We narrowed to 83,870 results for: myc
-
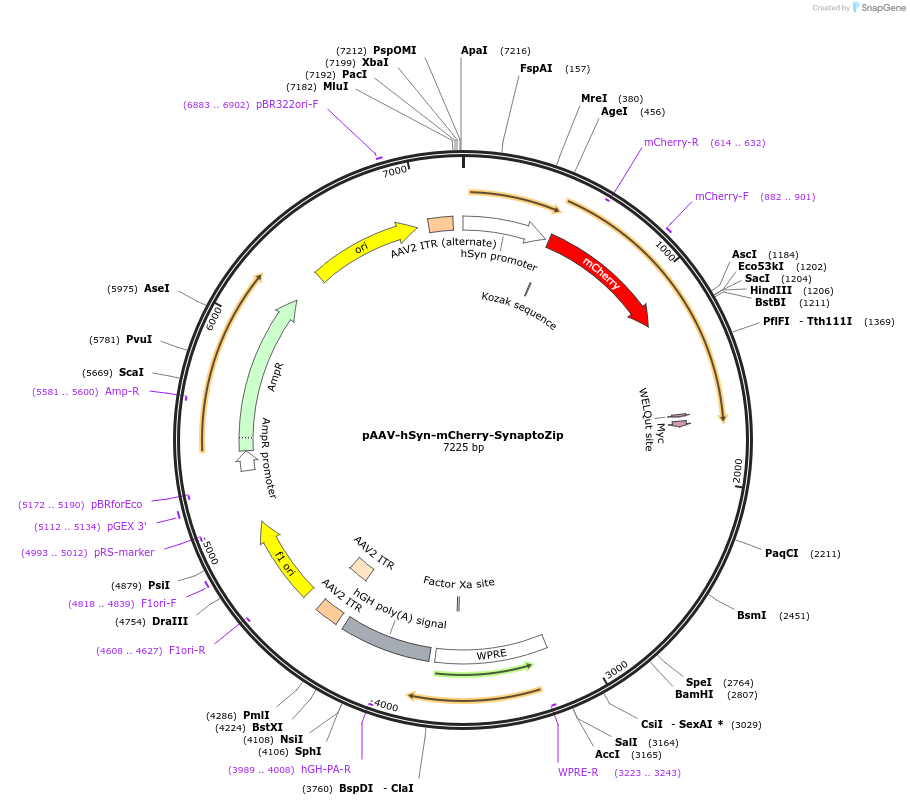
Plasmid#177318PurposeAAV expression of the synaptic activity-marker SynaptoZip fused to mCherry driven by human Synapsin promoterDepositorInsertmCherry-SynaptoZip (Vamp2 Rat)
UseAAVTagsMyc and mCherryExpressionMammalianPromoterhSynAvailable SinceJune 24, 2022AvailabilityAcademic Institutions and Nonprofits only -
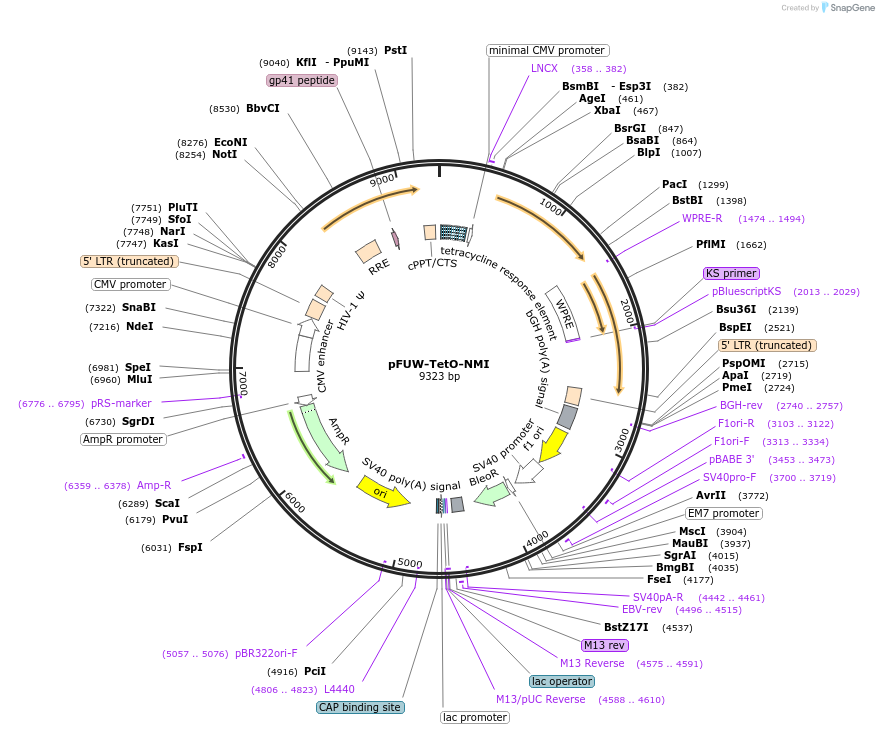
pFUW-TetO-NMI
Plasmid#178446Purposedoxycycline-inducible expression of Human NMI in mammalian cellsDepositorAvailable SinceMarch 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
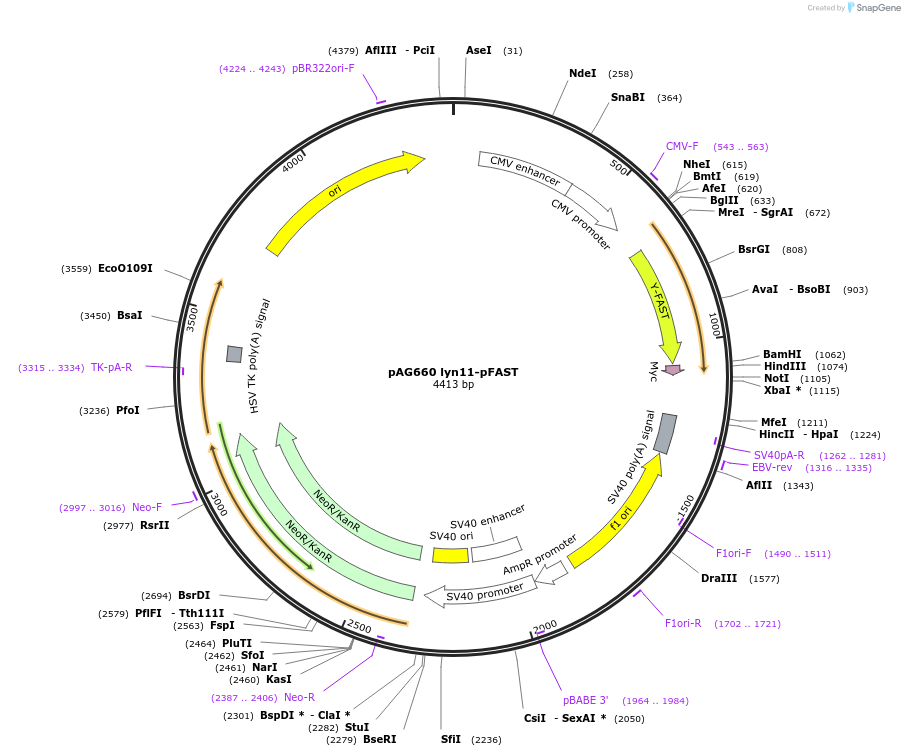
pAG660 lyn11-pFAST
Plasmid#172863Purposeexpresses lyn11-pFAST in mammalian cells (inner membrane labeling)DepositorInsertLyn11-pFAST
TagsMyc tag (C terminal on insert)ExpressionMammalianPromoterCMVAvailable SinceFeb. 3, 2022AvailabilityAcademic Institutions and Nonprofits only -
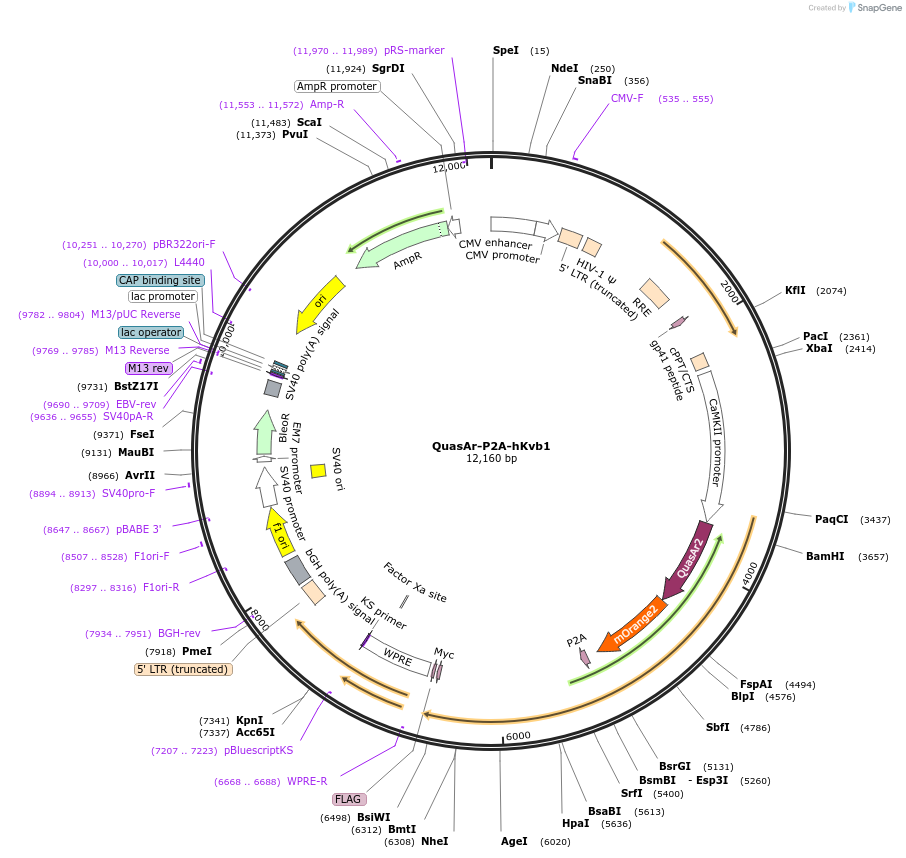
QuasAr-P2A-hKvb1
Plasmid#154083PurposeIndependent expression of QuasAr and human Kvbeta 1 in pFCK vectorDepositorInsertQuasAr-P2A-human Kvb1 (KCNAB1 Human, Synthetic)
UseLentiviralTagsmyc-FlagExpressionMammalianPromoterCaMKIIAvailable SinceNov. 2, 2020AvailabilityAcademic Institutions and Nonprofits only -
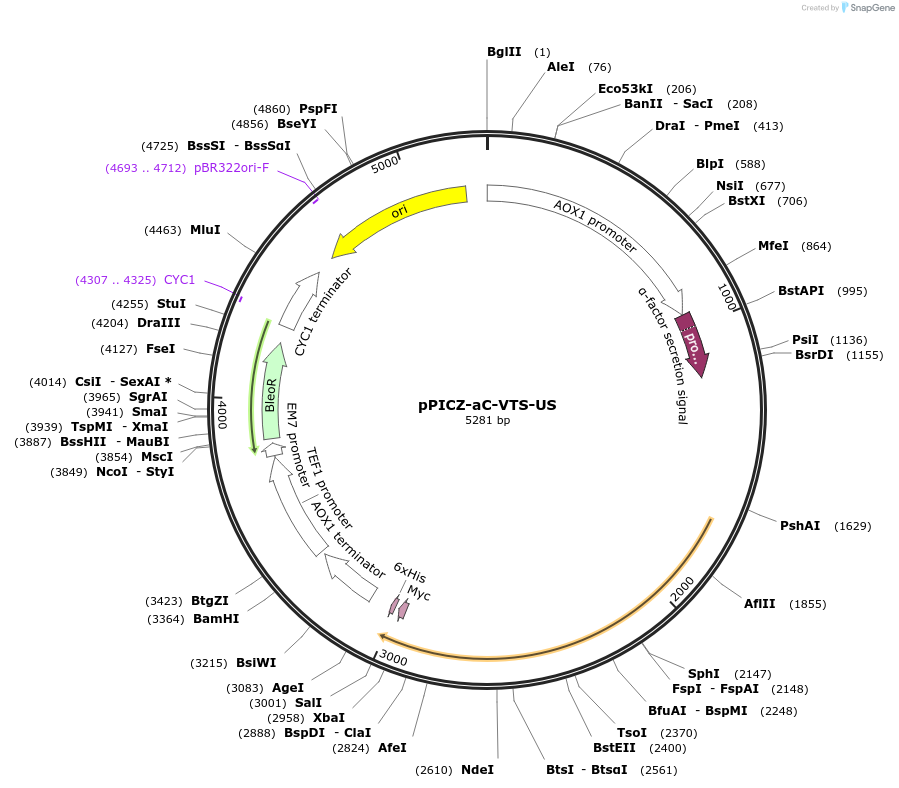
pPICZ-aC-VTS-US
Plasmid#160425PurposeExpression of c-luciferase from sp. Vargula tsujii (USA)DepositorInsertluciferase
UseLuciferaseTags6x Histidine tag, Yeast alpha secretion factor, c…ExpressionYeastMutationSee depositor comments belowPromoterAOXAvailable SinceOct. 23, 2020AvailabilityAcademic Institutions and Nonprofits only -
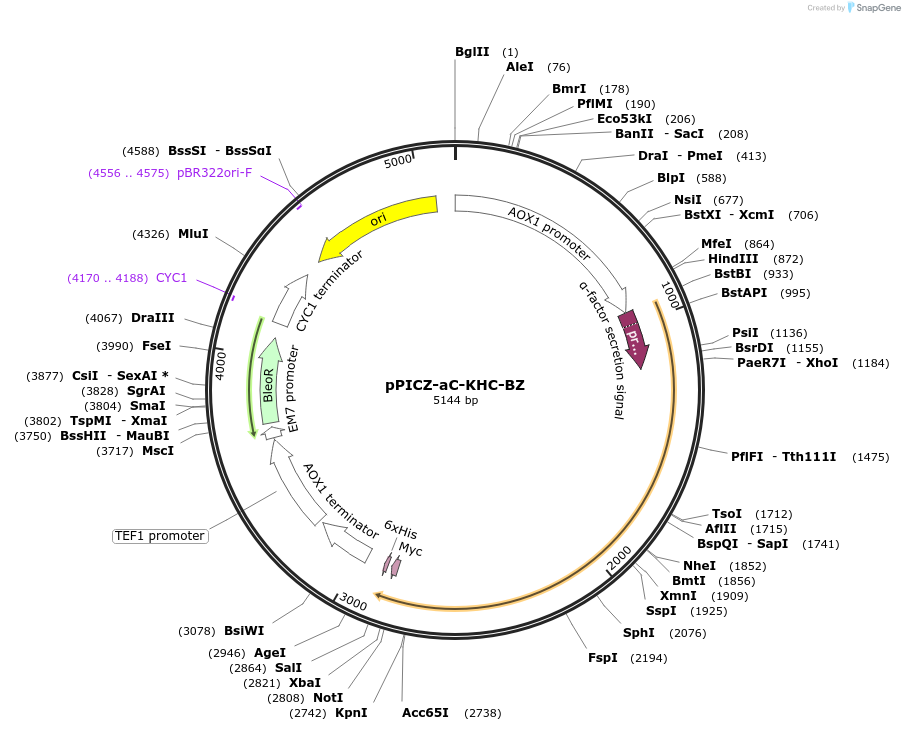
pPICZ-aC-KHC-BZ
Plasmid#160427PurposeExpression of c-luciferase from sp. Kornikeria hastingsi carriebowae (Belize)DepositorInsertluciferase
UseLuciferaseTags6x Histidine tag, Yeast alpha secretion factor, c…ExpressionYeastPromoterAOXAvailable SinceOct. 23, 2020AvailabilityAcademic Institutions and Nonprofits only -
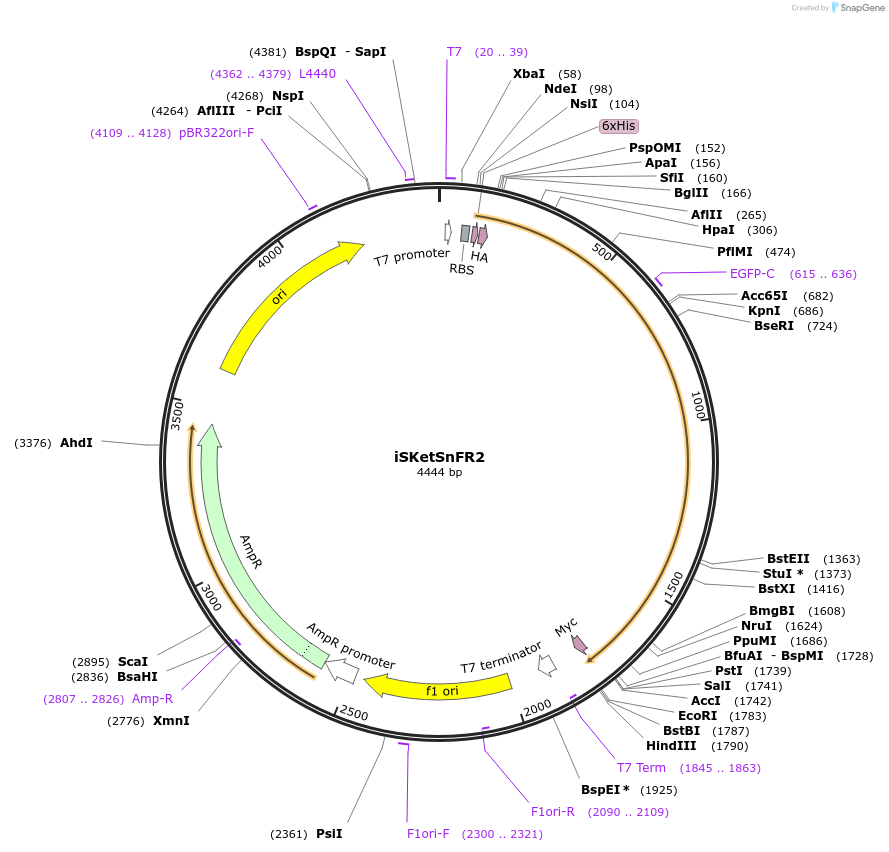
iSKetSnFR2
Plasmid#140467PurposeDevelop a family of genetically encoded fluorescent biosensors for S-ketamine, termed iSKetSnFR2, thus enabling optical subcellular pharmacokinetics for S-Ketamine.DepositorInsertiSKetSnFR2
TagsHis Tag, HA Tag and Myc tagExpressionBacterialPromoterT7Available SinceMay 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
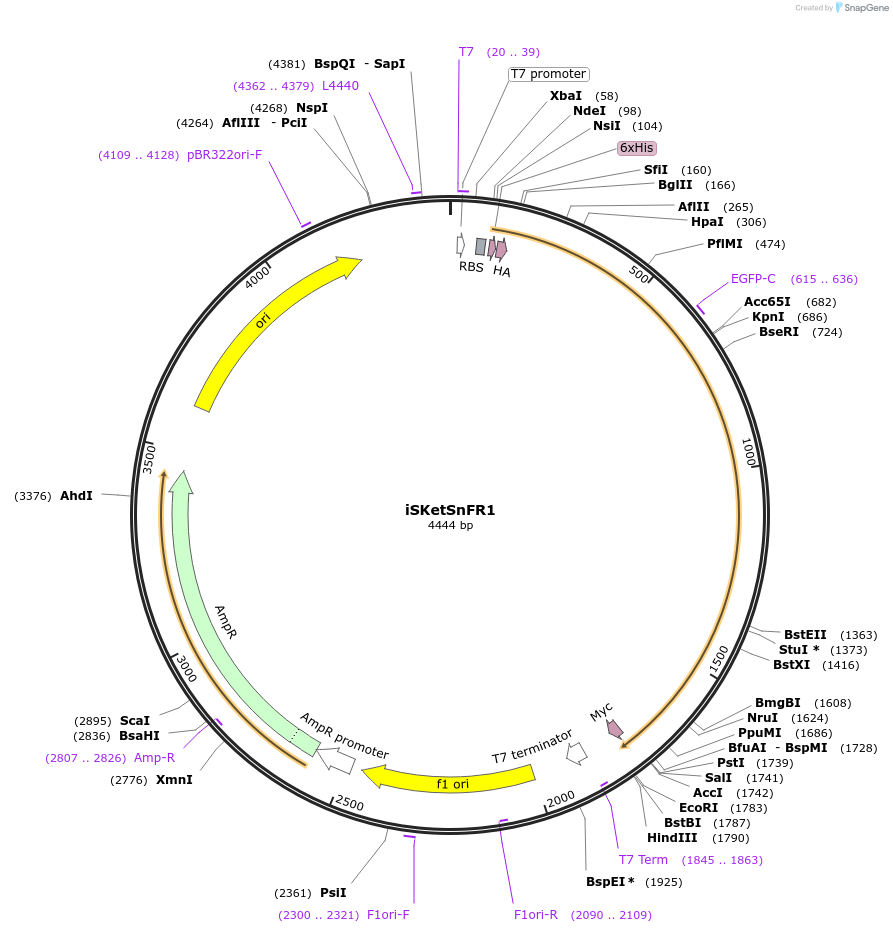
iSKetSnFR1
Plasmid#140466PurposeDevelop a family of genetically encoded fluorescent biosensors for S-ketamine, termed iSKetSnFR1, thus enabling optical subcellular pharmacokinetics for S-Ketamine.DepositorInsertiSKetSnFR1
TagsHis Tag, HA Tag and Myc tagExpressionBacterialPromoterT7Available SinceMay 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
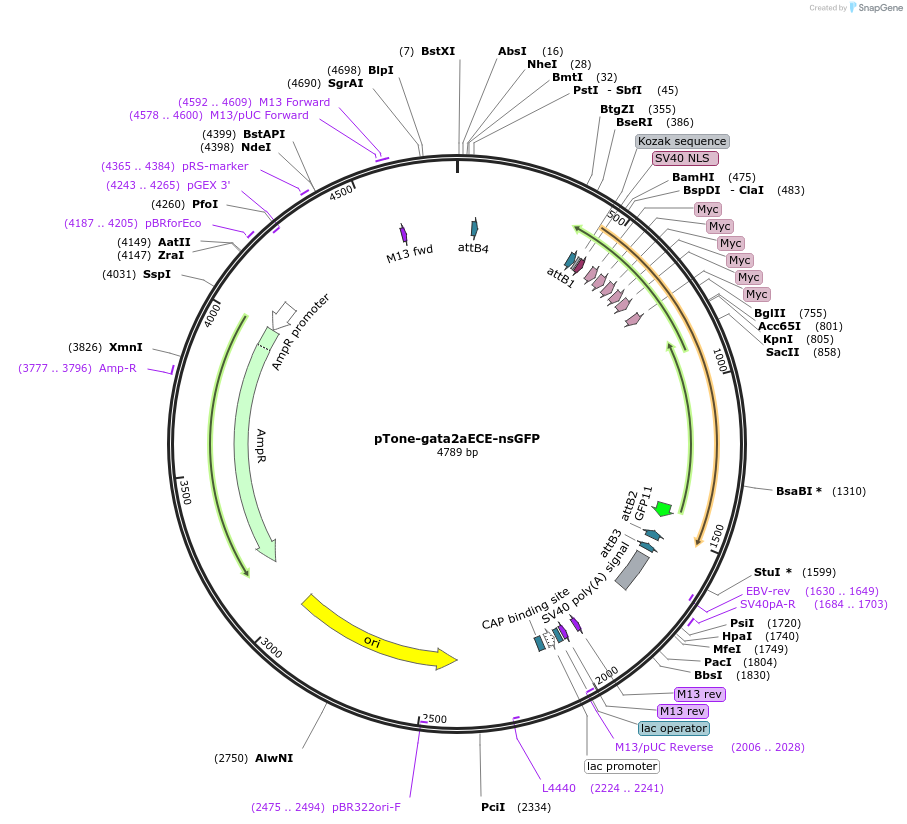
pTone-gata2aECE-nsGFP
Plasmid#132975Purposegata2a endothelial enhancer (x6) and basal promoter driving nuclear-localized sfGFP in pTol1 backboneDepositorInsertsnuclear localized sfGFP
gata2a endothelial enhancer (x6) with a carp b-actin basal promoter
UseUnspecified; Transposon-mediatedTags6x myc and SV40 NLSAvailable SinceDec. 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
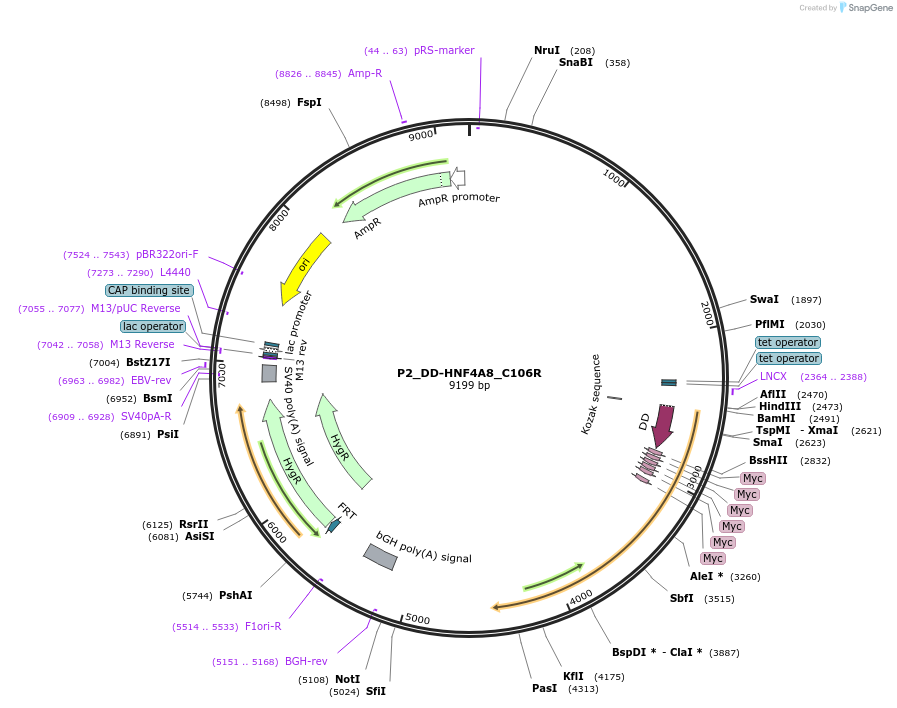
P2_DD-HNF4A8_C106R
Plasmid#31096DepositorInsertHNF4A (HNF4A Human)
UseFlp/frtTagsFKBP L106P and mycExpressionMammalianMutationsplice variant 8 derived from promoter P2 with zi…Available SinceOct. 14, 2011AvailabilityAcademic Institutions and Nonprofits only -
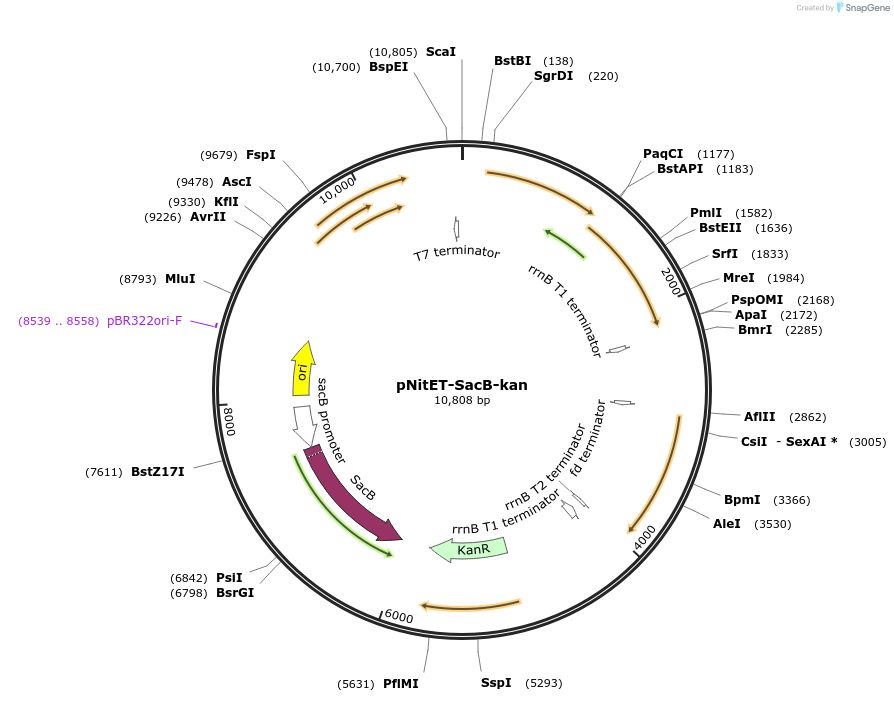
pNitET-SacB-kan
Plasmid#107692PurposeExpression of the phage Che9c RecET system for Recombineering in mycobacteriaDepositorInsertsChe9 phage RecET
PNit2-NitR
SacRB
ExpressionBacterialAvailable SinceMay 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
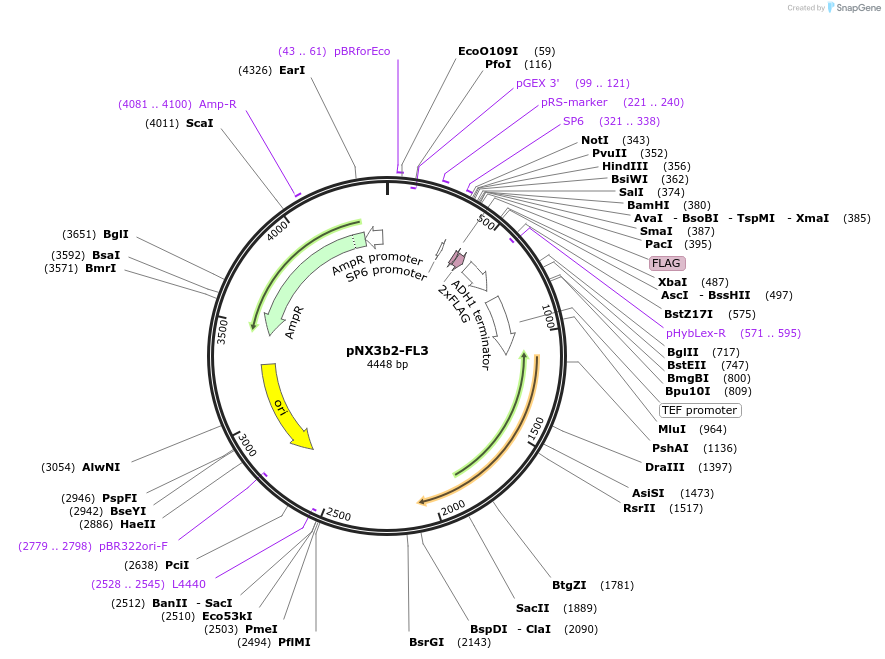
pNX3b2-FL3
Plasmid#78388Purposefission yeast epitope taggingDepositorInserts3xFLAG
hygML
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
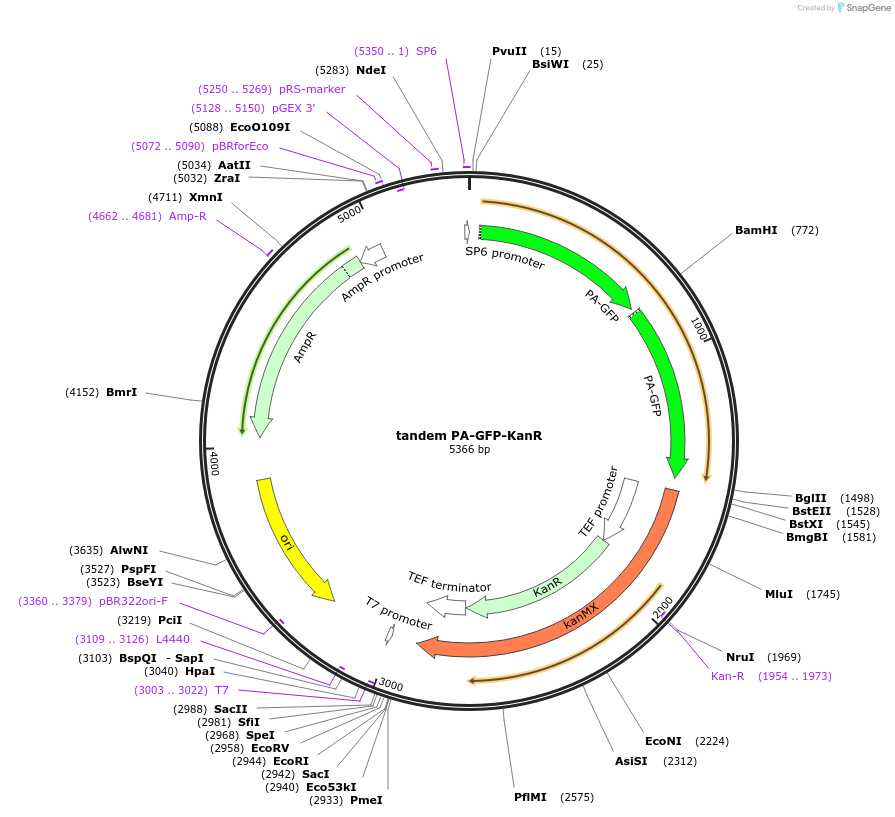
tandem PA-GFP-KanR
Plasmid#37548DepositorInserttandem PA-GFP
UseYeast taggingAvailable SinceJuly 12, 2012AvailabilityAcademic Institutions and Nonprofits only -
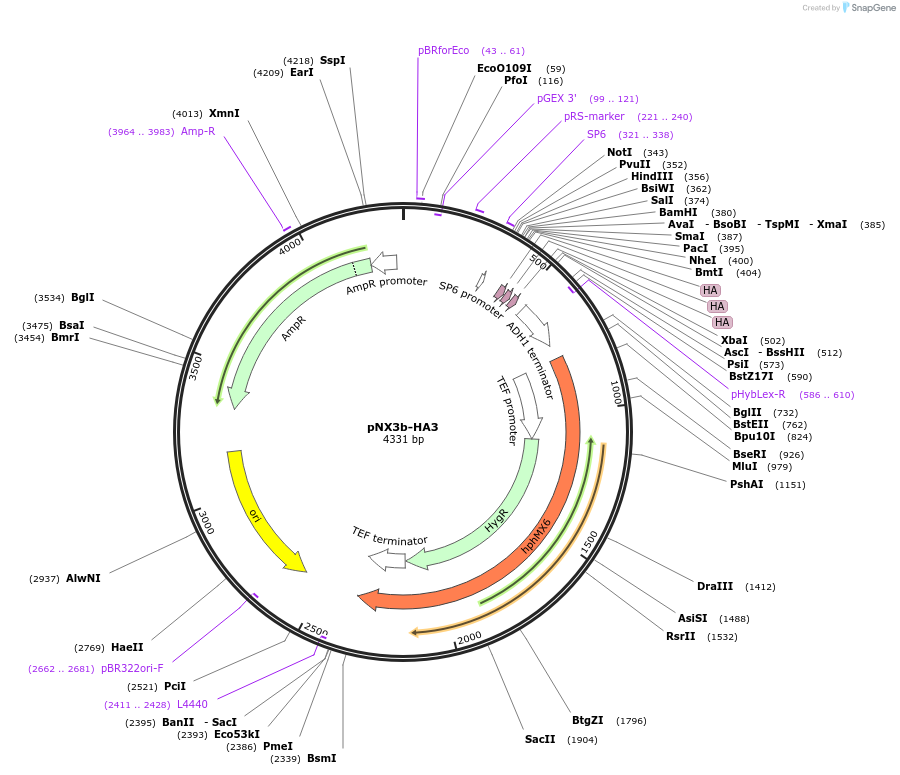
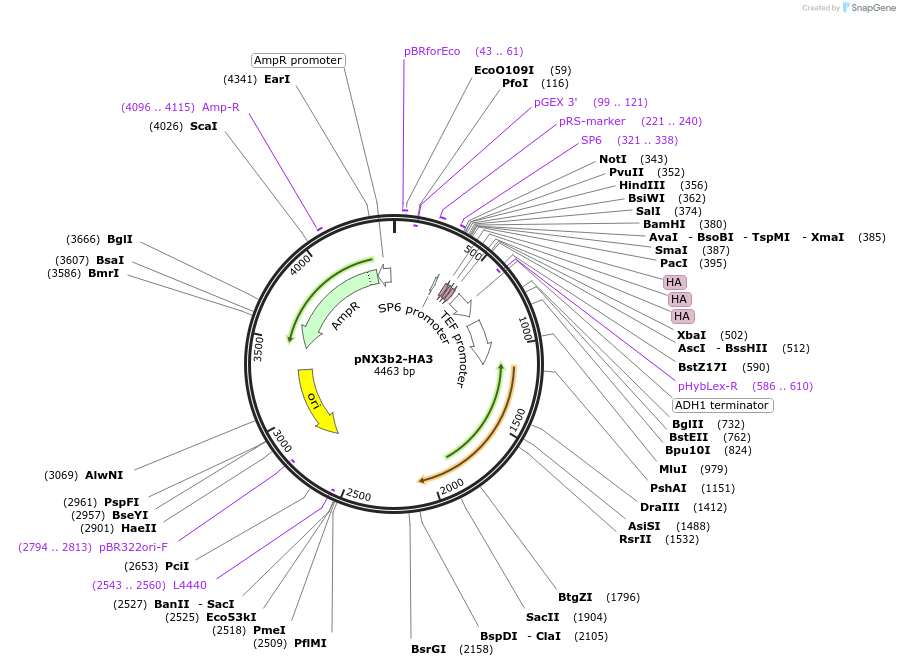
pNX3b-HA3
Plasmid#78366Purposefission yeast epitope taggingDepositorInserts3xHA
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
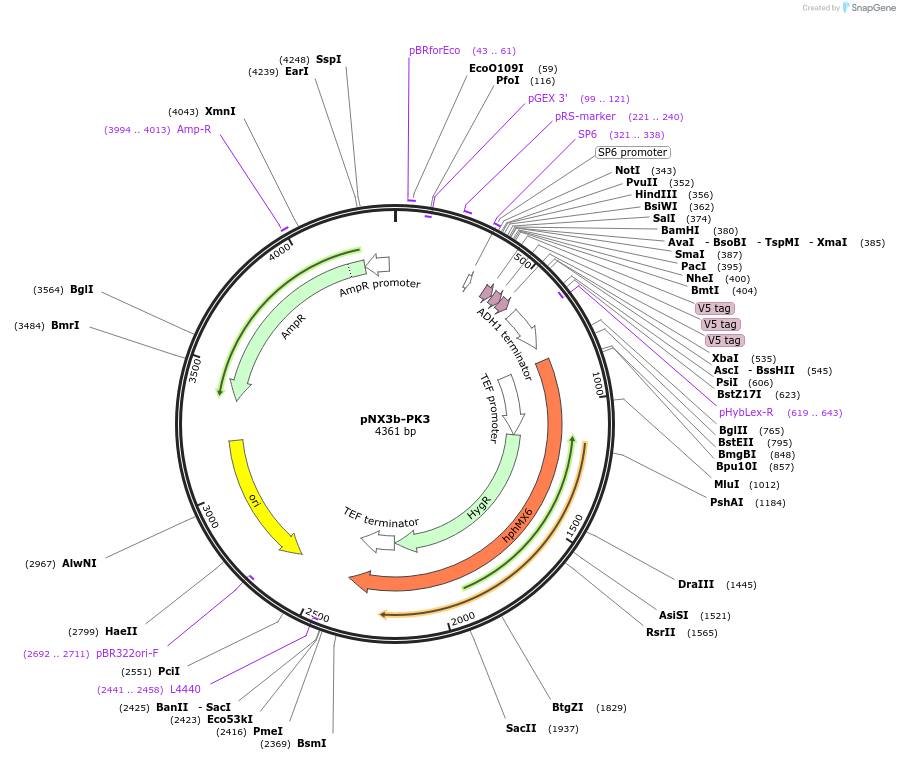
pNX3b-PK3
Plasmid#78361Purposefission yeast epitope taggingDepositorInserts3xPK
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
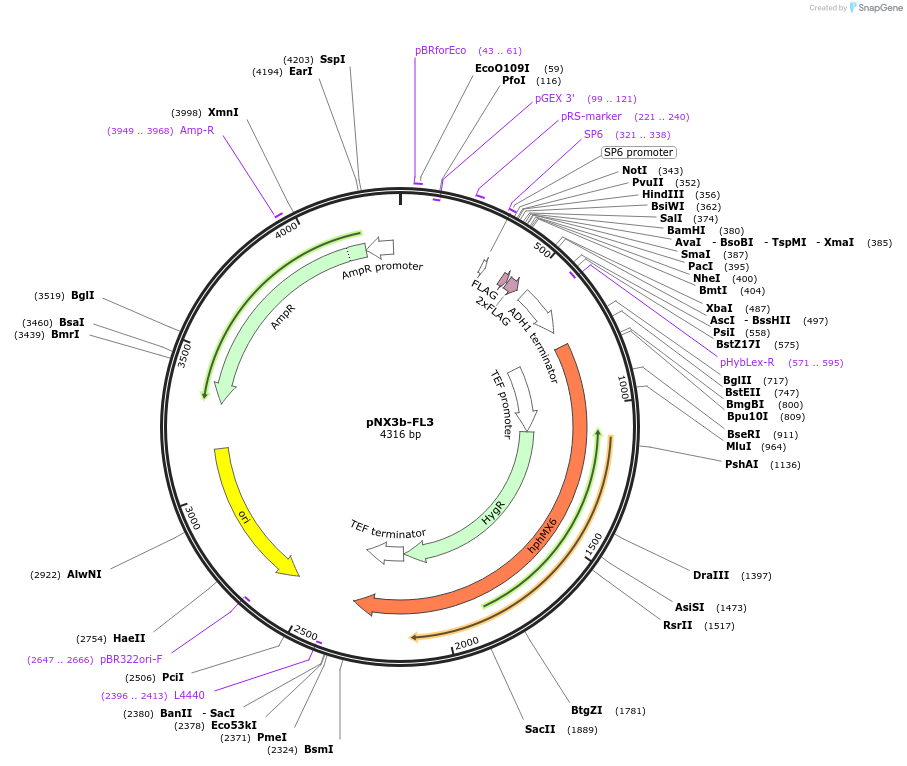
pNX3b-FL3
Plasmid#78365Purposefission yeast epitope taggingDepositorInserts3xFLAG
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
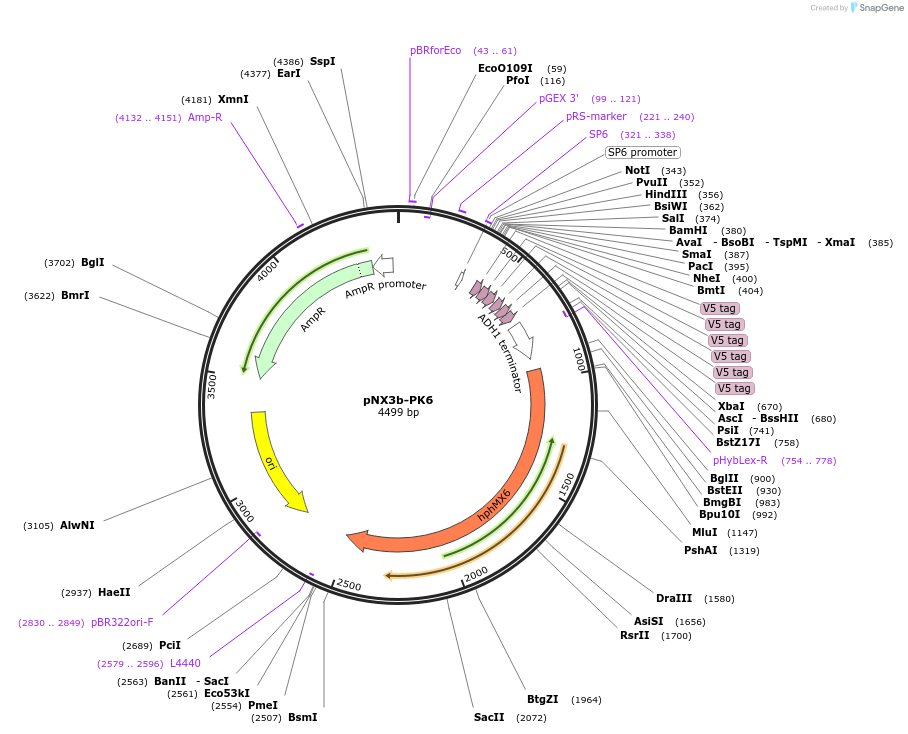
pNX3b-PK6
Plasmid#78362Purposefission yeast epitope taggingDepositorInserts6xPK
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
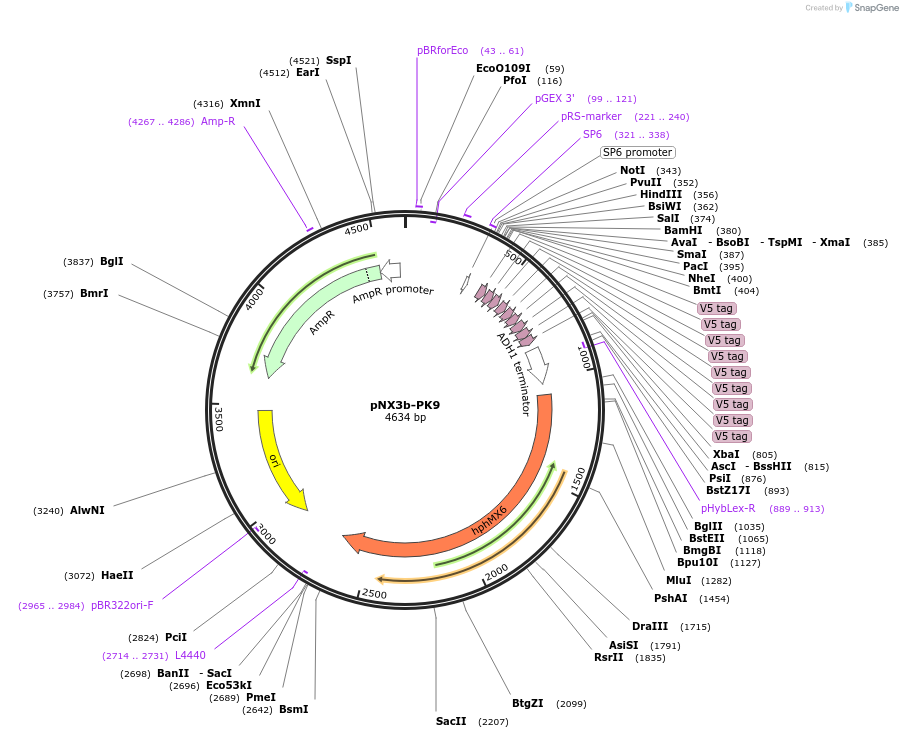
pNX3b-PK9
Plasmid#78363Purposefission yeast epitope taggingDepositorInserts9xPK
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
pNX3b2-HA3
Plasmid#78387Purposefission yeast epitope taggingDepositorInserts3xHA
hygML
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
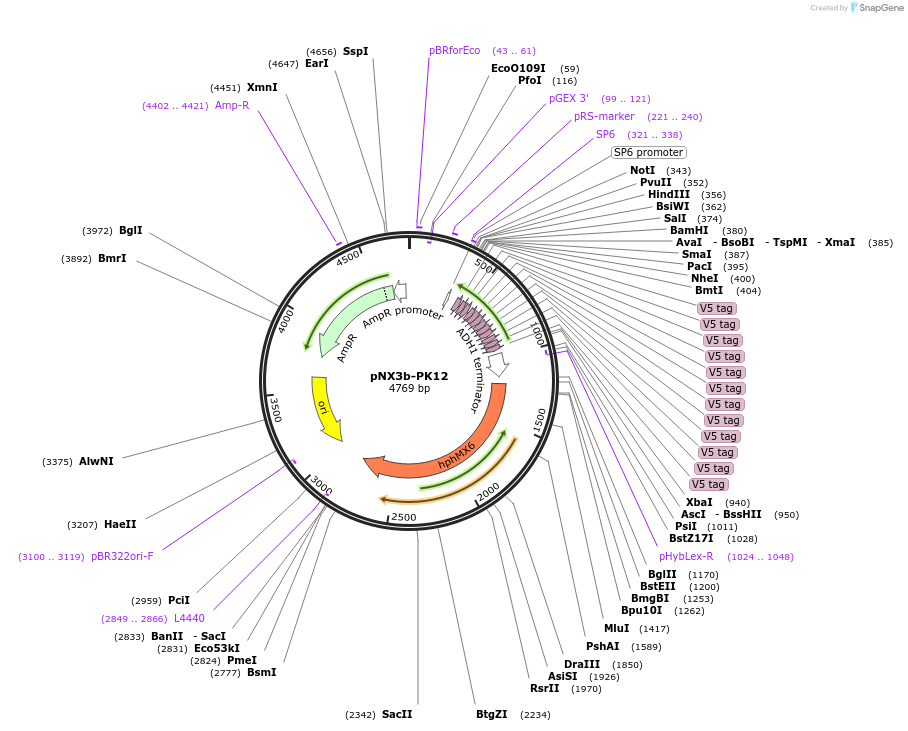
pNX3b-PK12
Plasmid#78364Purposefission yeast epitope taggingDepositorInserts12xPK
hygMX6
Available SinceNov. 17, 2016AvailabilityAcademic Institutions and Nonprofits only