) |
|---|---|---|
| Dronpa | Mutant Dronpa domains associate in the dark, inhibiting a fused protein, and dissociate upon illumination, allowing activity; Dronpa145N homotetramerizes, Dronpa145K and Dronpa145N dimerize with each other. | ~500 (dissociate)
~400 (reassociate) |
| PhoCl | Photocleavable protein engineered from a green-to-red photoconvertible fluorescent protein; irreversible | 380 |
Plan Your Optogenetics Experiment
Before starting, check the Methods section of the original report or recent publications using your chosen tool for more details. When designing your own optogenetics experiment, here are some key factors to consider:
Activation or Inhibition?
First things first: do you want to activate or silence the neurons in your experiment? Depending on your answer, you'd pick an excitatory or inhibitory opsin, respectively.
For experiments targeting other cell types or pathways using optical switches, you may have a lot more options to choose from. You could choose an existing tool or build your own, using optical components that activate or inhibit the target of interest by disrupting its localization or conformation (or that of its interacting partner).
Color of Illumination Light
Different tools require different activation wavelengths, which may have advantages or disadvantages depending on the equipment available or your experimental design. Red light exhibits better tissue penetrance, which may allow you to place the optic fiber outside of the brain, rendering the experimental procedure less invasive. It's also possible to combine multiple opsins with different activation wavelengths in the same experiment. For example, you could use different colors of light to activate or silence the same neuronal population, or activate different neural populations at distinct times. Or you might want to use your optogenetic tool alongside a biosensor that you image in a different color channel.
Time and kinetics
Temporal precision is key in optogenetic experiments. Your experimental design will determine whether you'll need short or long periods of activation/inactivation periods. These can range from milliseconds with hChR2 to seconds or minutes with stable step-function opsins (SSFOs).
Delivery
Two factors determine which neuronal population is manipulated in a given experiment: the expression of the opsin and the area that is being illuminated. There are several different ways to control opsin expression. Generally, the most robust and stable expression is achieved in a transgenic mouse line, such as for example the VGAT-ChR2 mouse, where ChR2 is expressed under the control of the vesicular gamma aminobutyric acid (GABA) transporter (VGAT) and ChR2 is thus expressed in all GABAergic neurons. In this case, the subpopulation of GABAergic neurons being activated by the light is controlled via the placement of the optic fiber.
Using viral vectors for opsin delivery results in more localized expression of the opsin. Depending on the virus and promoter system used, there is an incubation time (days to weeks) until peak expression of the opsin is reached. This approach is especially powerful when combined with site-specific recombinase systems like Cre-lox, which allows you to express an opsin only in a genetically defined subpopulation of cells at the viral injection site, rather than in all cells. For example, using a viral vector with a floxed opsin in a VGAT-Cre animal will result in expression of the opsin only in inhibitory neurons near the injection site.
Ready to start? Explore Addgene's Optogenetics Plasmid Collection to find the plasmids you need.
Looking for viral preps? We also offer ready-to-use AAV preparations of many optogenetics plasmids and AAV Packaged on Request for many more.
References and Additional Resources
- OptoBase (Link opens in a new window) annotated database of optical switches and their properties, from the Weber Lab
- Optogenetics Resources (Link opens in a new window) from the Boyden Lab
- Optogenetics Resources (Link opens in a new window) from the Deisseroth Lab
Berglund, K., Birkner, E., Augustine, G. J., & Hochgeschwender, U. (2013). Light-emitting channelrhodopsins for combined optogenetic and chemical-genetic control of neurons. PLoS ONE, 8(3), e59759. https://doi.org/10.1371/journal.pone.0059759 PMID: 23544095
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., & Deisseroth, K. (2005). Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci, 8(9), 1263–1268. https://doi.org/10.1038/nn1525 PMID: 16116447
Chuong, A. S., Miri, M. L., Busskamp, V., Matthews, G. A., Acker, L. C., Sørensen, A. T., Young, A., Klapoetke, N. C., Henninger, M. A., Kodandaramaiah, S. B., Ogawa, M., Ramanlal, S. B., Bandler, R. C., Allen, B. D., Forest, C. R., Chow, B. Y., Han, X., Lin, Y., Tye, K. M., Roska, B., Cardin, J. A., Boyden, E. S. (2014). Noninvasive optical inhibition with a red-shifted microbial rhodopsin. Nat Neurosci, 17(8), 1123–1129. https://doi.org/10.1038/nn.3752 PMID: 24997763
Deisseroth, K., Feng, G., Majewska, A. K., Miesenböck, G., Ting, A., & Schnitzer, M. J. (2006). Next-generation optical technologies for illuminating genetically targeted brain circuits. J Neurosci, 26(41), 10380–10386. https://doi.org/10.1523/JNEUROSCI.3863-06.2006 PMID: 17035522
Duan, X., Zhu, M., & Gao, S. (2025). Two Decades of Optogenetic Tools: A Retrospective and a Look Ahead. Adv Genet, 6(3), e00021. https://doi.org/10.1002/ggn2.202500021 PMID: 41036479
Emiliani, V., Entcheva, E., Hedrich, R., Hegemann, P., Konrad, K. R., Lüscher, C., Mahn, M., Pan, Z. H., Sims, R. R., Vierock, J., & Yizhar, O. (2022). Optogenetics for light control of biological systems. Nat Rev Methods Primers, 2, 55. https://doi.org/10.1038/s43586-022-00136-4 PMID: 37933248
Gradinaru, V., Zhang, F., Ramakrishnan, C., Mattis, J., Prakash, R., Diester, I., Goshen, I., Thompson, K. R., & Deisseroth, K. (2010). Molecular and cellular approaches for diversifying and extending optogenetics. Cell, 141(1), 154–165. https://doi.org/10.1016/j.cell.2010.02.037 PMID: 20303157
Han, X., & Boyden, E. S. (2007). Multiple-color optical activation, silencing, and desynchronization of neural activity, with single-spike temporal resolution. PLoS ONE, 2(3), e299. https://doi.org/10.1371/journal.pone.0000299 PMID: 17375185
Klapoetke, N. C., Murata, Y., Kim, S. S., Pulver, S. R., Birdsey-Benson, A., Cho, Y. K., Morimoto, T. K., Chuong, A. S., Carpenter, E. J., Tian, Z., Wang, J., Xie, Y., Yan, Z., Zhang, Y., Chow, B. Y., Surek, B., Melkonian, M., Jayaraman, V., Constantine-Paton, M., Wong, G. K., Boyden, E. S. (2014). Independent optical excitation of distinct neural populations. Nat Methods, 11(3), 338–346. https://doi.org/10.1038/nmeth.2836 PMID: 24509633
Lin J. Y. (2011). A user's guide to channelrhodopsin variants: features, limitations and future developments. Exp Physiol, 96(1), 19–25. https://doi.org/10.1113/expphysiol.2009.051961 PMID: 20621963
Mattis, J., Tye, K. M., Ferenczi, E. A., Ramakrishnan, C., O'Shea, D. J., Prakash, R., Gunaydin, L. A., Hyun, M., Fenno, L. E., Gradinaru, V., Yizhar, O., & Deisseroth, K. (2011). Principles for applying optogenetic tools derived from direct comparative analysis of microbial opsins. Nat Methods, 9(2), 159–172. https://doi.org/10.1038/nmeth.1808 PMID: 22179551
Piatkevich, K. D., & Boyden, E. S. (2023). Optogenetic control of neural activity: The biophysics of microbial rhodopsins in neuroscience. Q Rev Biophys, 57, e1. https://doi.org/10.1017/S0033583523000033 PMID: 37831008
Wietek, J., Beltramo, R., Scanziani, M., Hegemann, P., Oertner, T. G., & Wiegert, J. S. (2015). An improved chloride-conducting channelrhodopsin for light-induced inhibition of neuronal activity in vivo. Sci Rep, 5, 14807. https://doi.org/10.1038/srep14807 PMID: 26443033
Wietek, J., & Prigge, M. (2016). Enhancing Channelrhodopsins: An Overview. Methods Mol Biol, 1408, 141–165. https://doi.org/10.1007/978-1-4939-3512-3_10 PMID: 26965121
Yizhar, O., Fenno, L. E., Prigge, M., Schneider, F., Davidson, T. J., O'Shea, D. J., Sohal, V. S., Goshen, I., Finkelstein, J., Paz, J. T., Stehfest, K., Fudim, R., Ramakrishnan, C., Huguenard, J. R., Hegemann, P., & Deisseroth, K. (2011). Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature, 477(7363), 171–178. https://doi.org/10.1038/nature10360 PMID: 21796121
Yizhar, O., Fenno, L., Zhang, F., Hegemann, P., & Diesseroth, K. (2011). Microbial opsins: a family of single-component tools for optical control of neural activity. Cold Spring Harb Protoc, 2011(3), top102. https://doi.org/10.1101/pdb.top102 PMID: 21363959
Zhang, F., Prigge, M., Beyrière, F., Tsunoda, S. P., Mattis, J., Yizhar, O., Hegemann, P., & Deisseroth, K. (2008). Red-shifted optogenetic excitation: a tool for fast neural control derived from Volvox carteri. Nat Neurosci, 11(6), 631–633. https://doi.org/10.1038/nn.2120 PMID: 18432196
Credits
- Contributing Authors
- Written and reviewed by the Scientific Curation team at Addgene.
- Last Updated
- Content last reviewed on 23 Feburary 2026.


