We narrowed to 41,100 results for: MUT
-
Plasmid#180425Purposemammalian expression of human SEPT7 Gmut2 fused to monomeric superfolder GFPDepositorAvailable SinceDec. 1, 2022AvailabilityAcademic Institutions and Nonprofits only
-
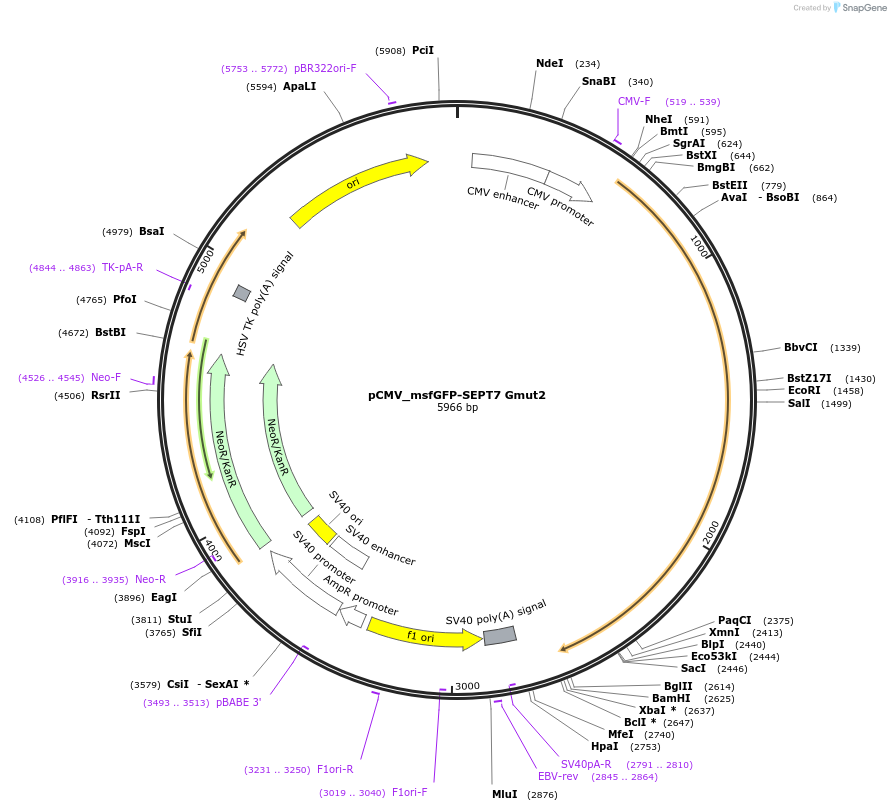
pCMV_msfGFP-SEPT7 Gmut2
Plasmid#180425Purposemammalian expression of human SEPT7 Gmut2 fused to monomeric superfolder GFPDepositorAvailable SinceDec. 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
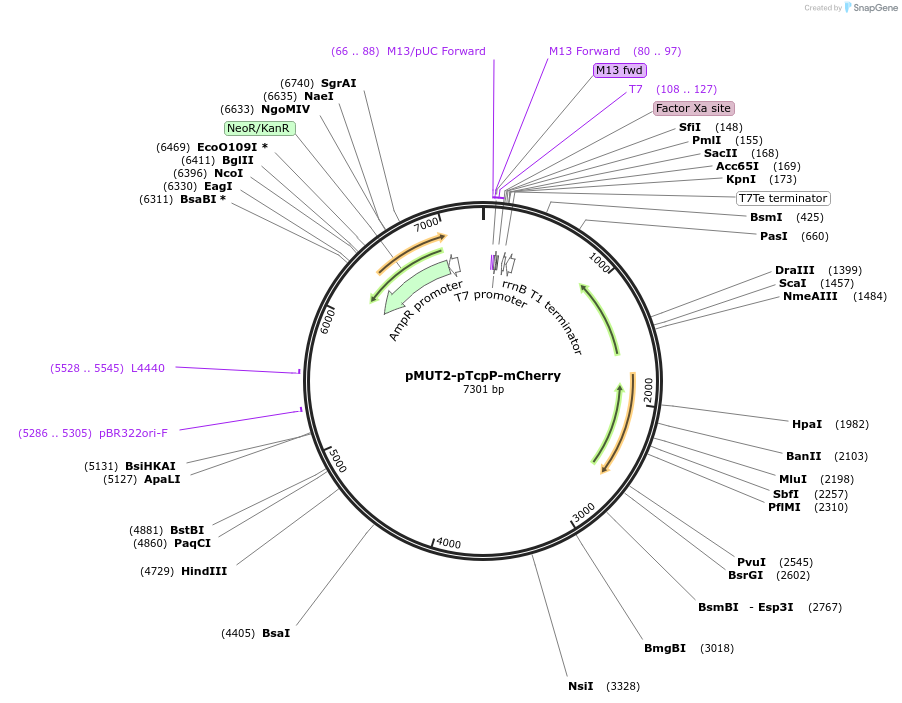
pMUT2-pTcpP-mCherry
Plasmid#192861PurposepMUT2 based vector for sensing human primary bile acids in E. coli Nissle 1917DepositorInsertTcpP-CadC fusion protein and cognate promoter inducible by human primary bile acids
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pMUT2-pTcpP-mCherry
Plasmid#192861PurposepMUT2 based vector for sensing human primary bile acids in E. coli Nissle 1917DepositorInsertTcpP-CadC fusion protein and cognate promoter inducible by human primary bile acids
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
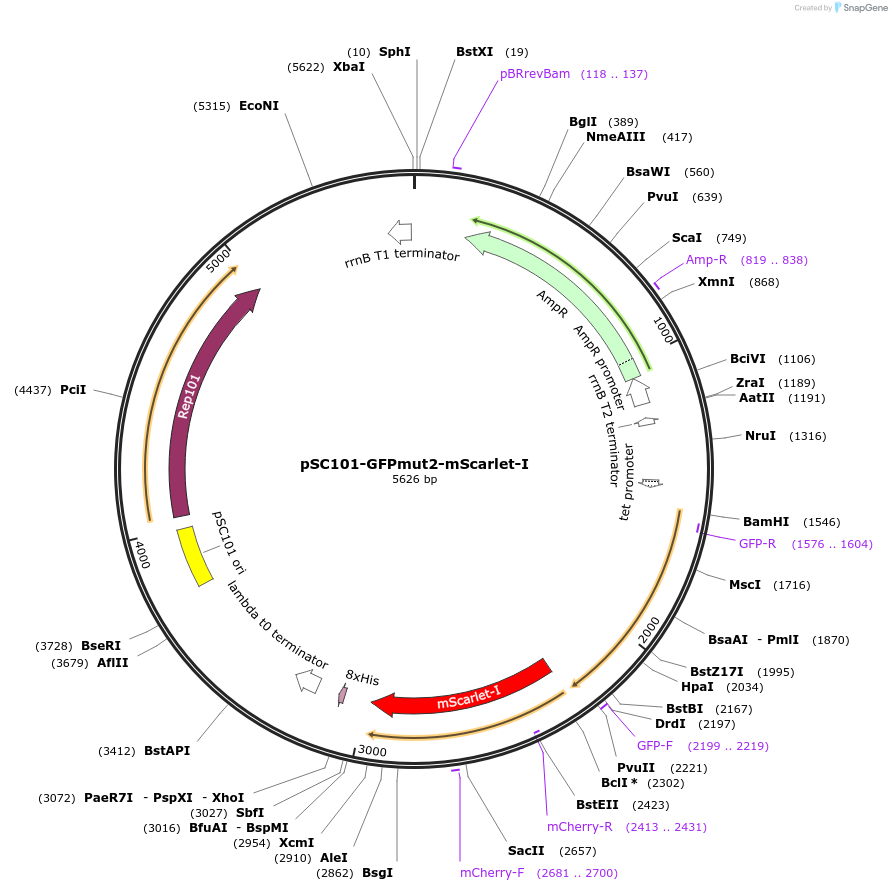
pSC101-GFPmut2-mScarlet-I
Plasmid#208183Purposemistranslation GFP and mScarlet-I positive controlDepositorInsertsGFPmut2
mScarlet-I
UseReporterExpressionBacterialPromoterPtet+dnaK P1Available SinceMarch 25, 2024AvailabilityAcademic Institutions and Nonprofits only -
pSC101-GFPmut2-mScarlet-I
Plasmid#208183Purposemistranslation GFP and mScarlet-I positive controlDepositorInsertsGFPmut2
mScarlet-I
UseReporterExpressionBacterialPromoterPtet+dnaK P1Available SinceMarch 25, 2024AvailabilityAcademic Institutions and Nonprofits only -
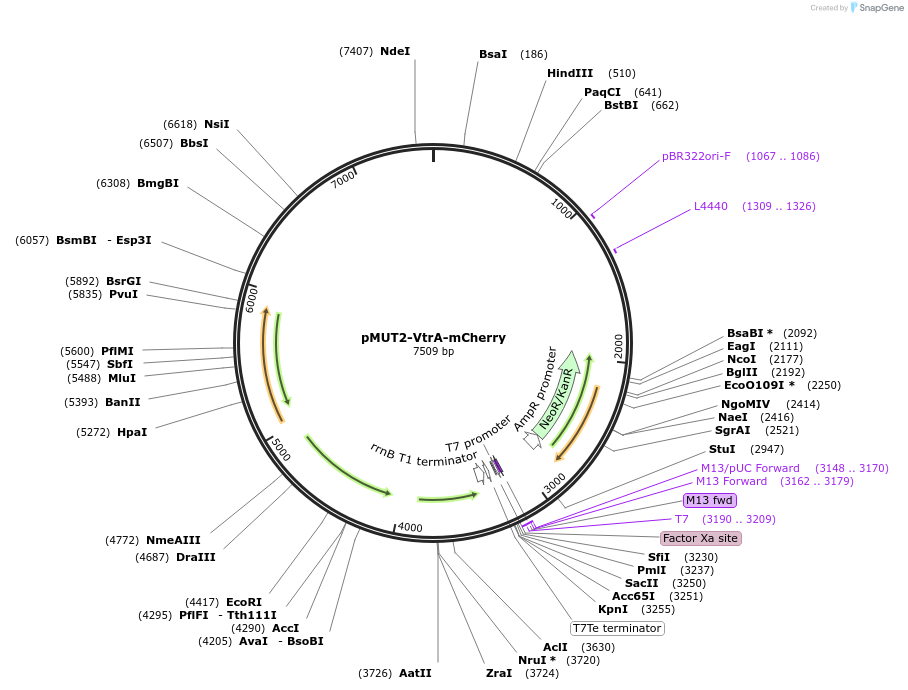
pMUT2-VtrA-mCherry
Plasmid#192862PurposepMUT2 based vector for sensing human secondary bile acids in E. coli Nissle 1917DepositorInsertVtrA-CadC fusion protein and cognate promoter inducible by secondary bile acids
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pMUT2-VtrA-mCherry
Plasmid#192862PurposepMUT2 based vector for sensing human secondary bile acids in E. coli Nissle 1917DepositorInsertVtrA-CadC fusion protein and cognate promoter inducible by secondary bile acids
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
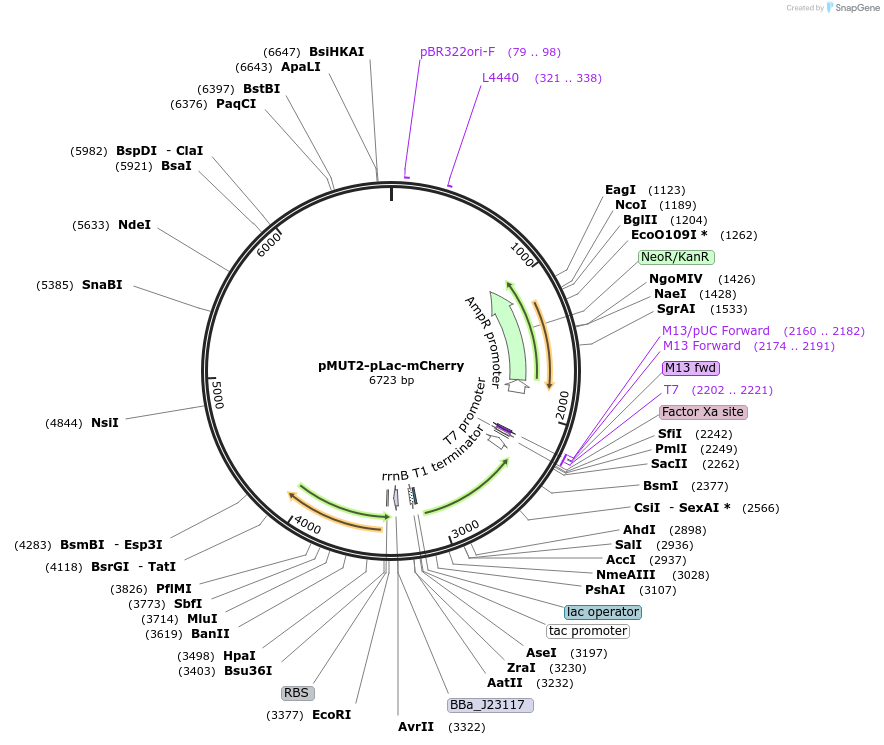
pMUT2-pLac-mCherry
Plasmid#192859PurposepMUT2 based vector for sensing lactic acid in E. coli Nissle 1917DepositorInsertlldR operon
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pMUT2-pLac-mCherry
Plasmid#192859PurposepMUT2 based vector for sensing lactic acid in E. coli Nissle 1917DepositorInsertlldR operon
ExpressionBacterialAvailable SinceApril 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
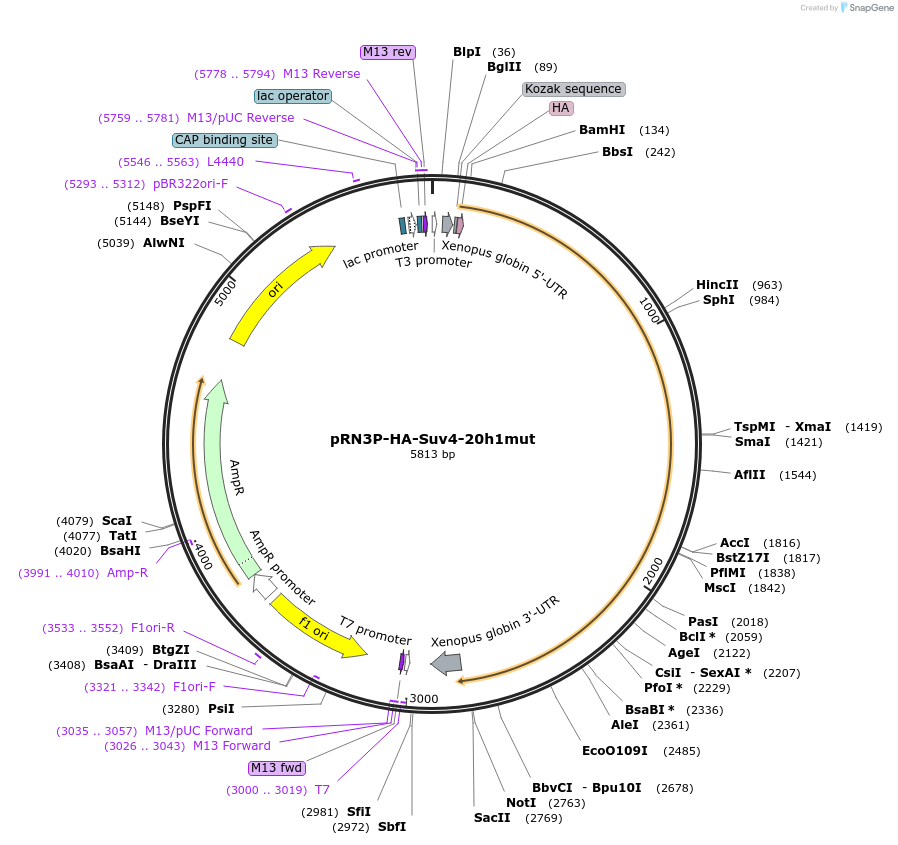
pRN3P-HA-Suv4-20h1mut
Plasmid#86690PurposeFor transcription of of Suv4-20h1mut mRNA preceded by an HA-tag in 5'DepositorInserthistone-lysine N-methyltransferase KMT5B mutant (Kmt5b Mouse)
TagsHAMutationmutated region NHDC (asparagine 273 to cysteine 2…PromoterT3Available SinceApril 26, 2017AvailabilityAcademic Institutions and Nonprofits only -
pRN3P-HA-Suv4-20h1mut
Plasmid#86690PurposeFor transcription of of Suv4-20h1mut mRNA preceded by an HA-tag in 5'DepositorInserthistone-lysine N-methyltransferase KMT5B mutant (Kmt5b Mouse)
TagsHAMutationmutated region NHDC (asparagine 273 to cysteine 2…PromoterT3Available SinceApril 26, 2017AvailabilityAcademic Institutions and Nonprofits only -
pgex-cIAP1 mut
Plasmid#8334DepositorAvailable SinceNov. 15, 2005AvailabilityAcademic Institutions and Nonprofits only -
pgex-cIAP1 mut
Plasmid#8334DepositorAvailable SinceNov. 15, 2005AvailabilityAcademic Institutions and Nonprofits only -
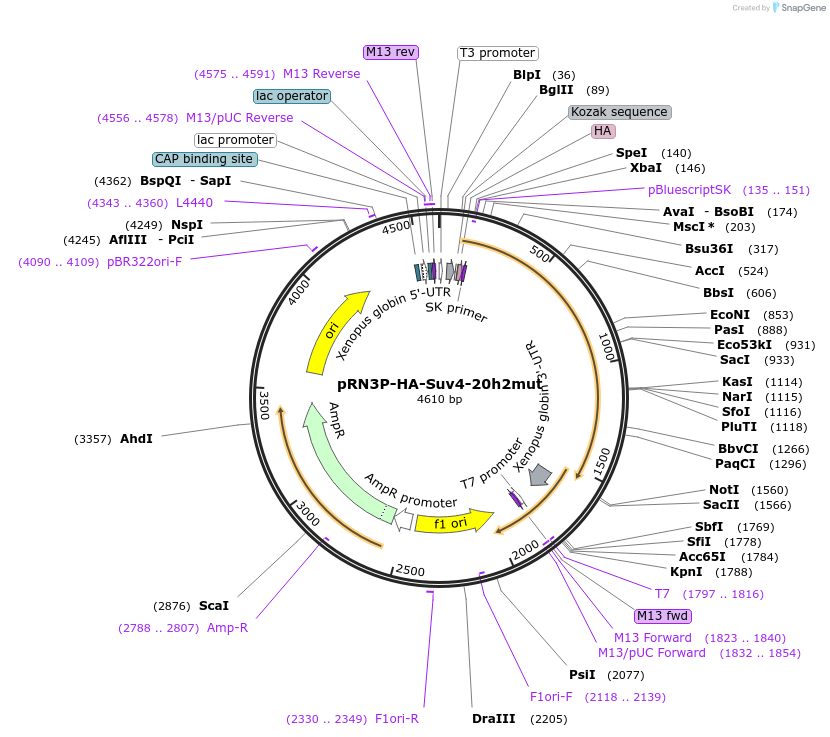
pRN3P-HA-Suv4-20h2mut
Plasmid#86692PurposeFor transcription of of Suv4-20h2mut mRNA preceded by an HA-tag in 5'DepositorInserthistone-lysine N-methyltransferase KMT5C mutant (Kmt5c Mouse)
TagsHAMutationMutated asparagine 273 to cysteine 276 (NHDC) int…PromoterT3Available SinceMarch 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
pRN3P-HA-Suv4-20h2mut
Plasmid#86692PurposeFor transcription of of Suv4-20h2mut mRNA preceded by an HA-tag in 5'DepositorInserthistone-lysine N-methyltransferase KMT5C mutant (Kmt5c Mouse)
TagsHAMutationMutated asparagine 273 to cysteine 276 (NHDC) int…PromoterT3Available SinceMarch 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
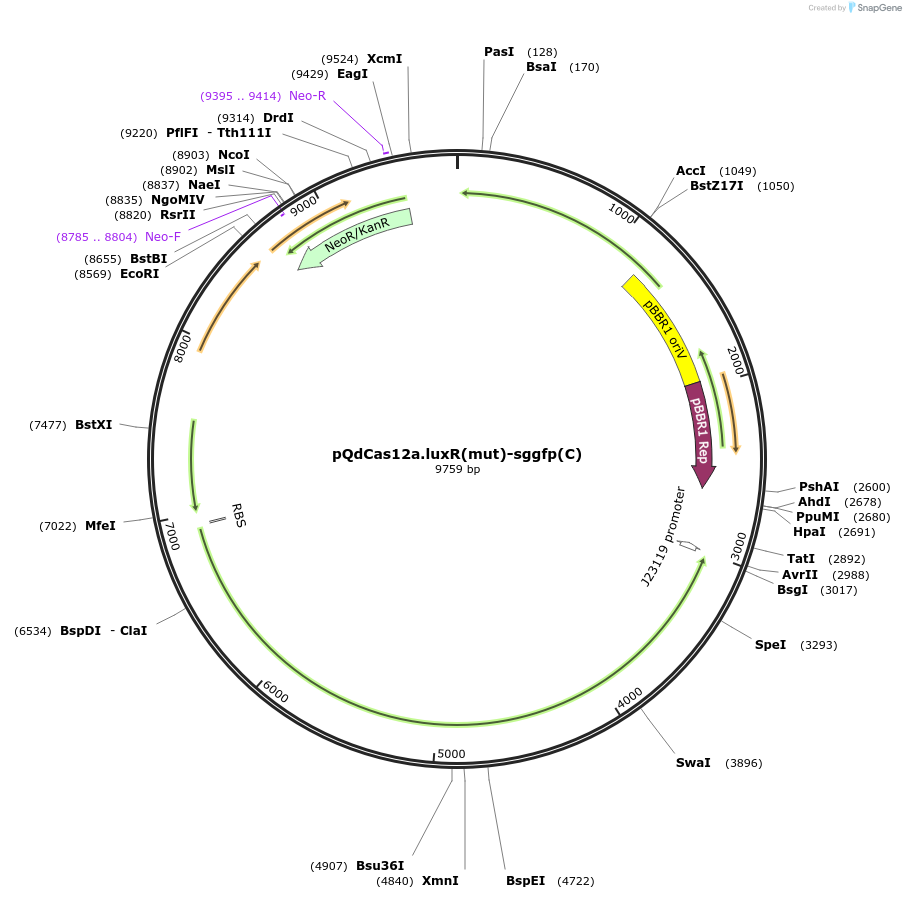
pQdCas12a.luxR(mut)-sggfp(C)
Plasmid#236043PurposeThe plasmid pQdCas12a.luxR(mut)-sggfp(C) expresses the dCas12a endonuclease and the sgRNA (design C) targeting the gfp gene. Additionally, it contains a mutation in the luxR gene.DepositorInsertquorum sensing cassette luxI, mutated luxR, and the LuxR-dependent promoter pLuxI , dCas12a and sgRNA design (C) of gfp gene
UseCRISPRPromoterquorum sensing promoterAvailable SinceMay 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
pQdCas12a.luxR(mut)-sggfp(C)
Plasmid#236043PurposeThe plasmid pQdCas12a.luxR(mut)-sggfp(C) expresses the dCas12a endonuclease and the sgRNA (design C) targeting the gfp gene. Additionally, it contains a mutation in the luxR gene.DepositorInsertquorum sensing cassette luxI, mutated luxR, and the LuxR-dependent promoter pLuxI , dCas12a and sgRNA design (C) of gfp gene
UseCRISPRPromoterquorum sensing promoterAvailable SinceMay 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
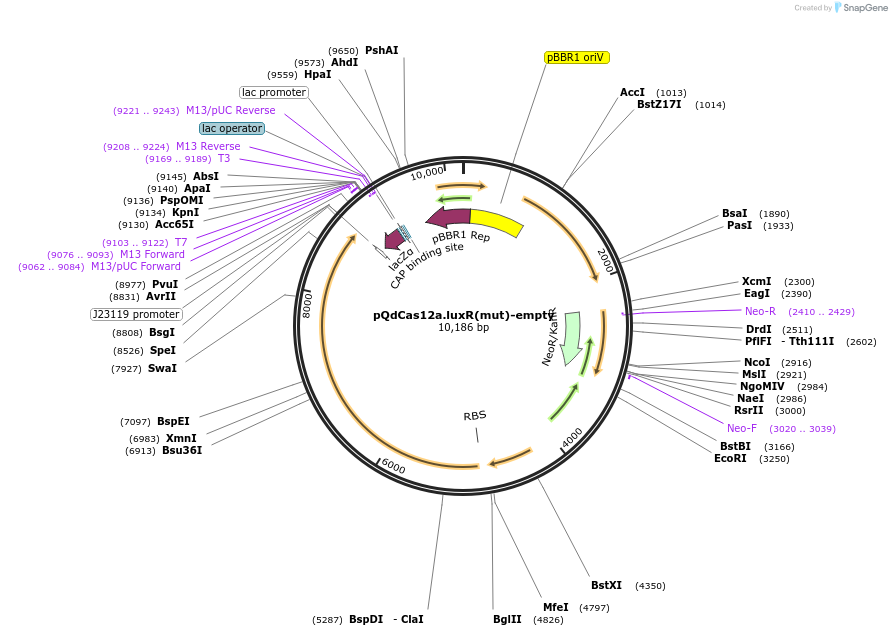
pQdCas12a.luxR(mut)-empty
Plasmid#236037PurposeThe plasmid pQdCas12a.luxR(mut)-empty expresses the dCas12a endonuclease. Additionally, it contains a mutation in the luxR gene.DepositorInsertquorum sensing cassette luxI, mutated luxR, and the LuxR-dependent promoter pLuxI, dcas12a, and lacZ-alpha (fragment).
UseCRISPRPromoterquorum sensing promoterAvailable SinceApril 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
pQdCas12a.luxR(mut)-empty
Plasmid#236037PurposeThe plasmid pQdCas12a.luxR(mut)-empty expresses the dCas12a endonuclease. Additionally, it contains a mutation in the luxR gene.DepositorInsertquorum sensing cassette luxI, mutated luxR, and the LuxR-dependent promoter pLuxI, dcas12a, and lacZ-alpha (fragment).
UseCRISPRPromoterquorum sensing promoterAvailable SinceApril 24, 2025AvailabilityAcademic Institutions and Nonprofits only