We narrowed to 2,874 results for: Star
-
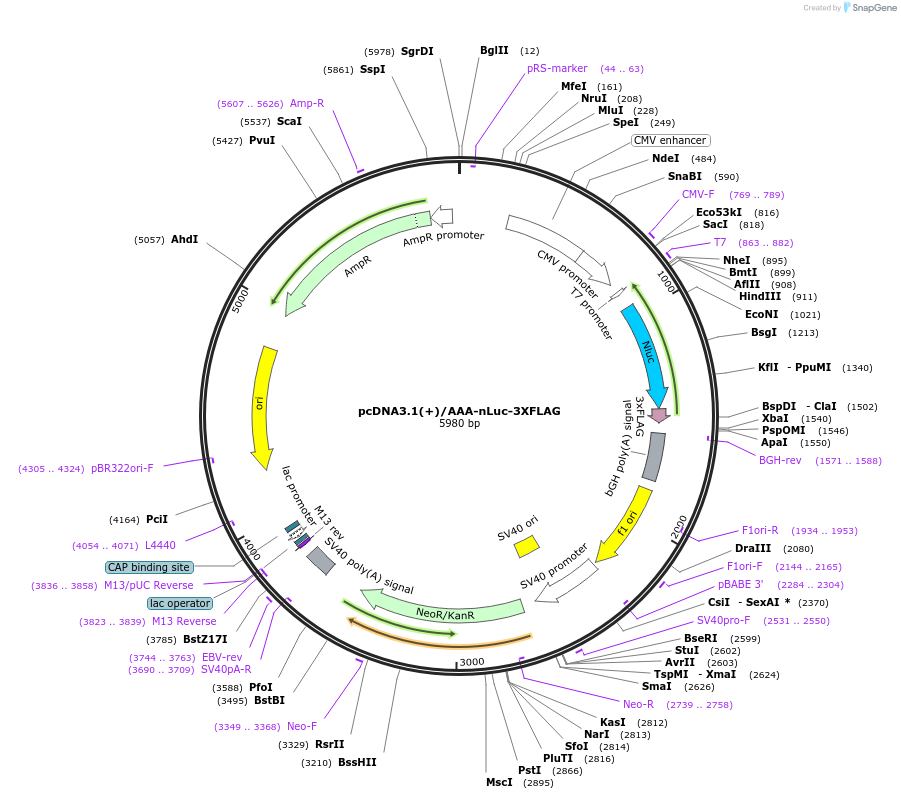
Plasmid#127306PurposeExpress 3xFLAG tagged nanoLuciferase (nLuc) from AAA start codon (Negative control plasmid)DepositorInsertnanoLuciferase
Tags3xFLAGExpressionMammalianPromoterCMVAvailable SinceJune 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
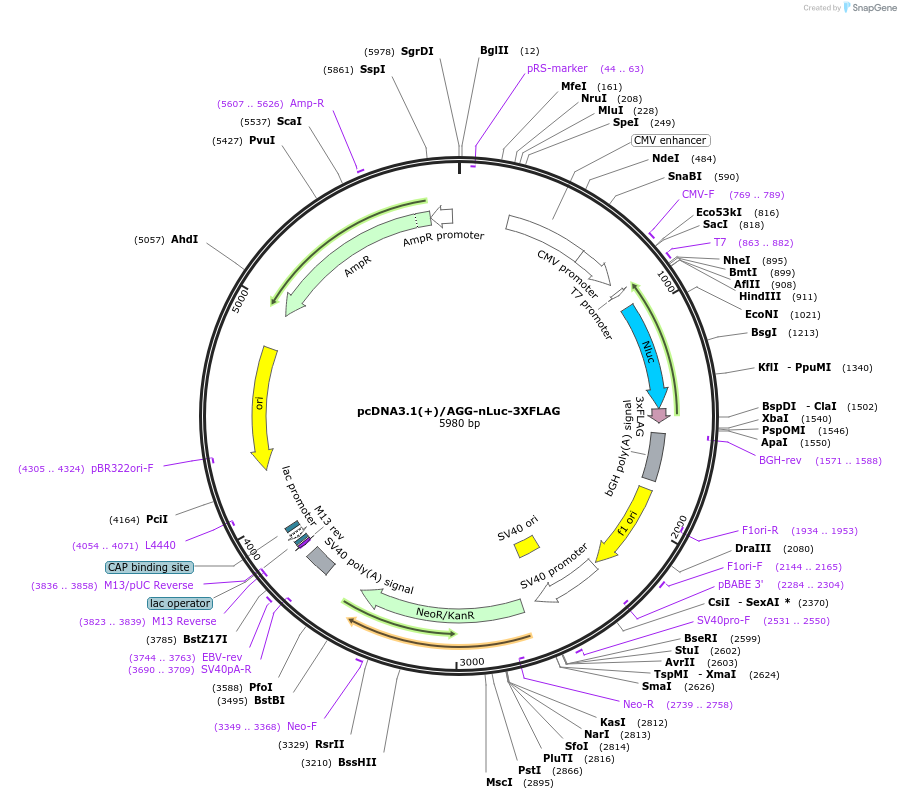
pcDNA3.1(+)/AGG-nLuc-3XFLAG
Plasmid#127304PurposeExpress 3xFLAG tagged nanoLuciferase (nLuc) from AGG start codonDepositorInsertnanoLuciferase
Tags3xFLAGExpressionMammalianPromoterCMVAvailable SinceJune 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
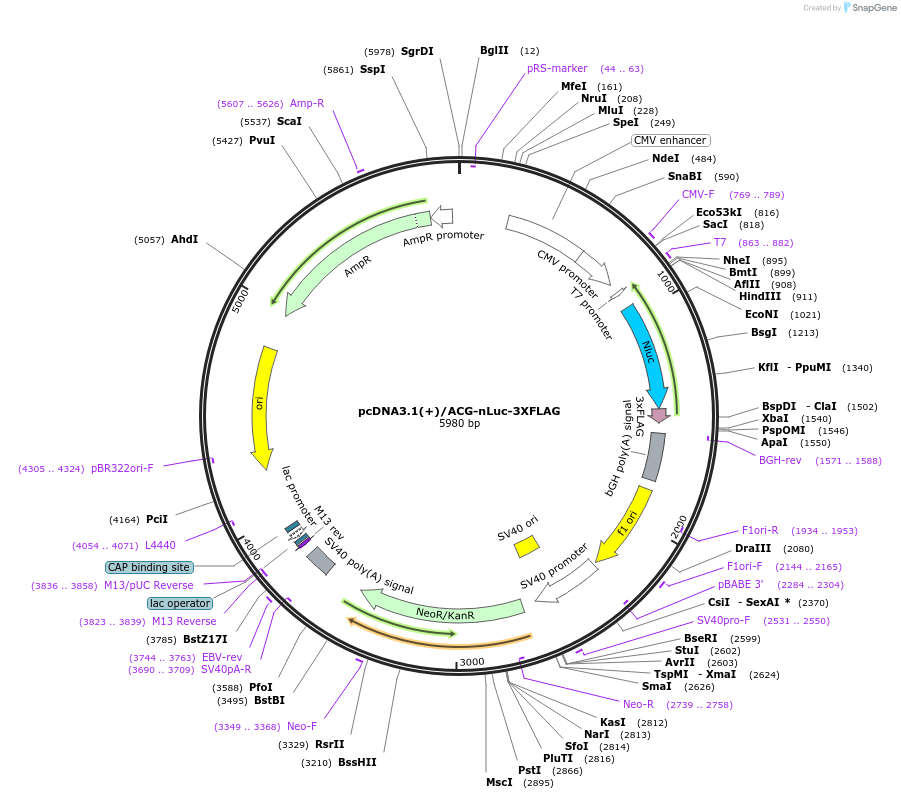
pcDNA3.1(+)/ACG-nLuc-3XFLAG
Plasmid#127303PurposeExpress 3xFLAG tagged nanoLuciferase (nLuc) from ACG start codonDepositorInsertnanoLuciferase
Tags3xFLAGExpressionMammalianPromoterCMVAvailable SinceJune 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
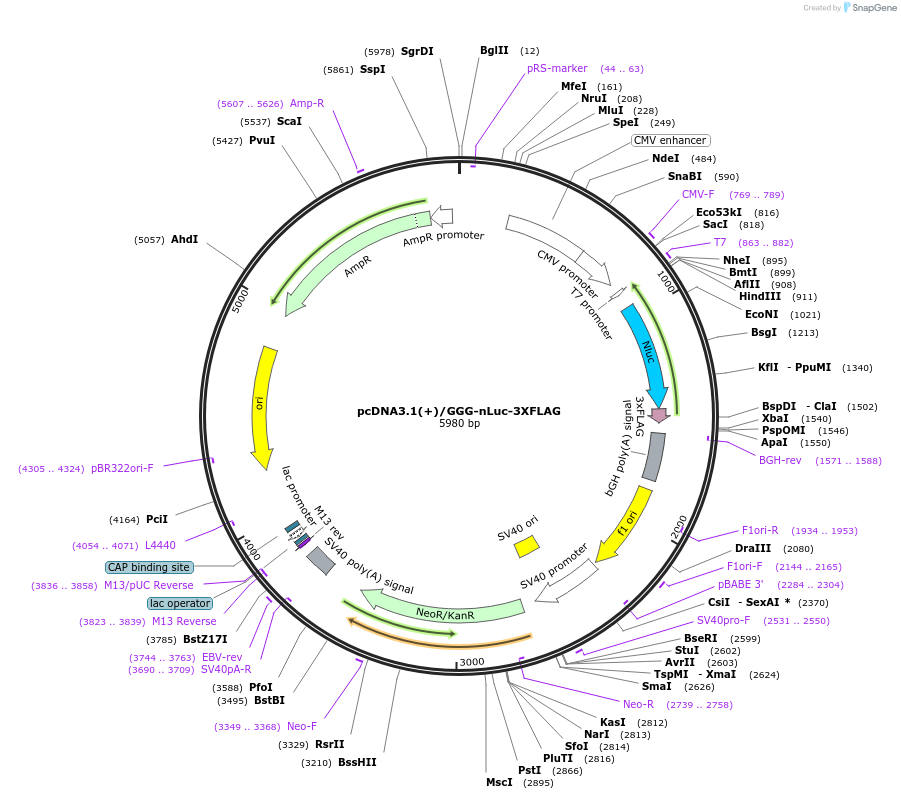
pcDNA3.1(+)/GGG-nLuc-3XFLAG
Plasmid#127307PurposeExpress 3xFLAG tagged nanoLuciferase (nLuc) from GGG start codon (Negative control plasmid)DepositorInsertnanoLuciferase
Tags3xFLAGExpressionMammalianPromoterCMVAvailable SinceJune 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
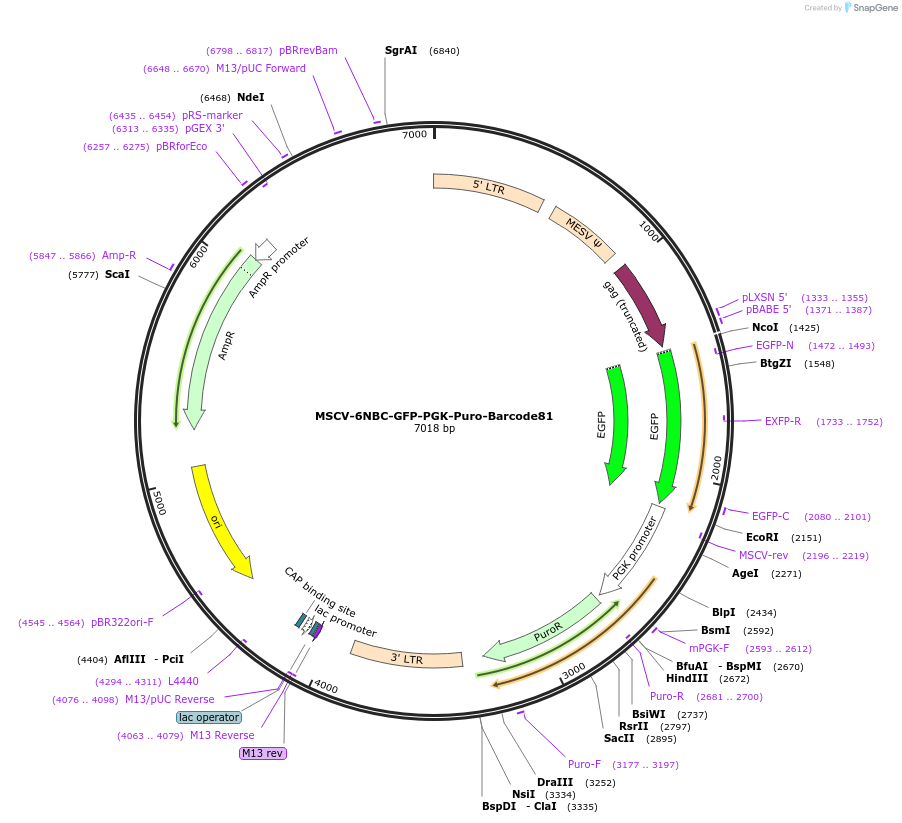
MSCV-6NBC-GFP-PGK-Puro-Barcode81
Plasmid#83758Purposeindividual 6N barcode to make barcoded cell line variantsDepositorInsertBarcode81
UseRetroviralMutationdeletion of T in starting ATG of GFP (please see …Available SinceNov. 10, 2016AvailabilityAcademic Institutions and Nonprofits only -
pHL 990
Plasmid#52992PurposepLtetO-1:DsrA, pCon:RhyB, pLlacO-1:rpoS::gfpDepositorInsertsDsrA sRNA
RhyB sRNA (competitor)
rpoS
UseSynthetic BiologyTagsgfpExpressionBacterialMutationincludes −150 to +30 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL 1022
Plasmid#52996PurposepLtetO-1:RhyB, pCon:OxyS, pLlacO-1:sodB::gfpDepositorInsertsRhyB sRNA
OxyS sRNA
sodB
UseSynthetic BiologyTagsgfpExpressionBacterialMutationcontains −56 to +141 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL 1020
Plasmid#52995PurposepLtetO-1:RhyB, pCon:DsrA, pLlacO-1:sodB::gfpDepositorInsertsRhyB sRNA
DsrA sRNA (competitor)
sodB
UseSynthetic BiologyTagsgfpExpressionBacterialMutationcontains −56 to +141 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL 1019
Plasmid#52994PurposepLtetO-1:RhyB, pCon:MicC, pLlacO-1:sodB::gfpDepositorInsertsRhyB sRNA
MicC sRNA (Competitor)
sodB
UseSynthetic BiologyTagsgfpExpressionBacterialMutationcontains −56 to +141 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL 1039
Plasmid#53000PurposepLtetO-1:DsrA, pCon:OxyS, pLlacO-1:rpoS::gfpDepositorInsertsDsrA sRNA
OxyS sRNA (competitor)
rpoS
UseSynthetic BiologyTagsgfpExpressionBacterialMutationcontains −150 to +30 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL 1394
Plasmid#53009PurposepLtetO-1:RhyB, pCon:T7(st3) (133bps)::mCherry, pLlacO-1:sodB::gfpDepositorInsertsRhyB sRNA
T7(st3) (133bps)
sodB
UseSynthetic BiologyTagsgfp and mCherryExpressionBacterialMutationcontains −56 to +141 bp relative to start codonPromoterpCon, pLlacO-1, and pLtetO-1Available SinceMay 14, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL1530
Plasmid#37569DepositorInsertsGFP
TetR
CsrA
UseSynthetic BiologyTagsglgC leader regionExpressionBacterialMutationstart codon removedPromoterpConNoHind and pLlacO-1Available SinceAug. 10, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1355
Plasmid#37566DepositorInsertsGFP
TetR
CsrA
UseSynthetic BiologyTagsglgC leader regionExpressionBacterialMutationstart codon removedPromoterpConNoHind and pConNoHindM12Available SinceJuly 26, 2012AvailabilityAcademic Institutions and Nonprofits only -
HOXB13 (human) Myc-tag-pSP72
Plasmid#8682DepositorInsertHOXB13 (HOXB13 Human)
UseTnt in vitroTagsMycExpressionBacterialMutationnew NsiI site at ATG startAvailable SinceMay 30, 2006AvailabilityAcademic Institutions and Nonprofits only -
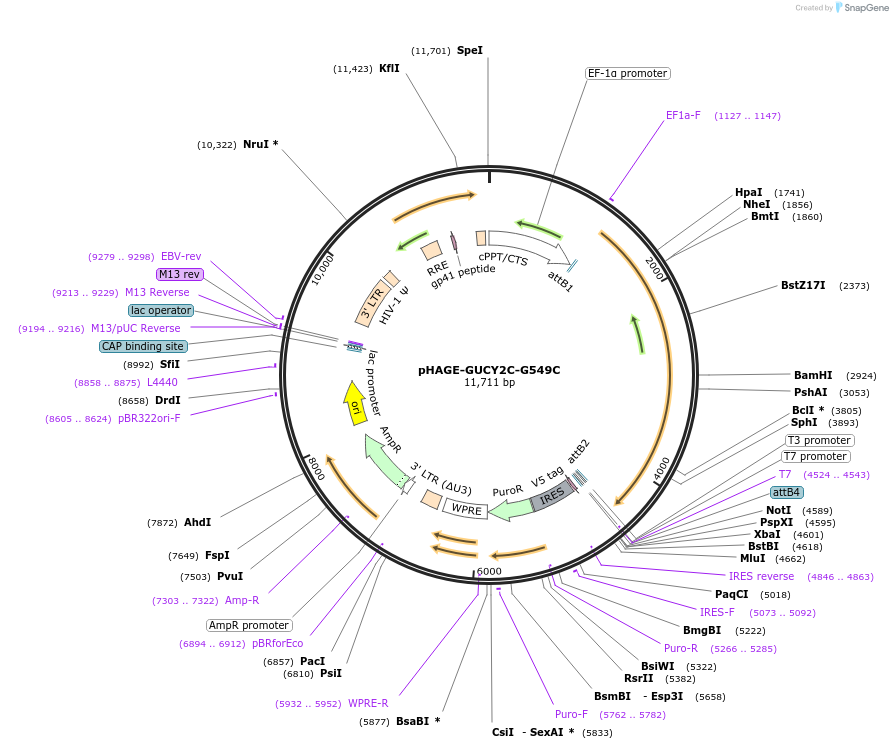
pHAGE-GUCY2C-G549C
Plasmid#116407PurposeLentiviral expression of GUCY2C G549CDepositorAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
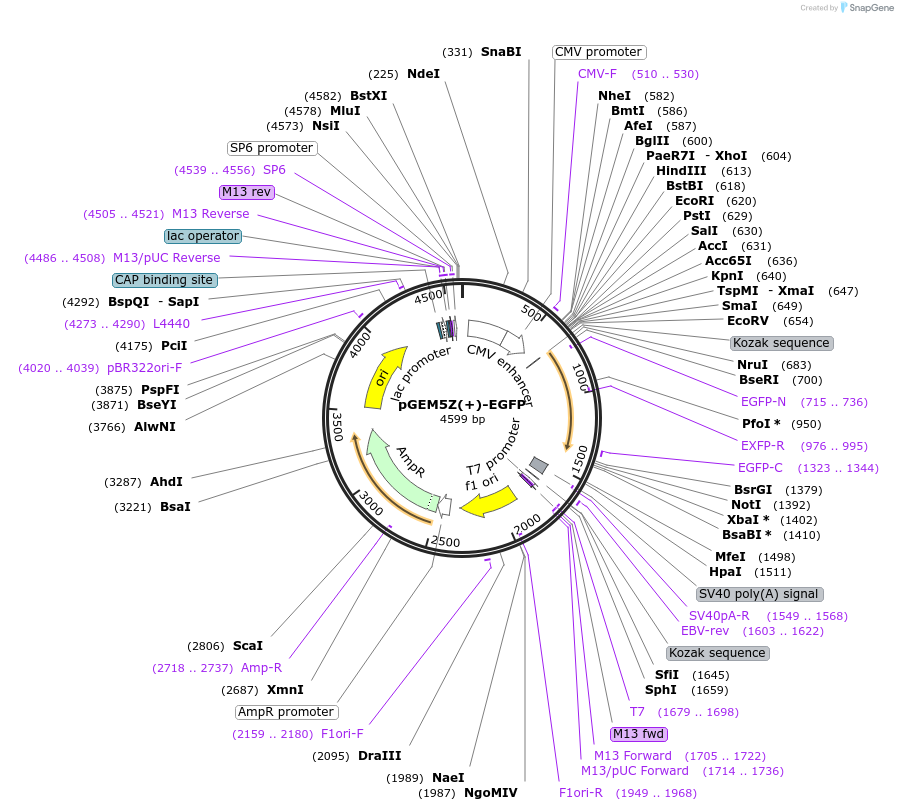
pGEM5Z(+)-EGFP
Plasmid#65206PurposeEGFP expression plasmid (also called p111) for the creation of a heteroduplex DNA repair assay plasmid with p189. *Please see notes in comments section.DepositorInsertEGFP
ExpressionBacterial and MammalianMutationRestriction enzyme recognition sites added either…PromoterCMVAvailable SinceJune 29, 2015AvailabilityAcademic Institutions and Nonprofits only -
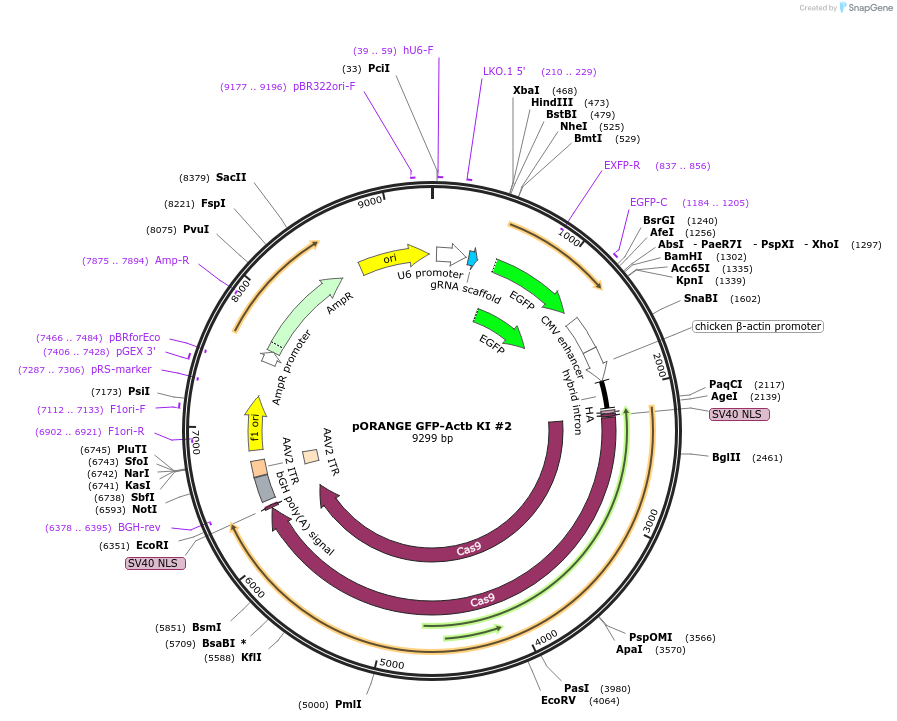
pORANGE GFP-Actb KI #2
Plasmid#139666PurposeEndogenous tagging of β-actin: N-terminal (amino acid position: before startcodon)DepositorInsertgRNA and GFP donor
ExpressionMammalianPromoterU6 and CbhAvailable SinceApril 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
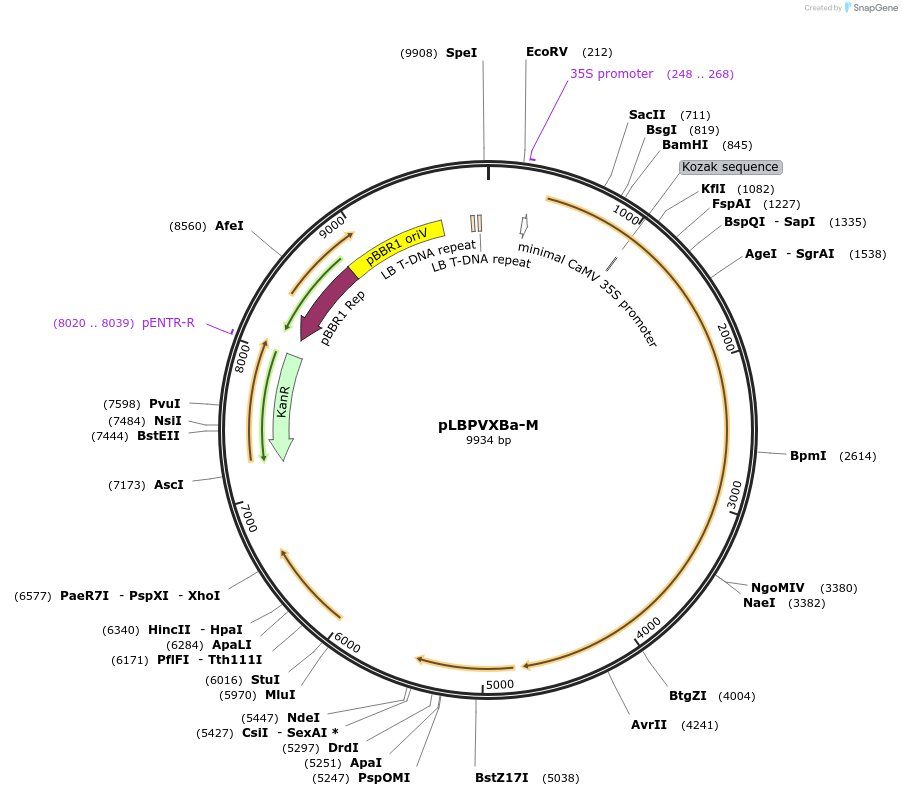
pLBPVXBa-M
Plasmid#229079PurposeNew version of a PVX vector to express heterologous sequences in plantsDepositorInsertPotato virus X
ExpressionPlantMutationMutation coat protein start codon (ATG to AGG); d…Available SinceFeb. 26, 2025AvailabilityAcademic Institutions and Nonprofits only -
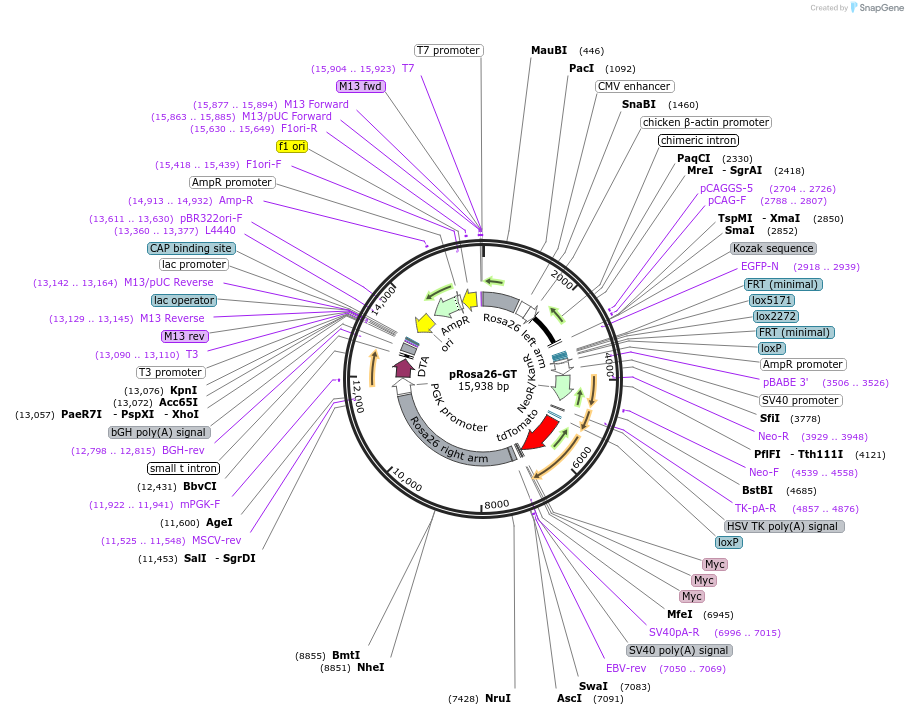
pRosa26-GT
Plasmid#40025DepositorInsertsGFP
beta-globin intron
Neo
tdT-3Myc
diphteria toxin A
Tags3 Myc tagsExpressionMammalianMutationdeleted nucleotides after nucleotide 274, inserti…PromoterCAG (chicken beta globin promoter and CMV enhance…Available SinceOct. 22, 2012AvailabilityAcademic Institutions and Nonprofits only -
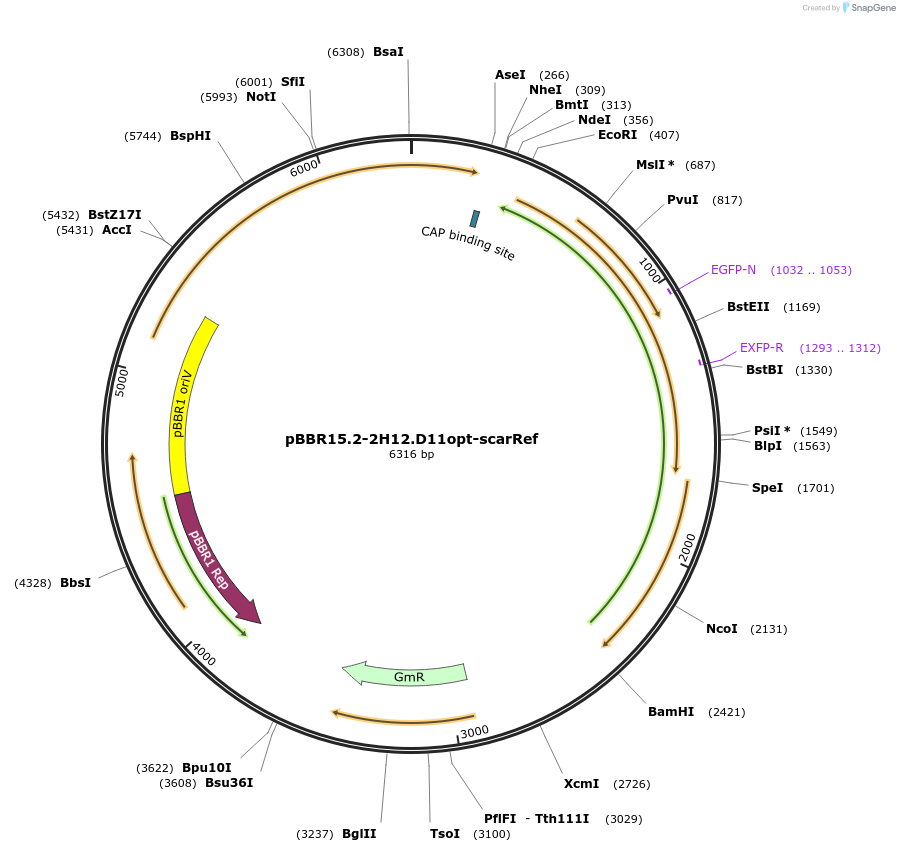
pBBR15.2-2H12.D11opt-scarRef
Plasmid#221150PurposecdGreen2.1 (optimized for Pseudomonas GC-content) expressed from a constitutive promoter in an operon with mScarlet-I (downstream) as a reference FP (Internal Lab ID: UJ11241)DepositorInsertscdGreen2.1
mScarlet-I
ExpressionBacterialMutationN-terminus changed: starts with MSKKYGEAVIKE ...PromoterConstitutive and None (same as cdGreen2.1)Available SinceJune 25, 2024AvailabilityAcademic Institutions and Nonprofits only