We narrowed to 3,681 results for: mpo
-
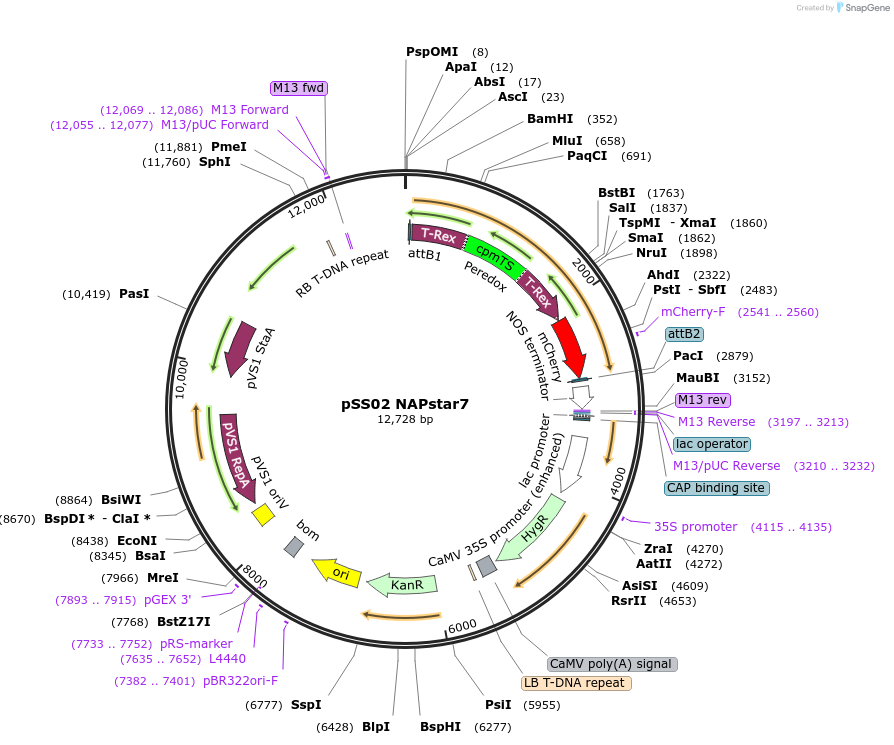
Plasmid#241224PurposeExpresses NADPH/NADP+ biosensor NAPstar7 in Arabidopsis thalianaDepositorInsertcyt-NAPstar7
ExpressionPlantAvailable SinceSept. 3, 2025AvailabilityAcademic Institutions and Nonprofits only -
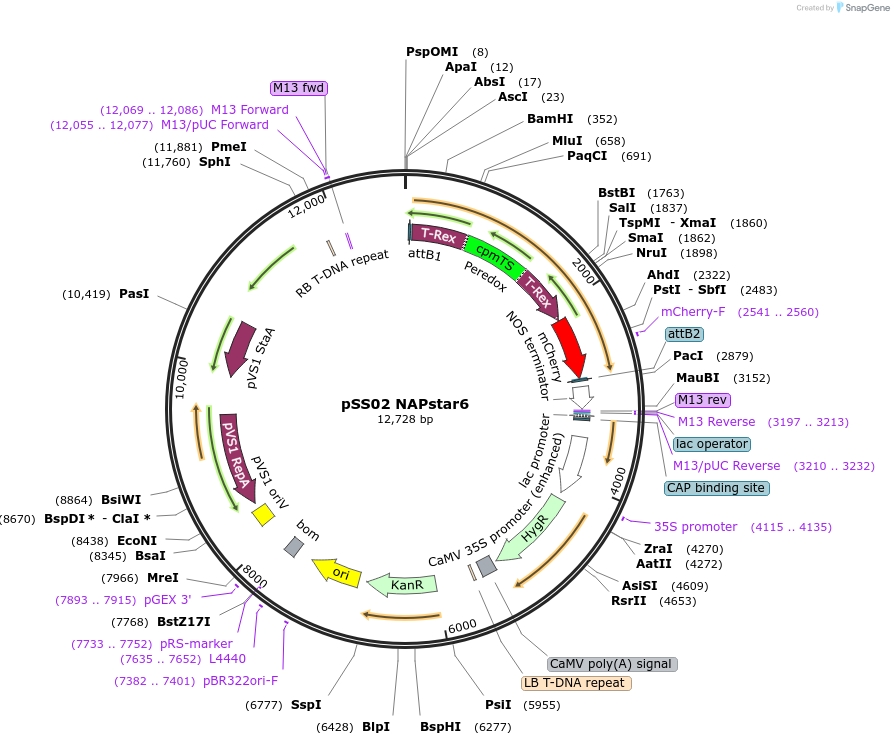
pSS02 NAPstar6
Plasmid#241223PurposeExpresses NADPH/NADP+ biosensor NAPstar6 in Arabidopsis thalianaDepositorInsertcyt-NAPstar6
ExpressionPlantAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
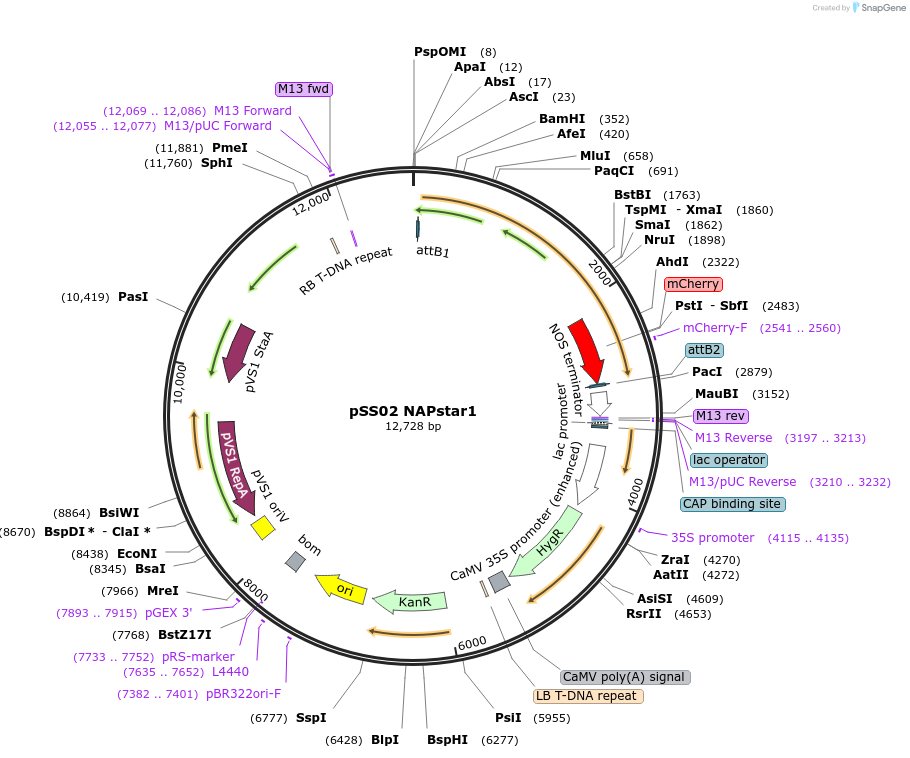
pSS02 NAPstar1
Plasmid#241270PurposeExpresses NADPH/NADP+ biosensor NAPstar1 in Arabidopsis thalianaDepositorInsertcyt-NAPstar1
ExpressionPlantAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
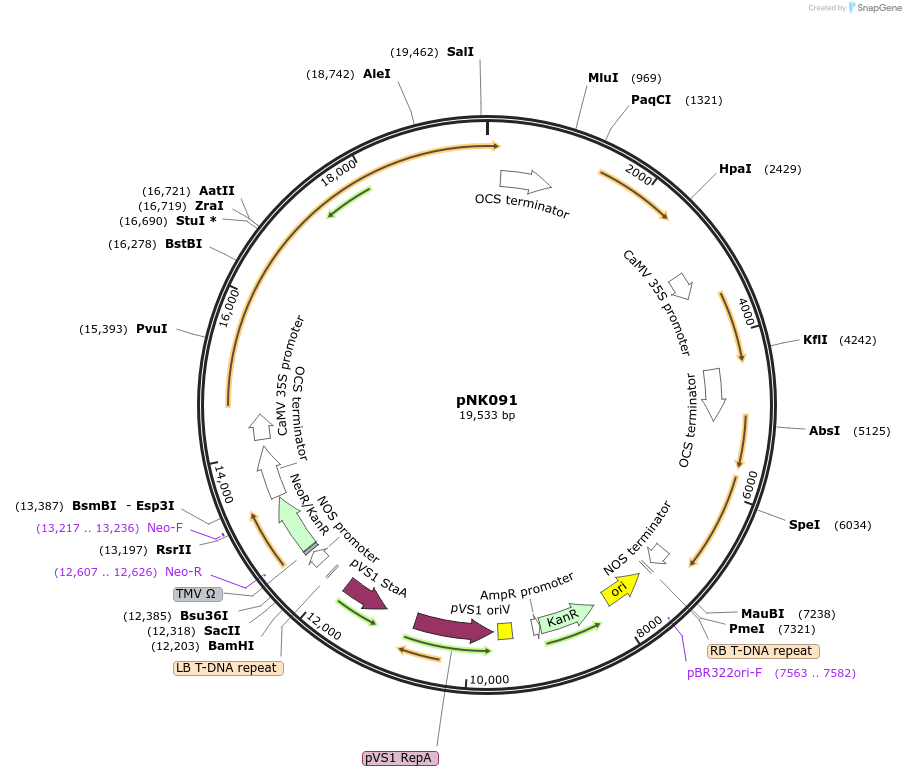
pNK091
Plasmid#219769PurposeMoClo-compatible Level P vector carrying fungal bioluminescence system lacking luciferase geneDepositorInsertpNos-KanR-ocsT | p35S-nnHispS-ocsT | pCmYLCV-npgA-ATPT | p35S-nnCPH-ocsT | pFMV-nnH3H-nosT
UseSynthetic BiologyExpressionPlantPromoterpNos-KanR-ocsT | p35S-nnHispS-ocsT | pCmYLCV-npgA…Available SinceJuly 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
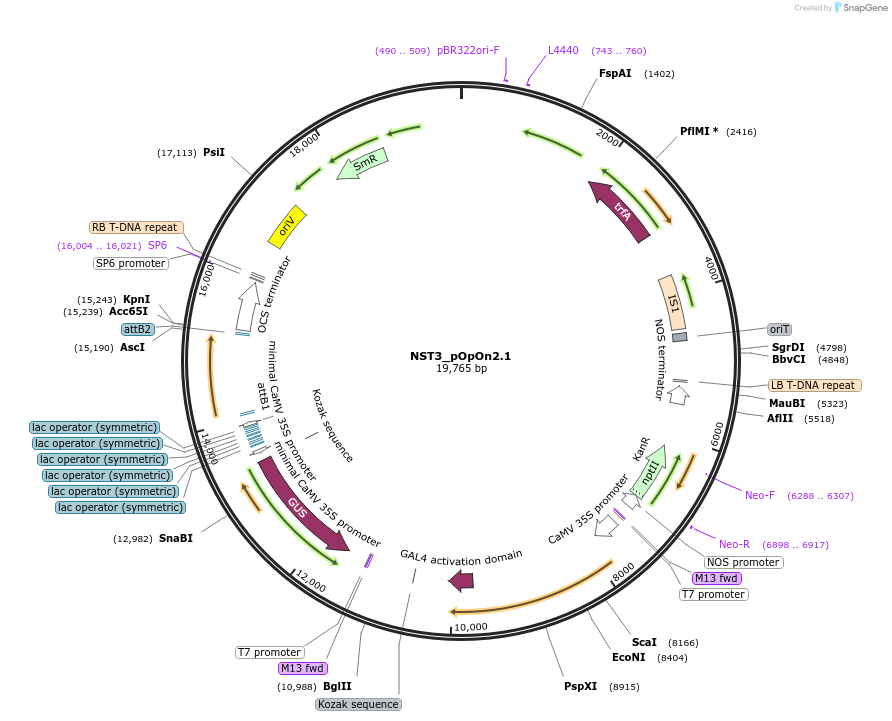
NST3_pOpOn2.1
Plasmid#228066PurposeExpresses NST3 in different cell types in Arabidopsis thaliana upon induction with dexamethasone.DepositorInsertNAC SECONDARY WALL THICKENING PROMOTING FACTOR3, NST3 (NAC012 Mustard Weed)
UseDexamethasone inducible vectorExpressionPlantPromoterCaMV35S for LhGR and pOp6-minimal CaMV35S for NST3Available SinceJan. 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
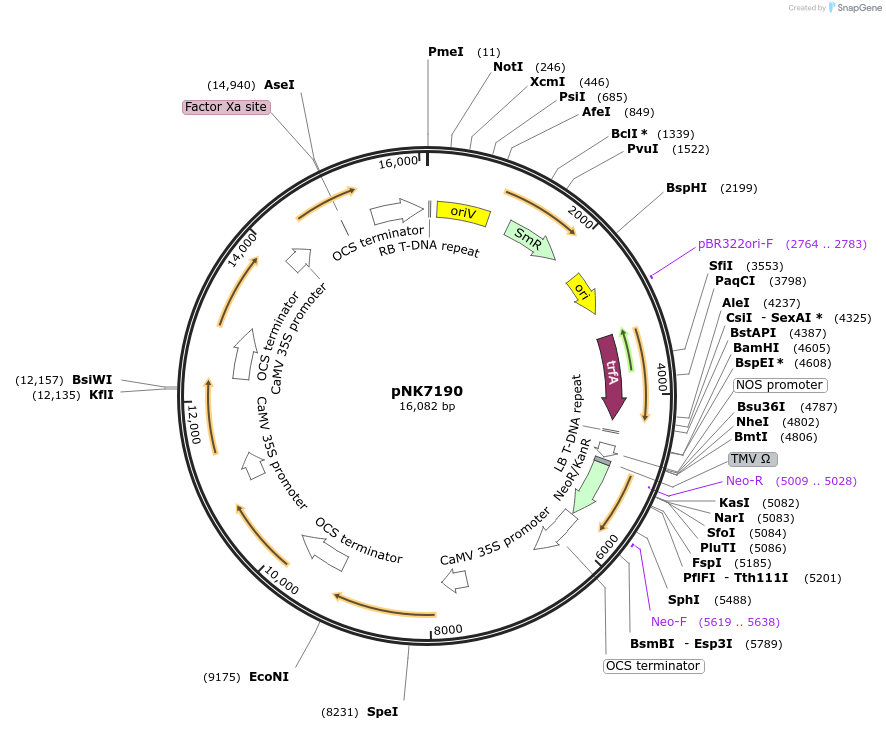
pNK7190
Plasmid#219763PurposeMoClo-compatible Level M for expression nnLuz, nnH3H, nnCPH in plants (N.benthamiana, BY2 cells)DepositorInsertnnLuz, nnH3H, nnCPH
ExpressionPlantAvailable SinceOct. 17, 2024AvailabilityAcademic Institutions and Nonprofits only -
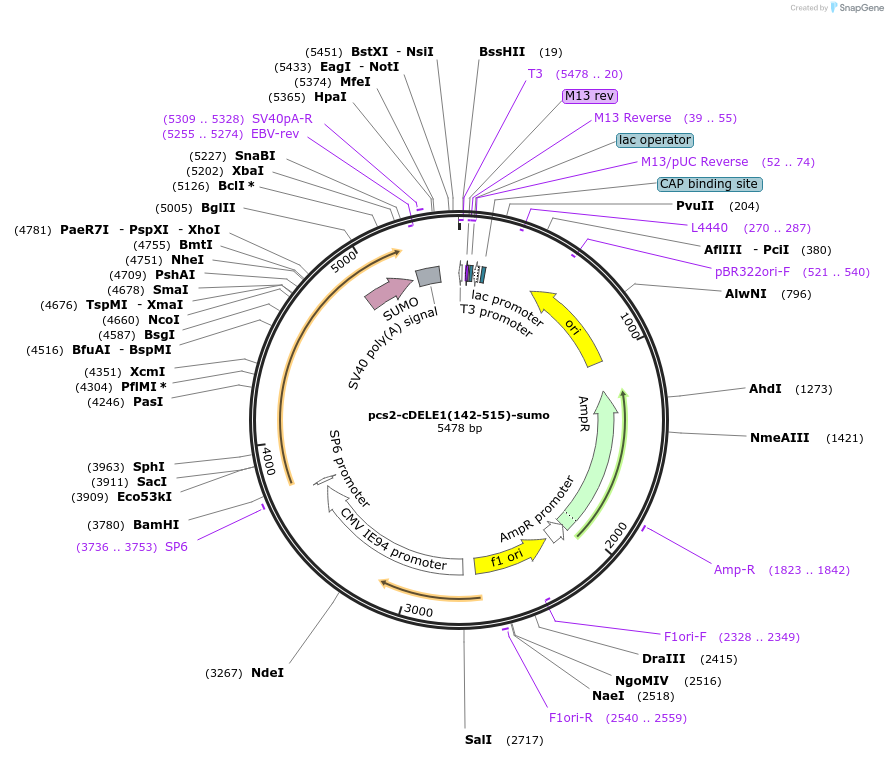
pcs2-cDELE1(142-515)-sumo
Plasmid#226107PurposecDELE1 (corresponding to mitochondrially imported, cleaved DELE1, residues 142-515) fused to sumo used for in-vitro translation by wheat germ extract in in-vitro ubiquitylation assaysDepositorInsertDELE1 (DELE1 Human)
TagsSumoExpressionMammalianMutationencodes only residues 142-515 of DELE1 (represent…Available SinceOct. 16, 2024AvailabilityAcademic Institutions and Nonprofits only -
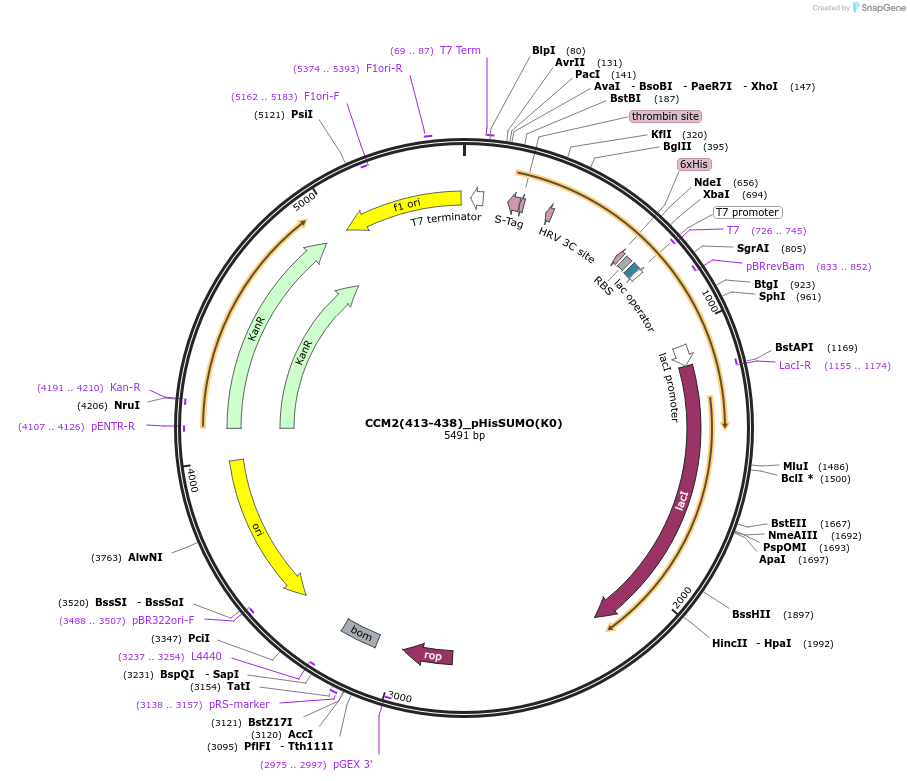
CCM2(413-438)_pHisSUMO(K0)
Plasmid#224065PurposeBacterial expression of the C-terminal helix of CCM2 (residues 413-438) fused to an N-terminal hexa-histidine tag and SUMO(K0) [all lysines in SUMO mutated to arginines]. HRV-3C cleavage siteDepositorAvailable SinceAug. 26, 2024AvailabilityIndustry, Academic Institutions, and Nonprofits -
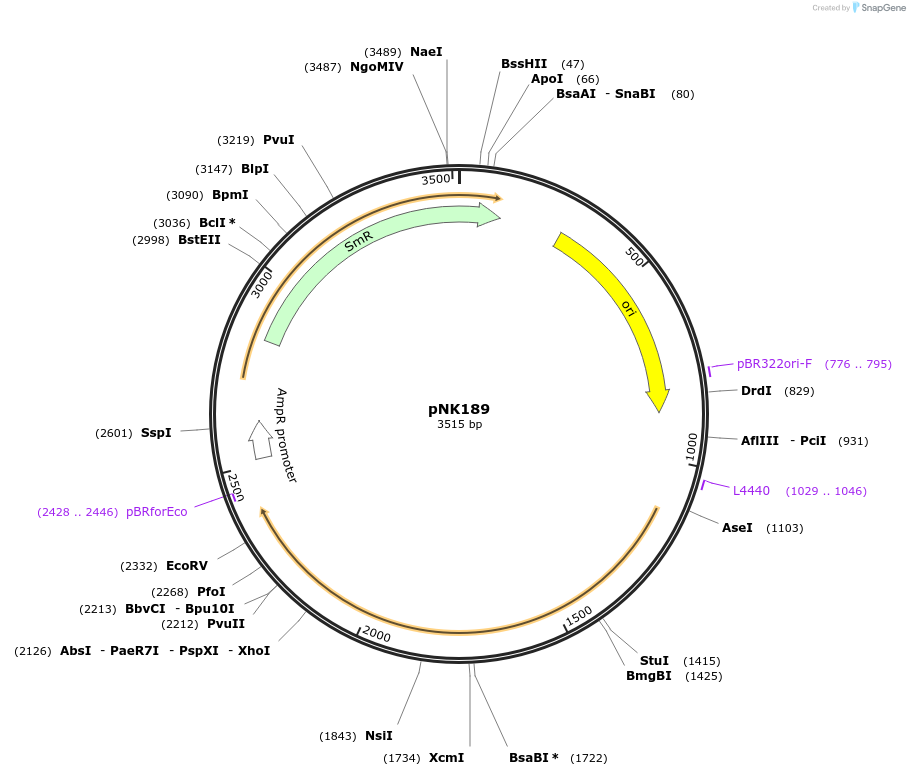
pNK189
Plasmid#219743PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi hispidin-3-hydroxylase nnH3H_v2 codon-optimised for expression in Homo sapiensDepositorInsertmutant of fungal hispidin-3-hydroxylase
UseSynthetic BiologyMutationD37E, V181I, S323M, M385KAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
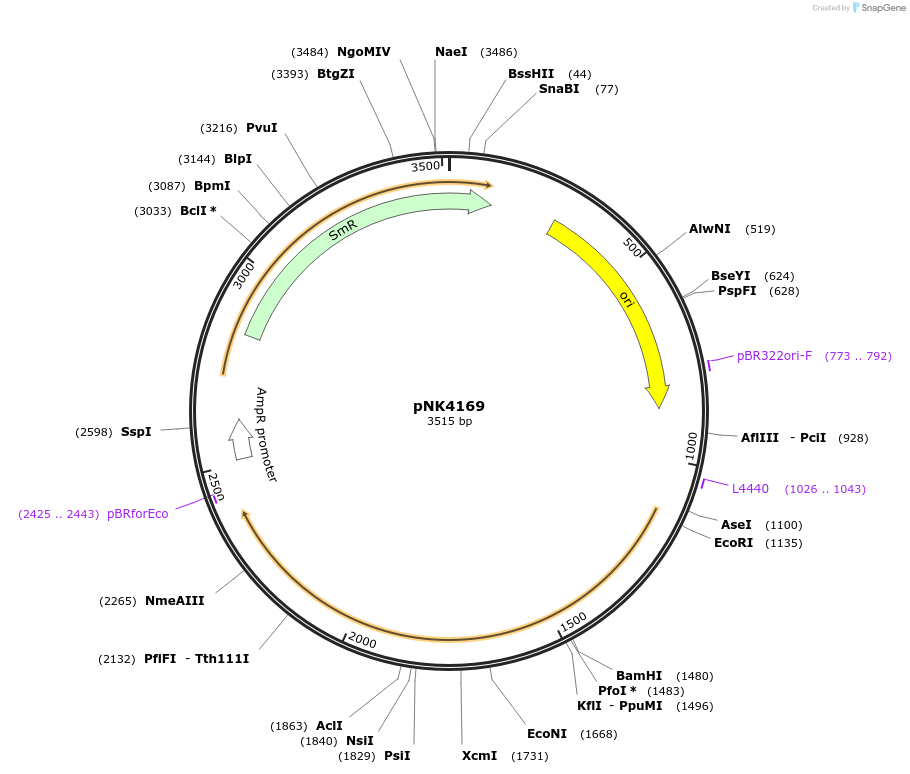
pNK4169
Plasmid#219744PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi hispidin-3-hydroxylase nnH3H_v2 codon-optimised for expression in Pichia pastorisDepositorInsertmutant of fungal hispidin-3-hydroxylase
UseSynthetic BiologyMutationD37E, V181I, S323M, M385KAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only