4,625 results
-
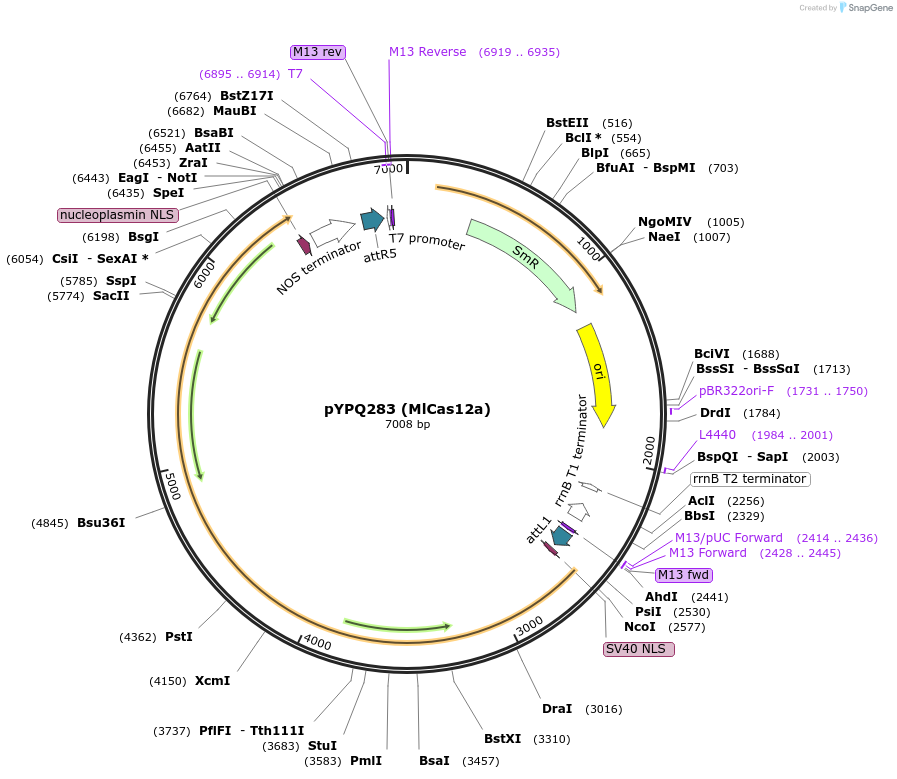
Plasmid#138115PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized MlCas12a without promoterDepositorInsertMlCas12a
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
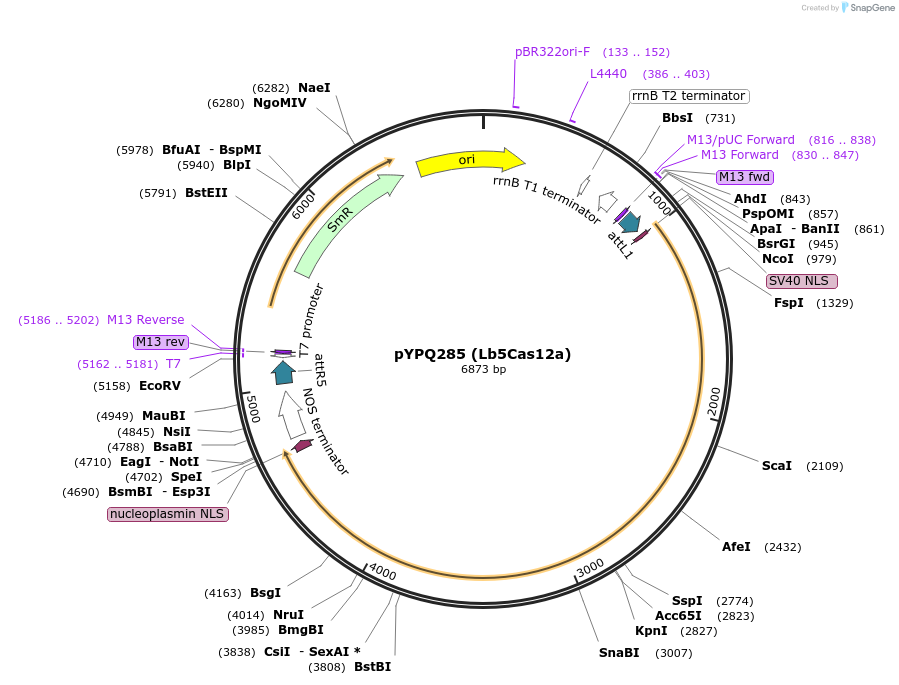
pYPQ285 (Lb5Cas12a)
Plasmid#138120PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized Lb5Cas12a without promoterDepositorInsertLb5Cas12a
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
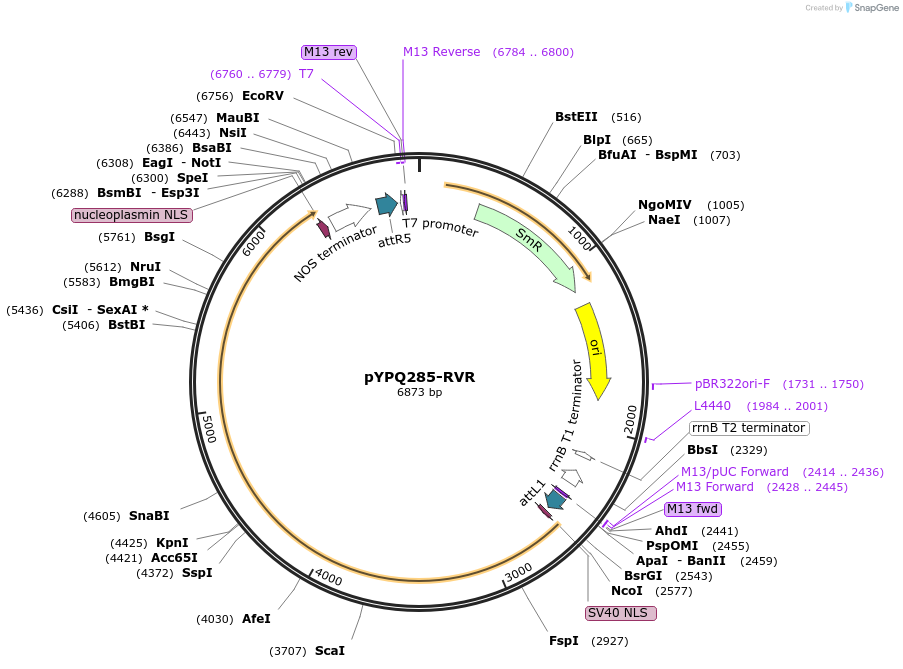
pYPQ285-RVR
Plasmid#138121PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized Lb5Cas12a with N512R, K518V and N522R mutations, without promoterDepositorInsertLb5Cas12a RVR variant
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantMutationN512R, K518V and N522RAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
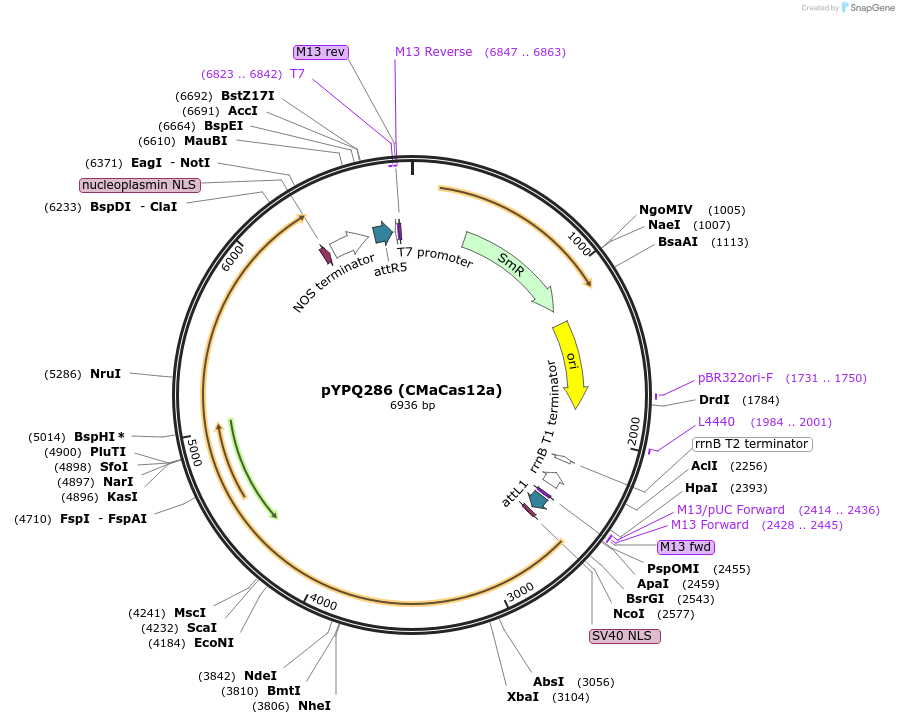
pYPQ286 (CMaCas12a)
Plasmid#138122PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized CMaCas12a without promoterDepositorInsertCMaCas12a
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
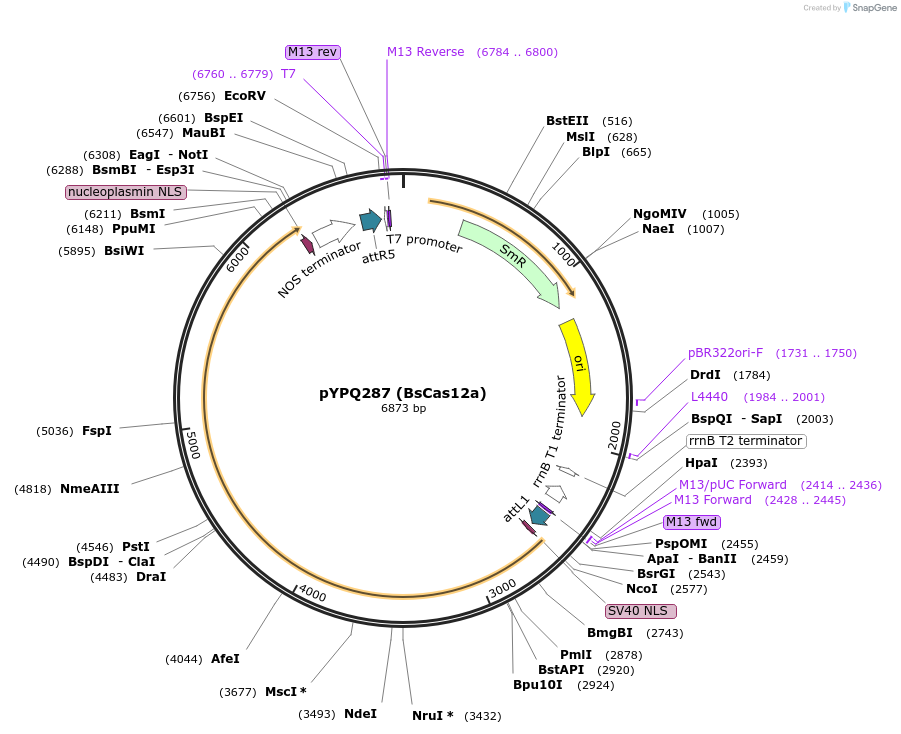
pYPQ287 (BsCas12a)
Plasmid#138123PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized BsCas12a without promoterDepositorInsertBsCas12a
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
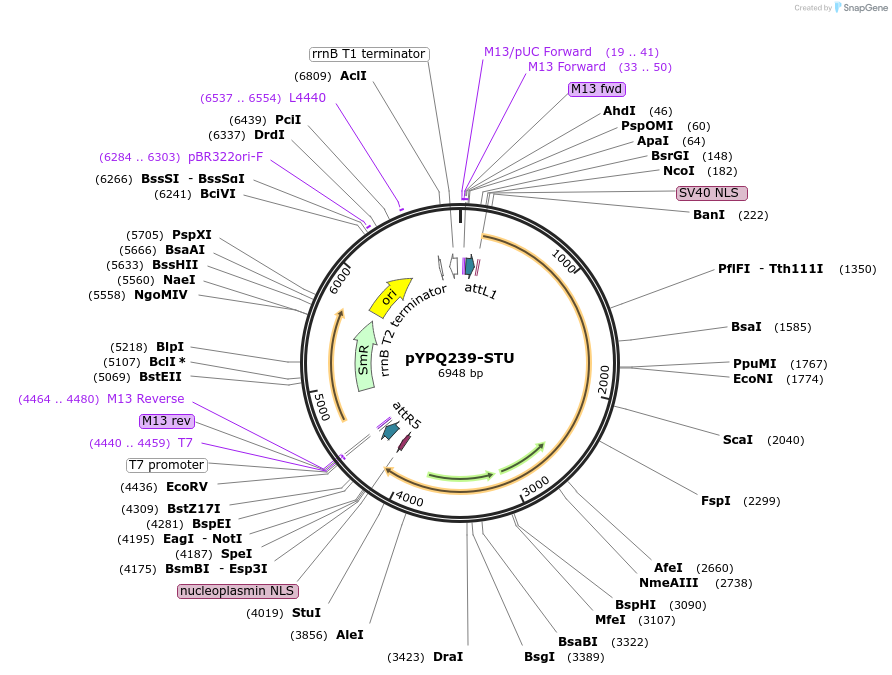
pYPQ239-STU
Plasmid#138112PurposeGateway entry plasmid (attL1 & attR5) expressing rice codon optimized FnCas12a without promoter and terminatorDepositorInsertFnCas12a
UseCRISPR; Gateway compatible cas12a entry cloneTagsNLSExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
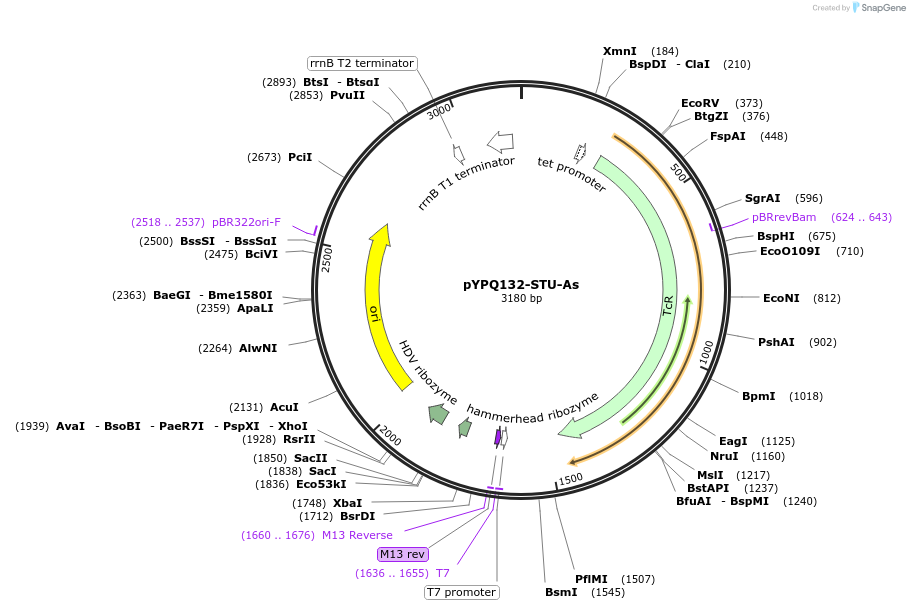
pYPQ132-STU-As
Plasmid#138097PurposeGolden Gate entry vector for 2nd crRNA cloning with AsCas12a crRNA scaffold flanked by HH and HDV ribozymesDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
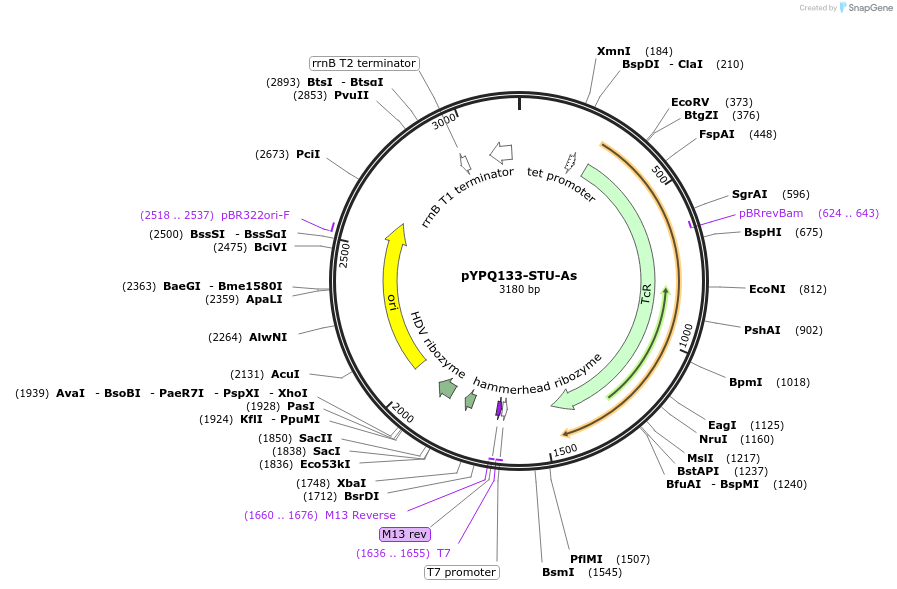
pYPQ133-STU-As
Plasmid#138100PurposeGolden Gate entry vector for 3rd crRNA cloning with AsCas12a crRNA scaffold flanked by HH and HDV ribozymesDepositorTypeEmpty backboneUseCRISPRExpressionPlantAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
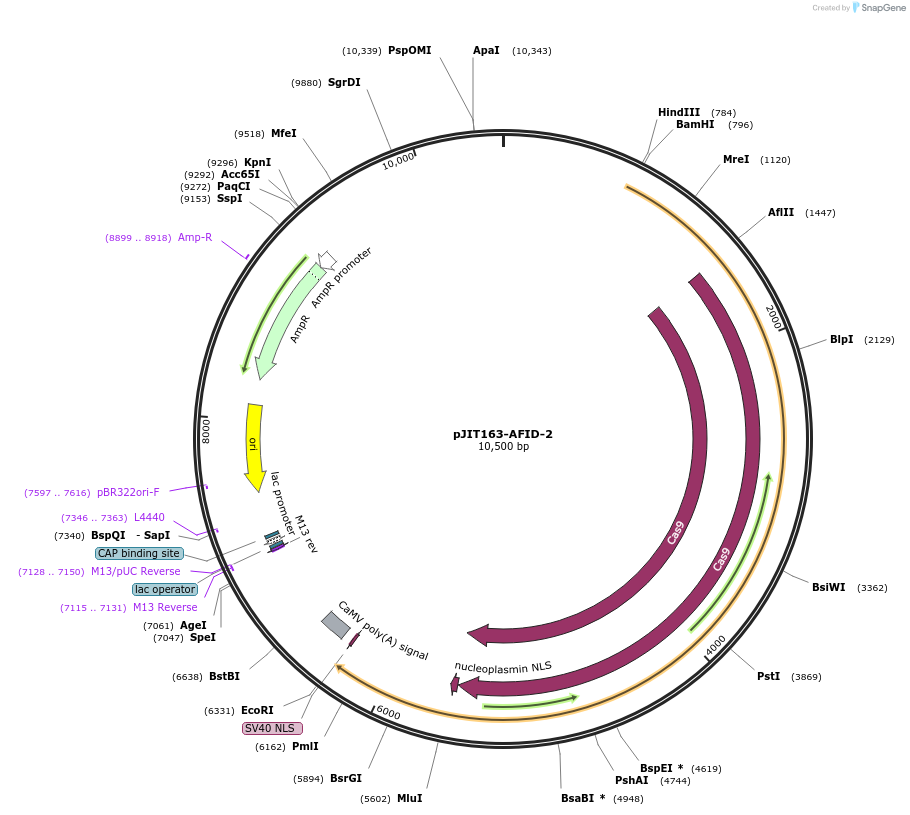
pJIT163-AFID-2
Plasmid#167358PurposeFor inducing predictable deletions in protoplasts and wheat plantsDepositorInsertA3A-Cas9-UDG
ExpressionPlantMutationWTAvailable SinceMarch 31, 2021AvailabilityAcademic Institutions and Nonprofits only -
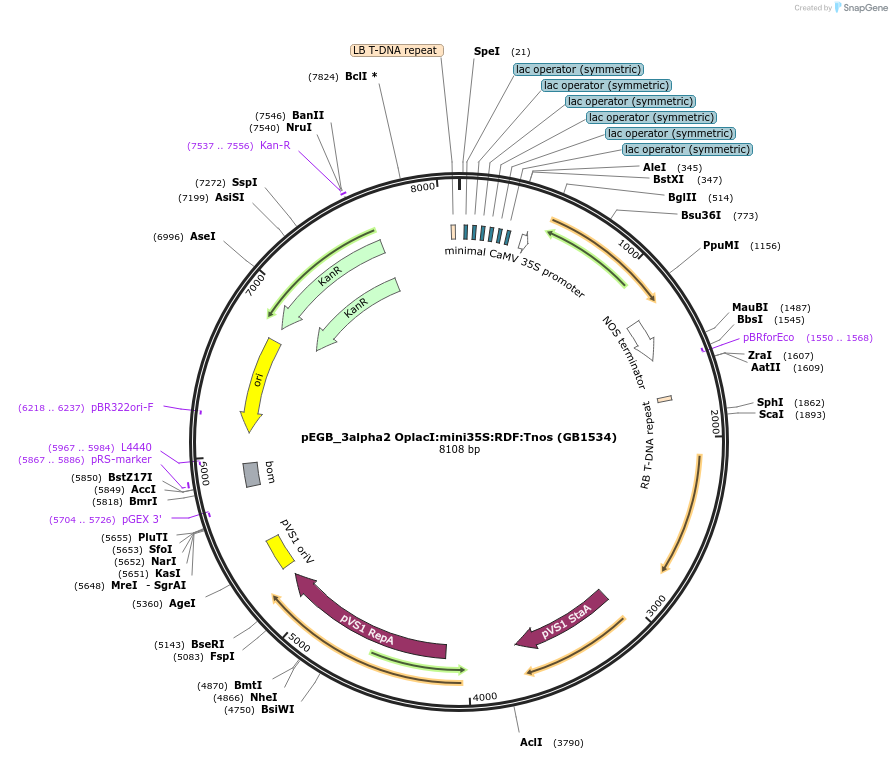
pEGB_3alpha2 OplacI:mini35S:RDF:Tnos (GB1534)
Plasmid#160617PurposeTU for the regulated expression of the PhiC31 phage recombination directionality factor (RDF) gene driven by a synthetic promoter consisting of the LacI operator fused to the minimal 35S promoter.DepositorInsertRDF
UseSynthetic BiologyExpressionPlantMutationBsaI and BsmBI sites removedPromotermini35sAvailable SinceMarch 25, 2021AvailabilityAcademic Institutions and Nonprofits only -
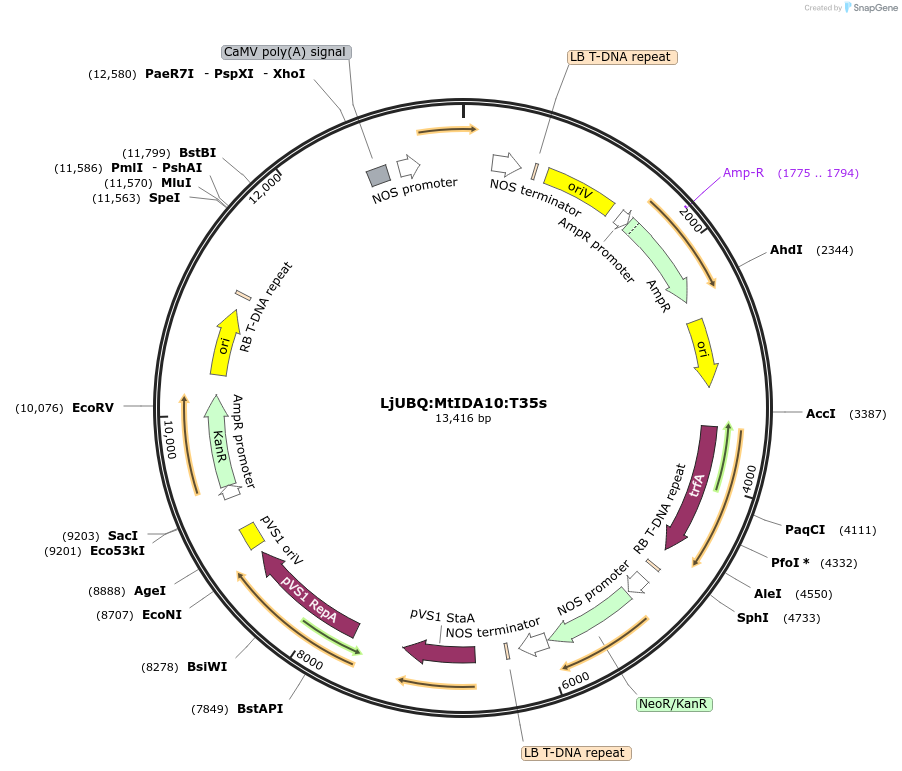
LjUBQ:MtIDA10:T35s
Plasmid#164743PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr3g094130.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 25, 2021AvailabilityAcademic Institutions and Nonprofits only -
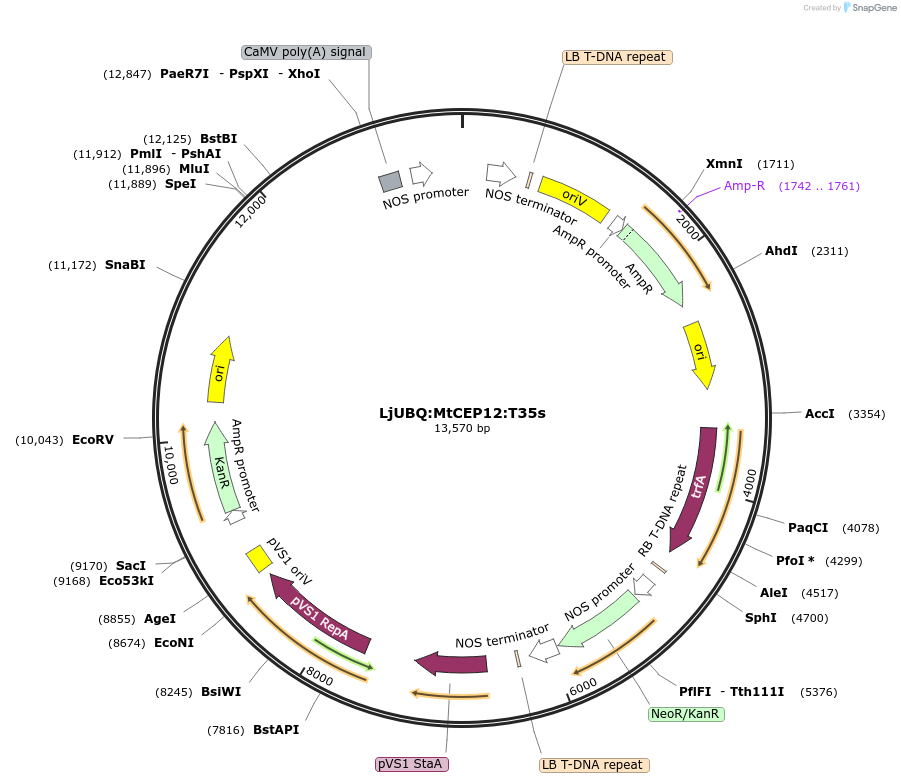
LjUBQ:MtCEP12:T35s
Plasmid#164730PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr1g020020.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 25, 2021AvailabilityAcademic Institutions and Nonprofits only -
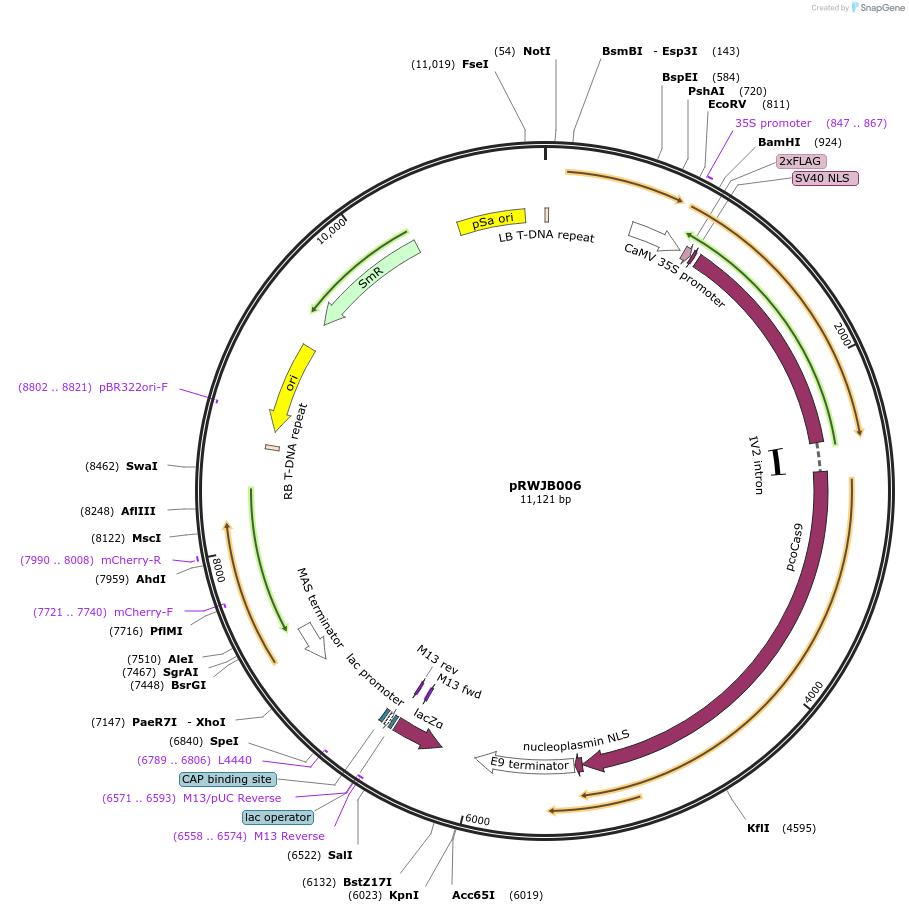
pRWJB006
Plasmid#160113PurposeDestination vector expressing plant-codon-optimized Cas9 under 35S promoter (BamHI and NotI), with sgRNAs be shuffled in; seed coat specific red fluorescence for screening trangene freeDepositorTypeEmpty backboneUseCRISPRExpressionBacterial and PlantPromoter35SAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
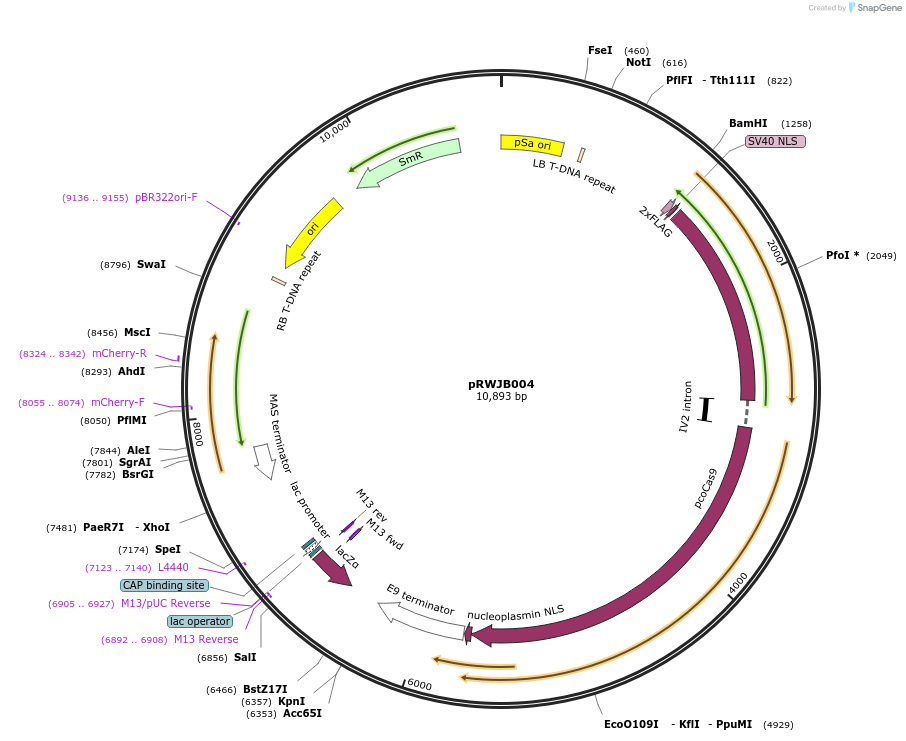
pRWJB004
Plasmid#160112PurposeDestination vector expressing plant-codon-optimized Cas9 under UBQ10 promoter (BamHI and NotI), with gRNAs be shuffled in; seed coat specific red fluorescence for screening trangene free plants;DepositorTypeEmpty backboneUseCRISPRExpressionBacterial and PlantPromoterUBQ10Available SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
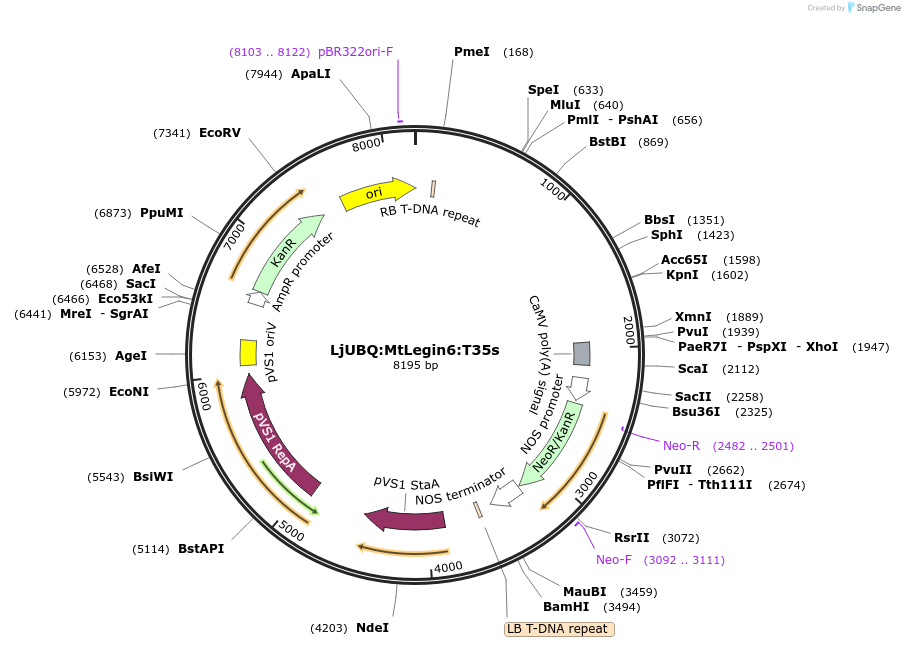
LjUBQ:MtLegin6:T35s
Plasmid#164751PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr3g067445.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
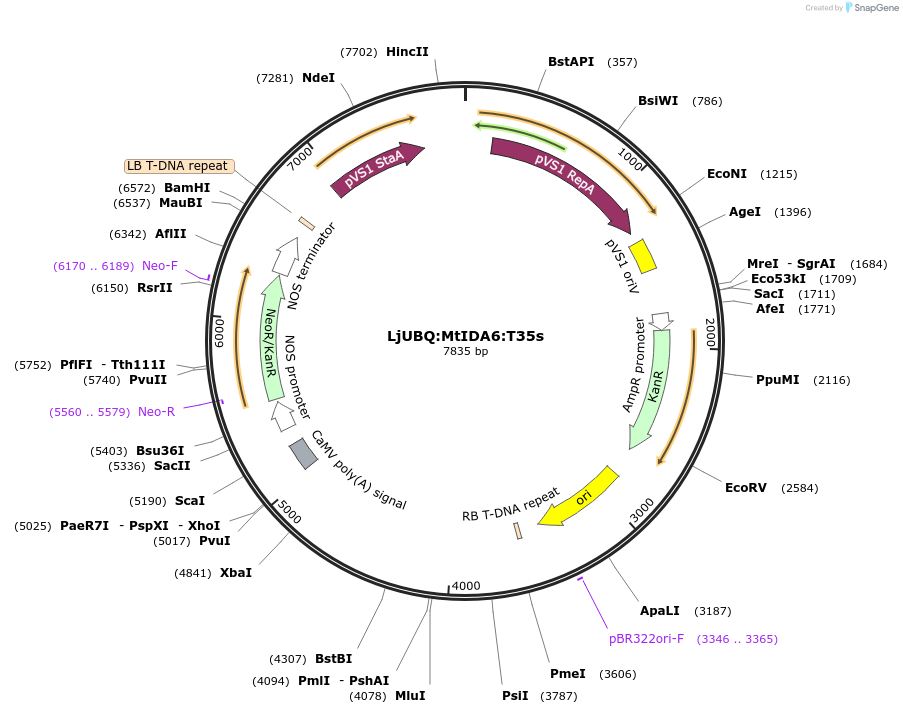
LjUBQ:MtIDA6:T35s
Plasmid#164749PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr1g099210.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
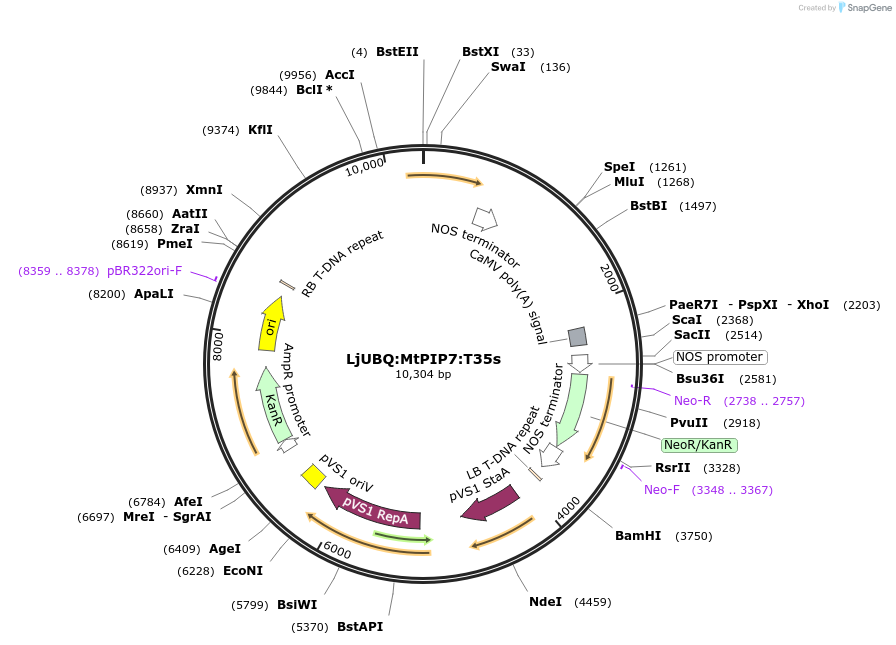
LjUBQ:MtPIP7:T35s
Plasmid#164766PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT4Noble_020581.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
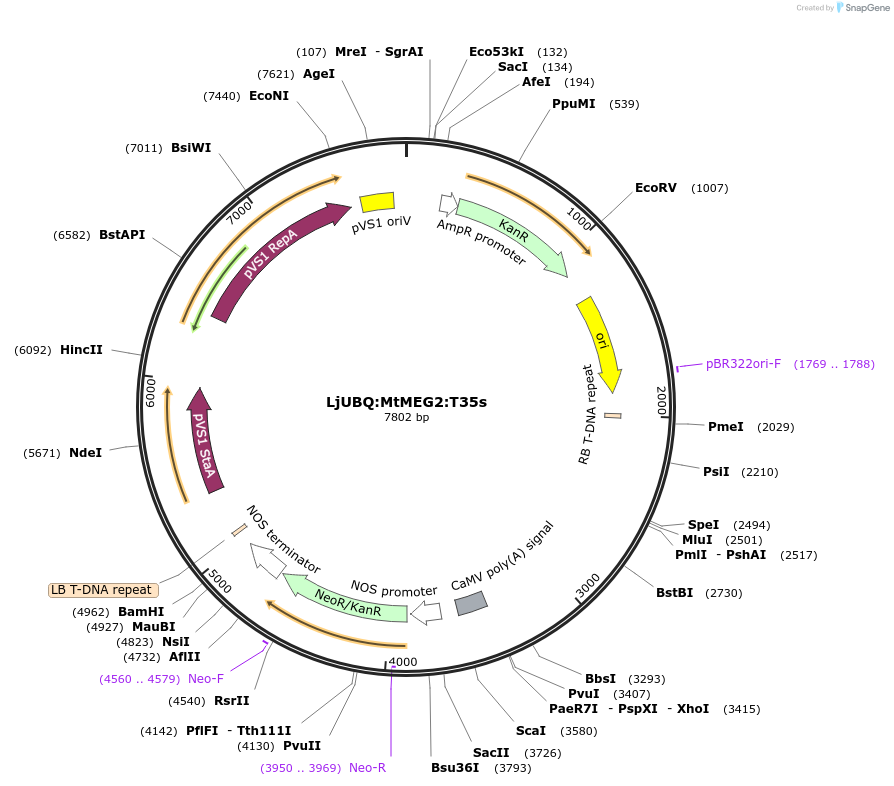
LjUBQ:MtMEG2:T35s
Plasmid#164754PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT4Noble_004794.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
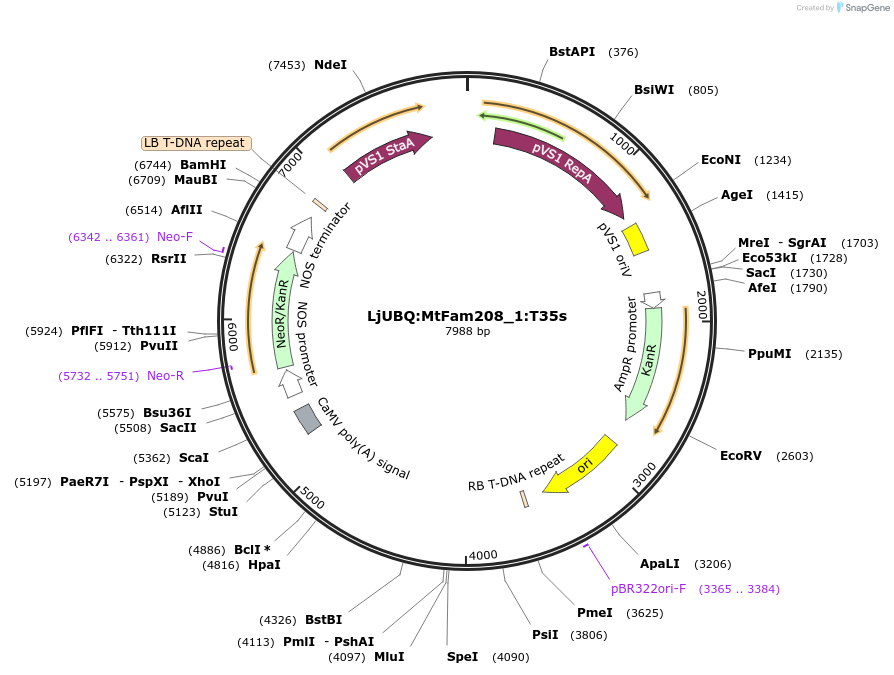
LjUBQ:MtFam208_1:T35s
Plasmid#164735PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr7g015340.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
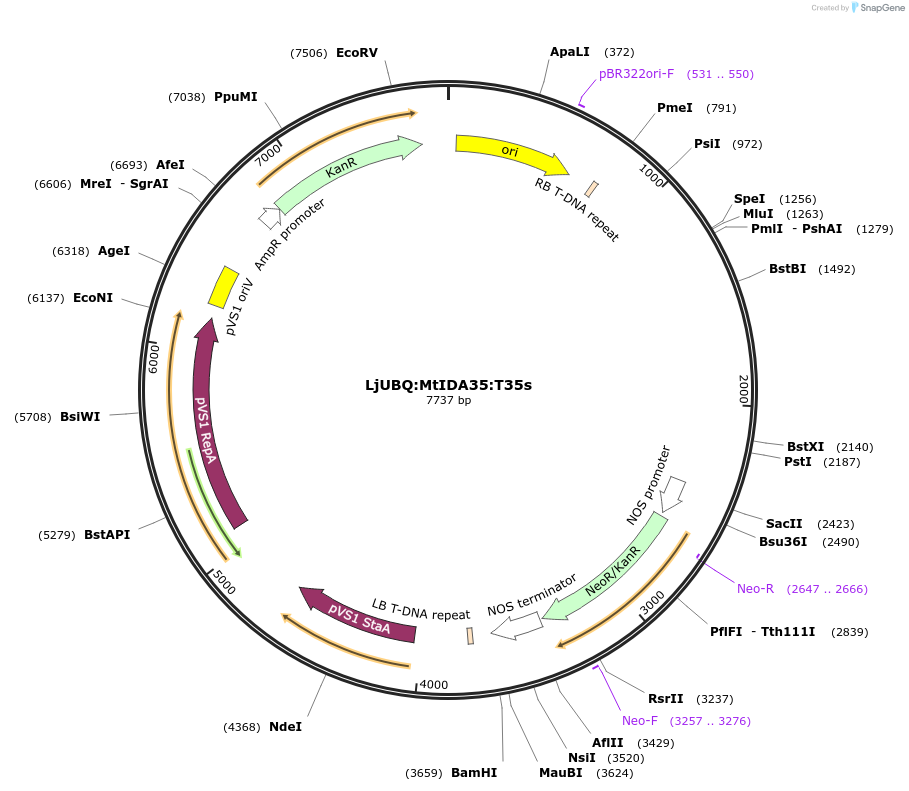
LjUBQ:MtIDA35:T35s
Plasmid#164747PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr7g115360.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
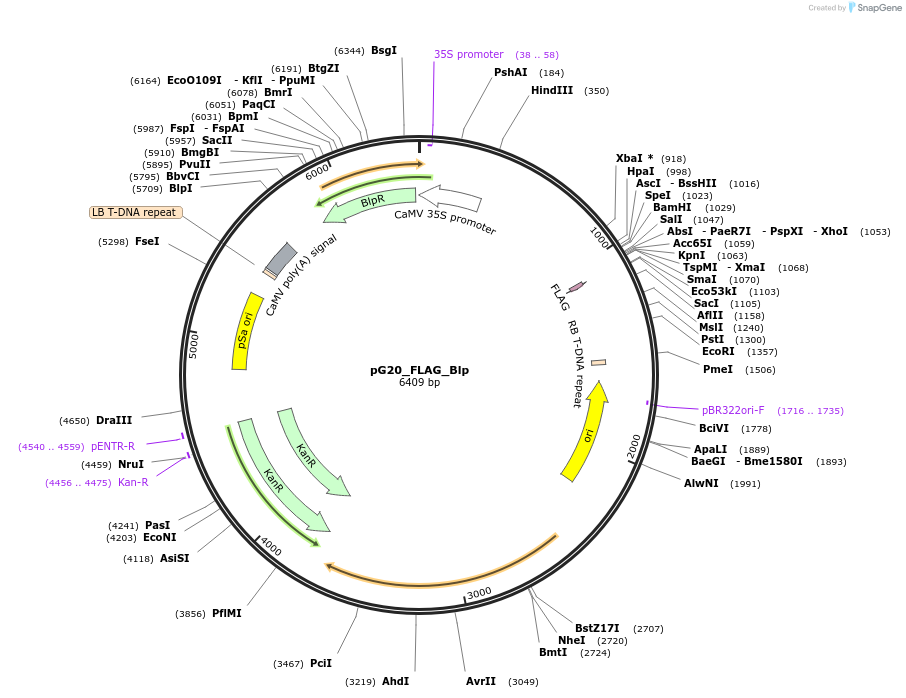
pG20_FLAG_Blp
Plasmid#159714PurposepGREEN-based cloning construct for expression in plantaDepositorTypeEmpty backboneTagsFLAGExpressionPlantAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
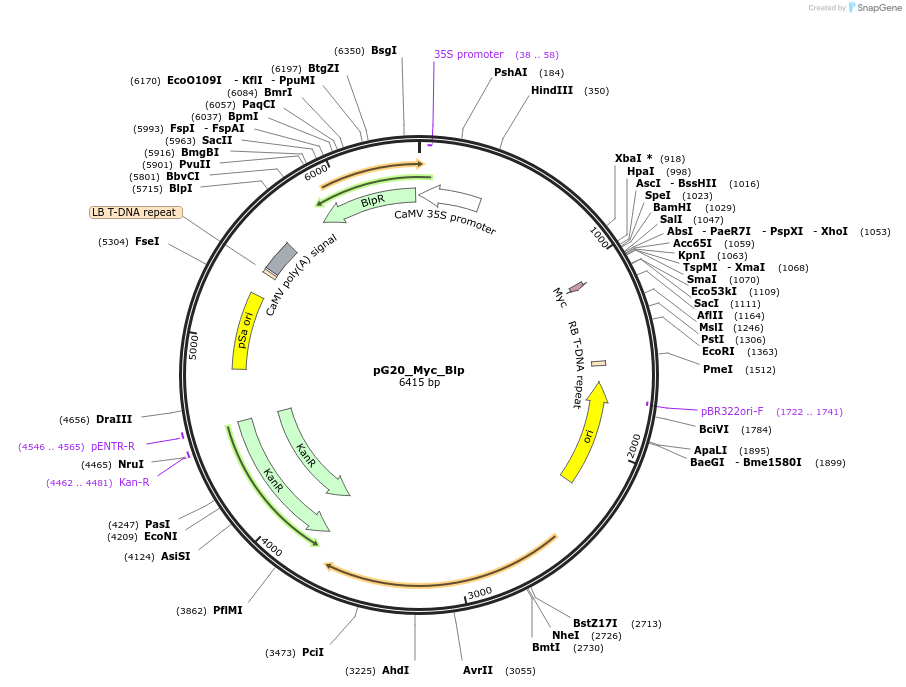
pG20_Myc_Blp
Plasmid#159716PurposepGREEN-based cloning construct for expression in plantaDepositorTypeEmpty backboneTagsMycExpressionPlantAvailable SinceFeb. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
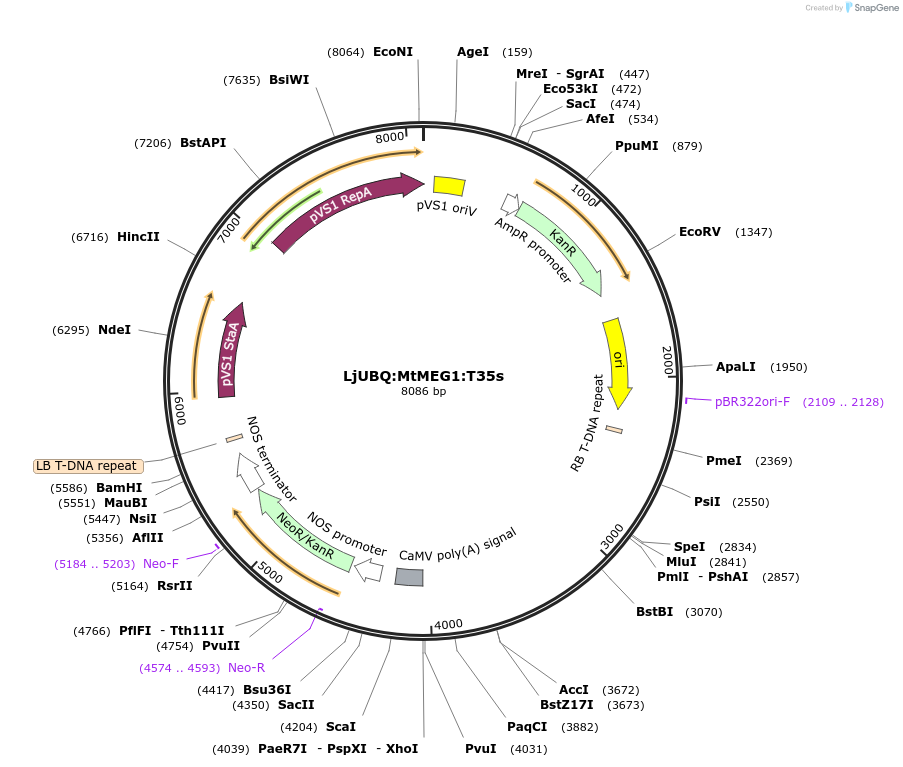
LjUBQ:MtMEG1:T35s
Plasmid#164753PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr7g079830.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 23, 2021AvailabilityAcademic Institutions and Nonprofits only -
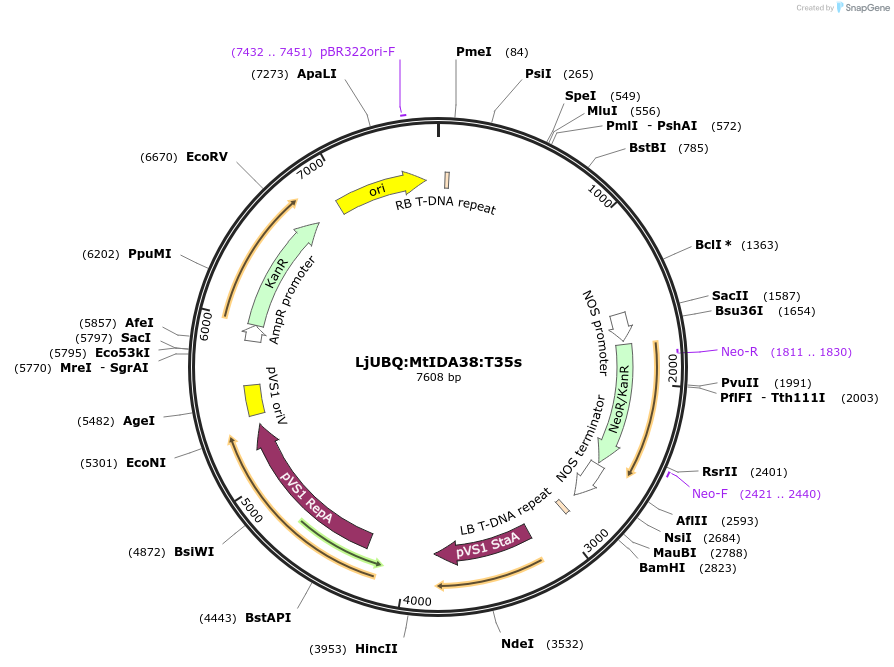
LjUBQ:MtIDA38:T35s
Plasmid#164748PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr7g096410.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 23, 2021AvailabilityAcademic Institutions and Nonprofits only -
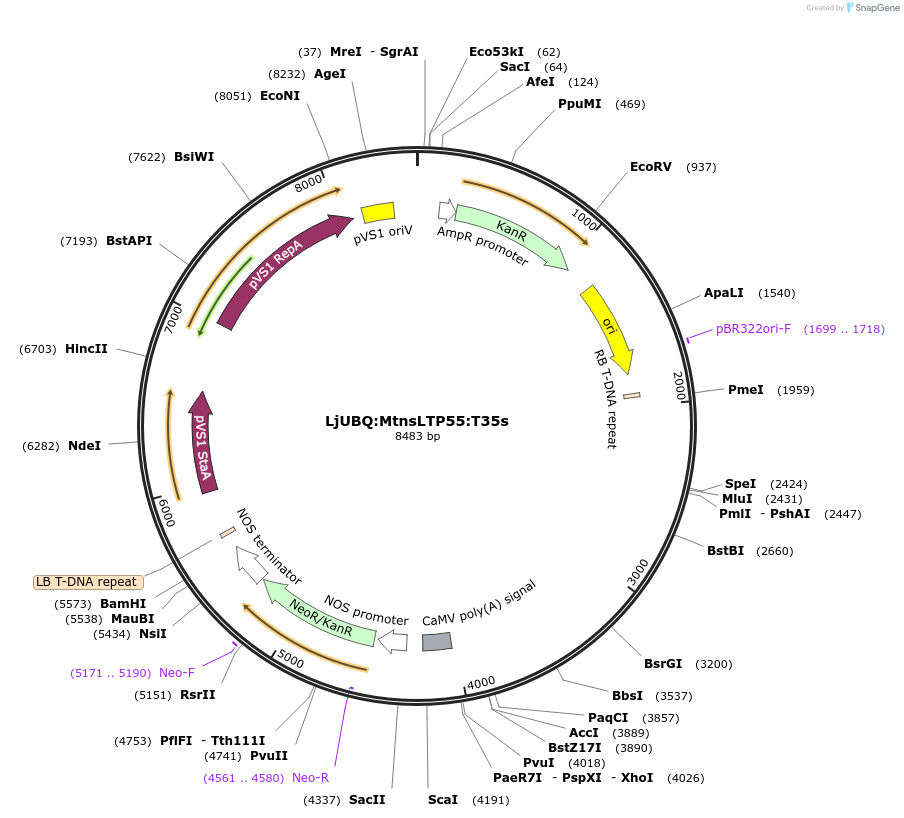
LjUBQ:MtnsLTP55:T35s
Plasmid#164759PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr1g071730.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
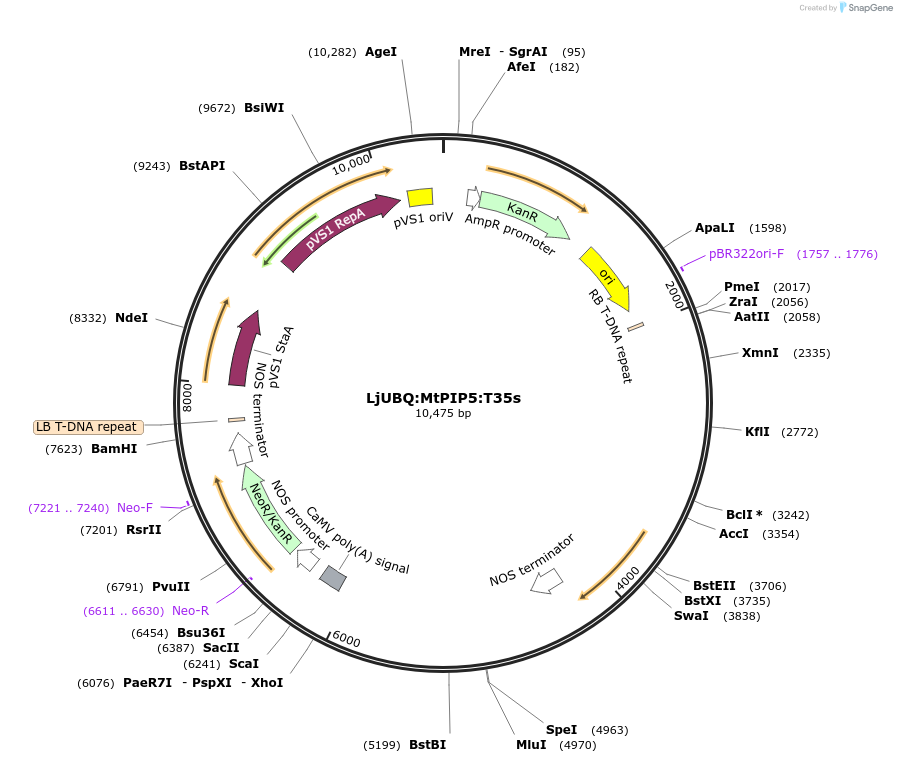
LjUBQ:MtPIP5:T35s
Plasmid#164764PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr4g095002.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
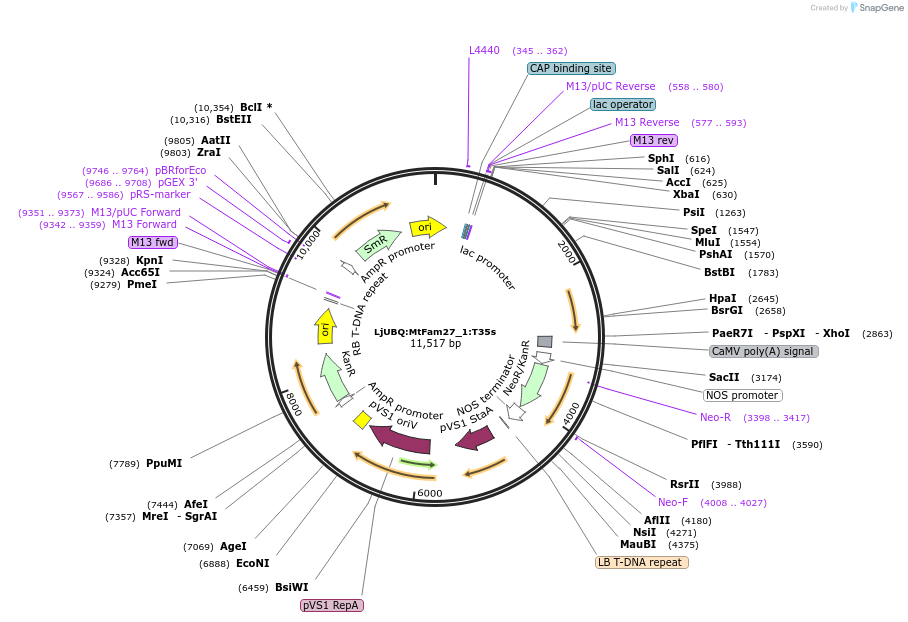
LjUBQ:MtFam27_1:T35s
Plasmid#164736PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr4g035905.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 18, 2021AvailabilityAcademic Institutions and Nonprofits only -
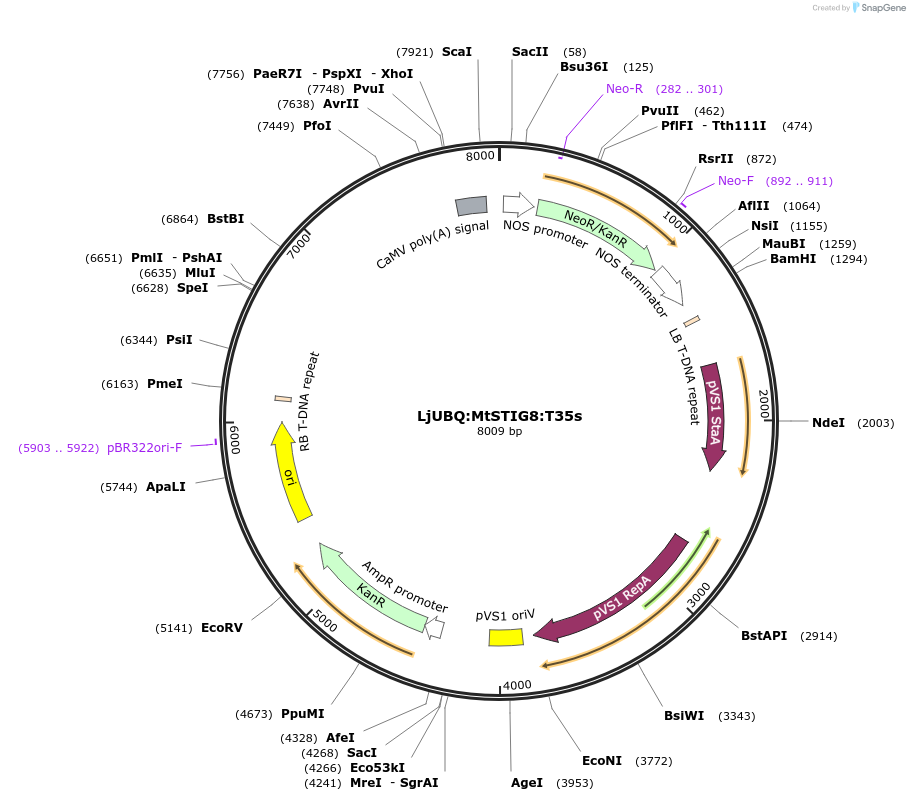
LjUBQ:MtSTIG8:T35s
Plasmid#164772PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr2g100600.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
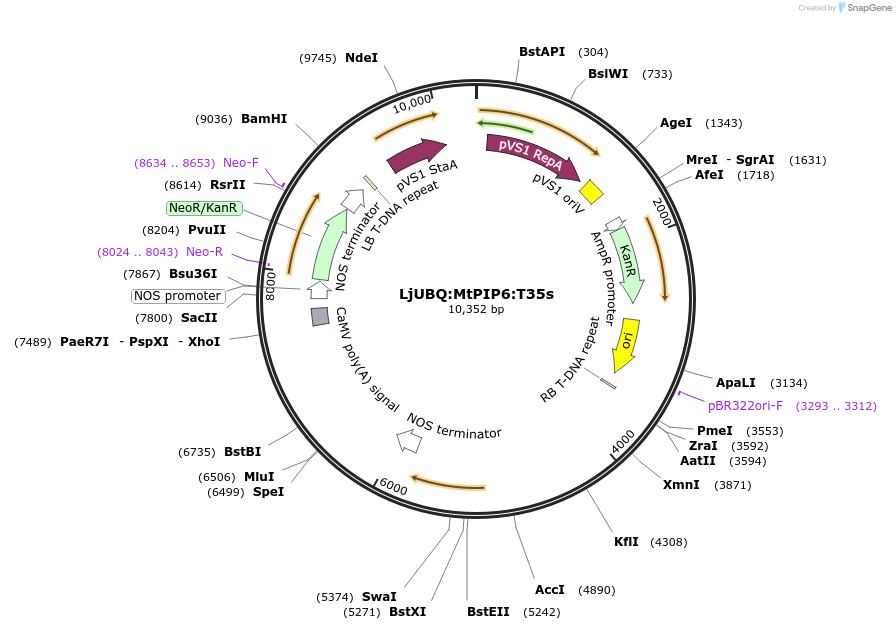
LjUBQ:MtPIP6:T35s
Plasmid#164765PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr5g004930.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
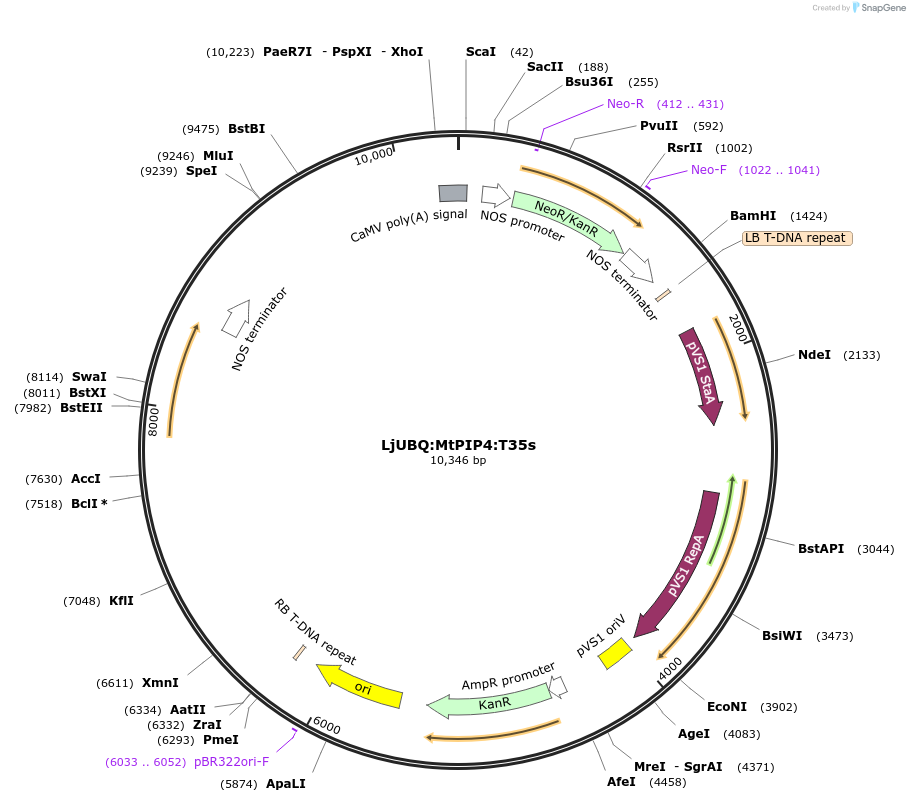
LjUBQ:MtPIP4:T35s
Plasmid#164763PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr5g016470.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
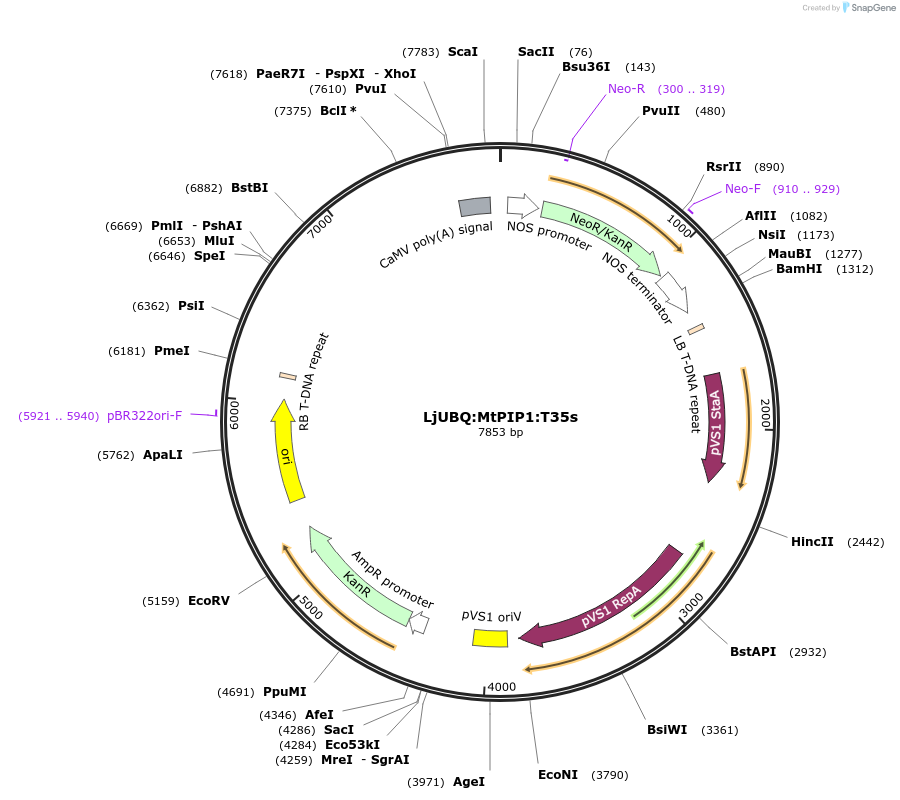
LjUBQ:MtPIP1:T35s
Plasmid#164761PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr4g068220.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
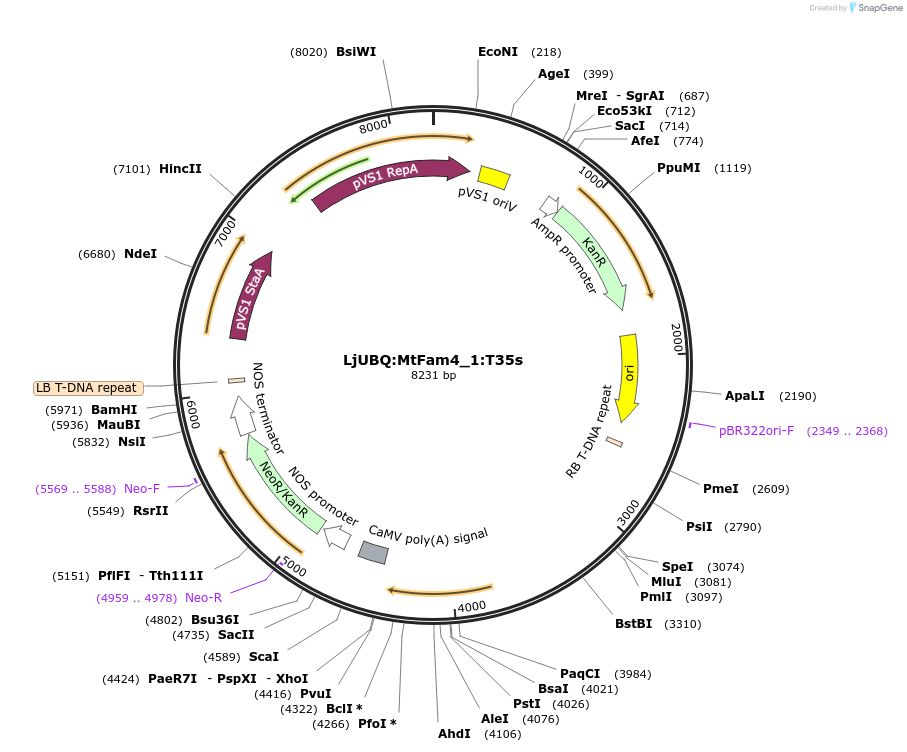
LjUBQ:MtFam4_1:T35s
Plasmid#164737PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMedtr8g020590.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
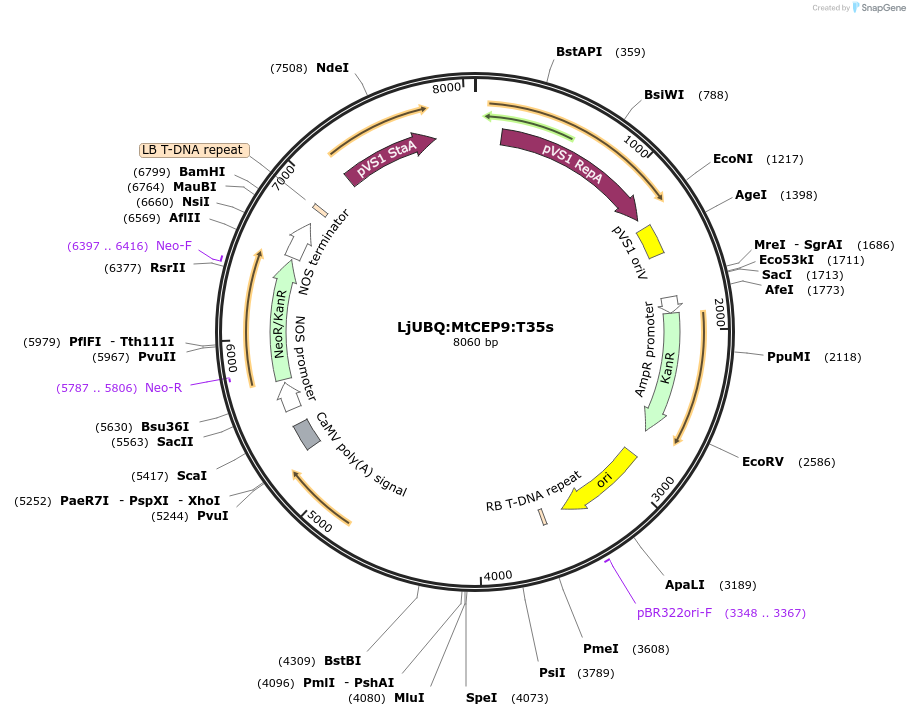
LjUBQ:MtCEP9:T35s
Plasmid#164733PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT35v5_AC233112_1015.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
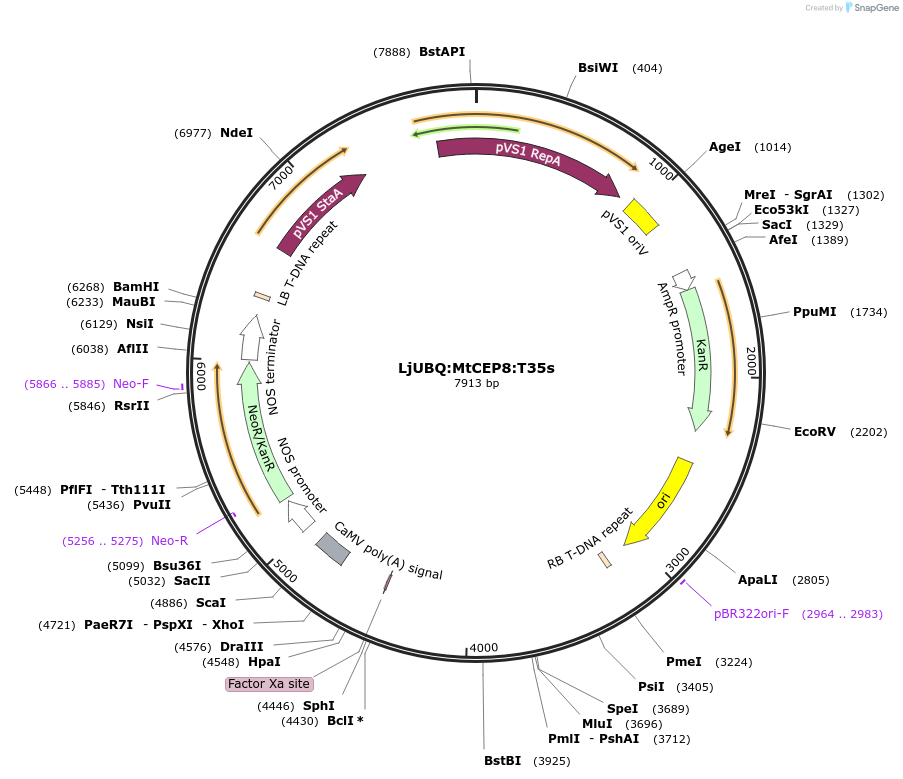
LjUBQ:MtCEP8:T35s
Plasmid#164732PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT35v5_AC233112_1014.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
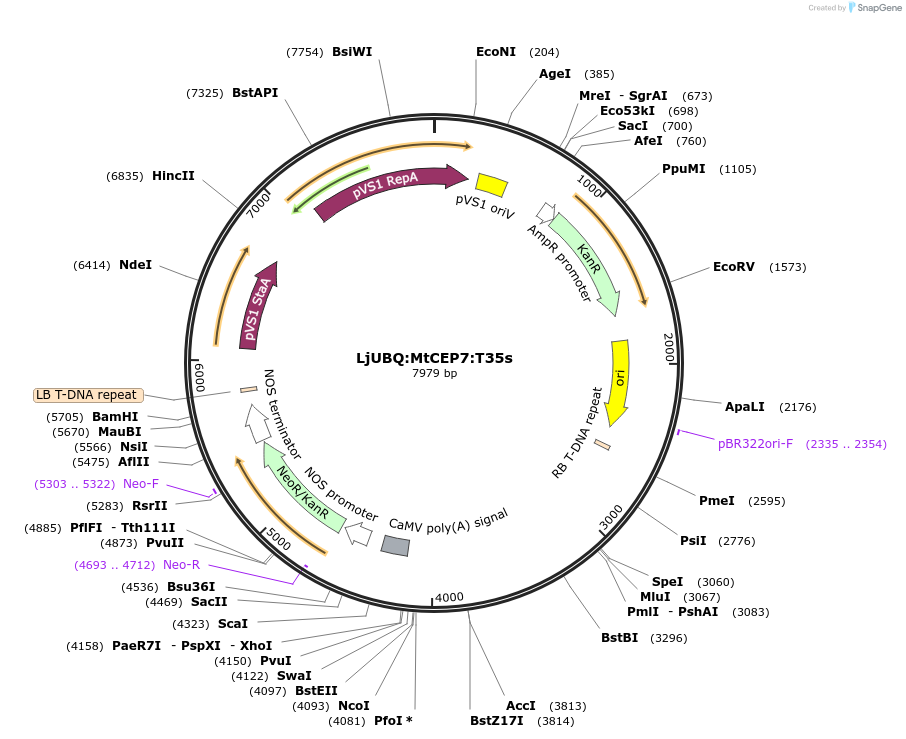
LjUBQ:MtCEP7:T35s
Plasmid#164731PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT35v5_AC233112_1013.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
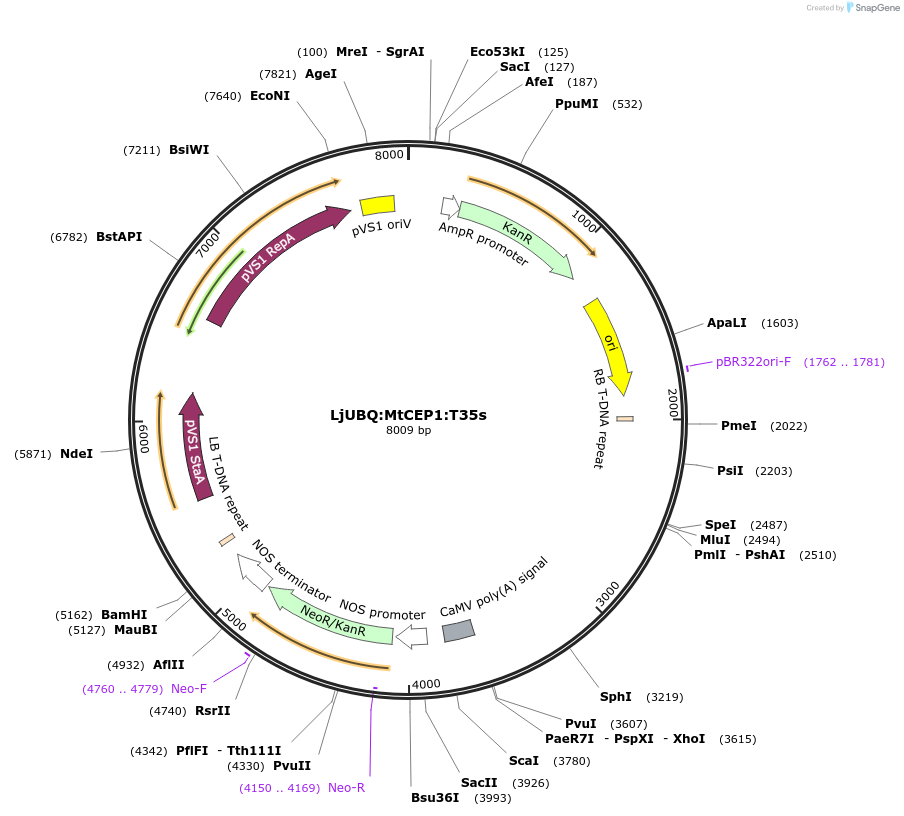
LjUBQ:MtCEP1:T35s
Plasmid#164729PurposeOverexpression of Peptide coding gene (Binary vector)DepositorInsertMT35v5_contig_59554_1.1
TagsTerminator 35SExpressionPlantPromoterLotus japonicus UbiquitinAvailable SinceFeb. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
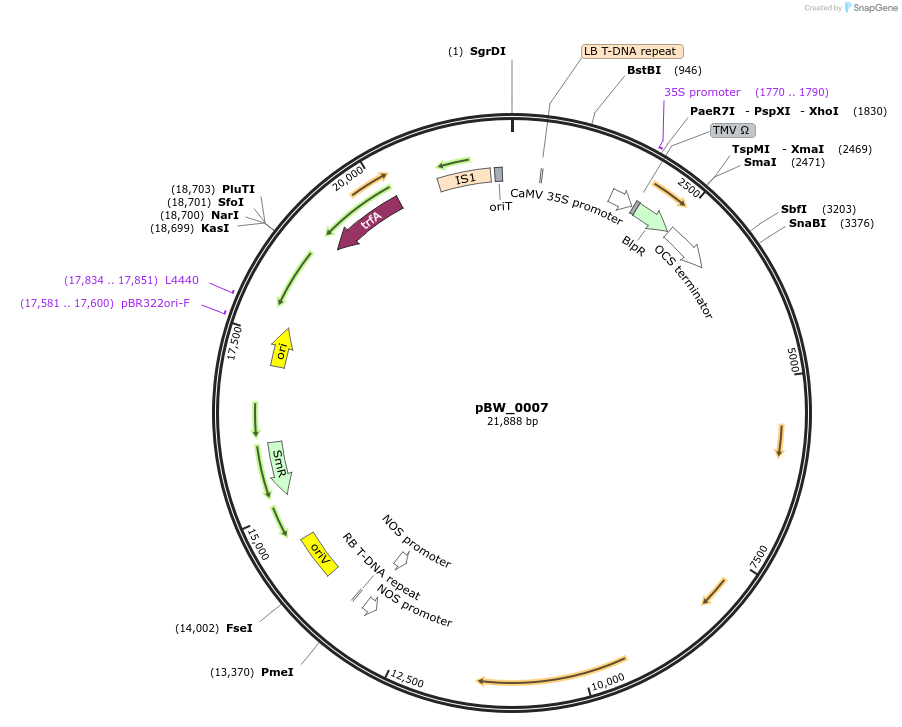
pBW_0007
Plasmid#102839PurposeBinary vector containing Sr22 PI573523 allele driven by Sr33 promoter, Sr33 terminatorDepositorAvailable SinceFeb. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
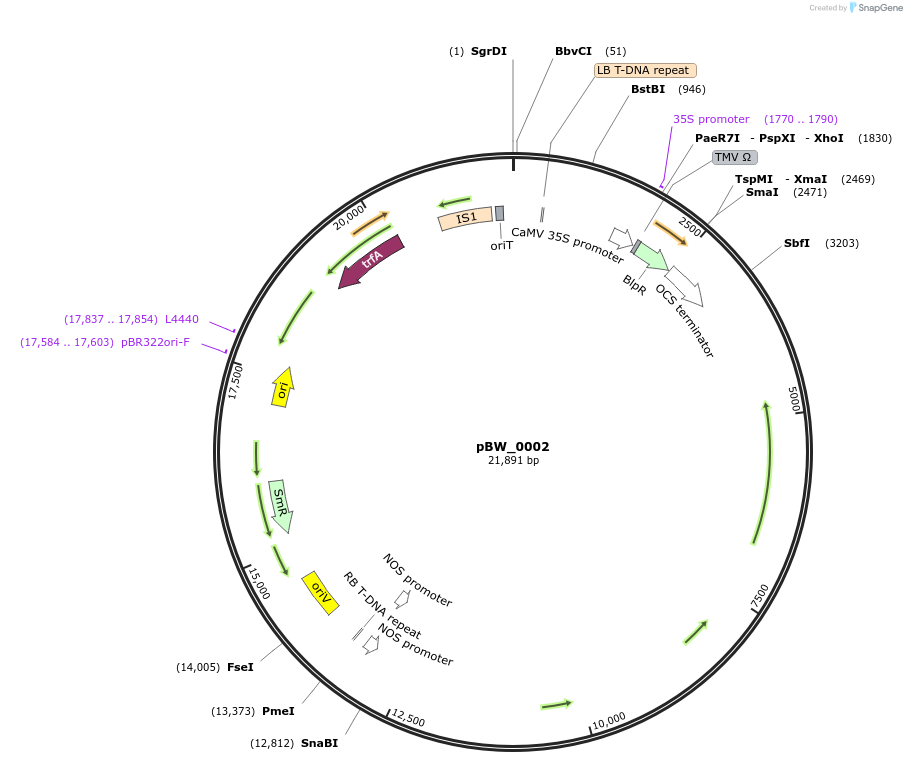
pBW_0002
Plasmid#102834PurposeBinary vector containing Sr22 Schomburgk allele driven by Sr33 promoter, Sr33 terminatorDepositorAvailable SinceFeb. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
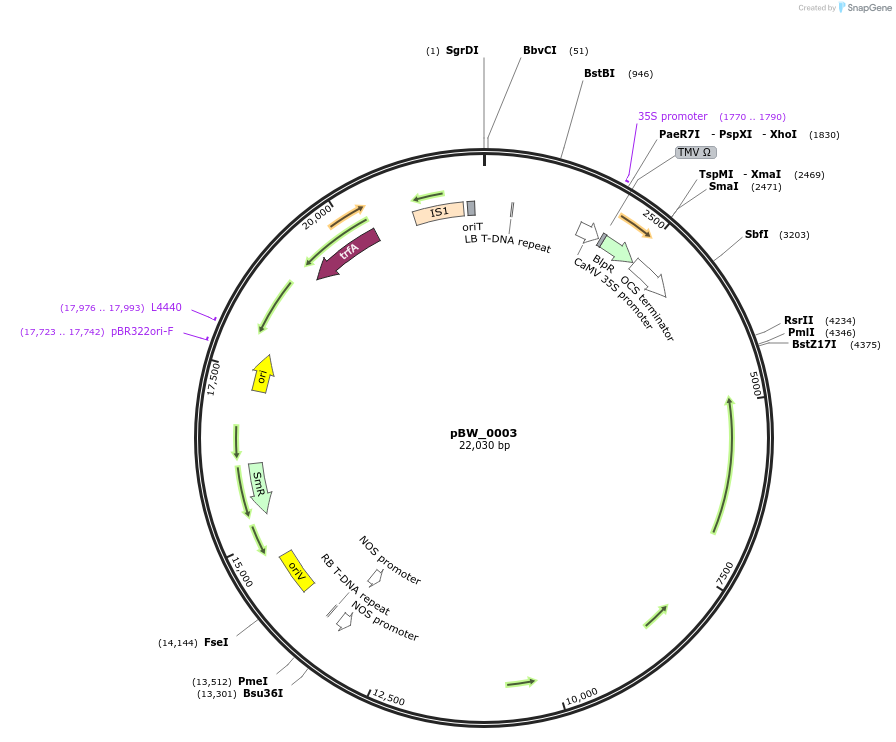
pBW_0003
Plasmid#102835PurposeBinary vector containing Sr22 Schomburgk allele driven by Sr33 promoter, Sr22 terminatorDepositorAvailable SinceFeb. 5, 2021AvailabilityAcademic Institutions and Nonprofits only -
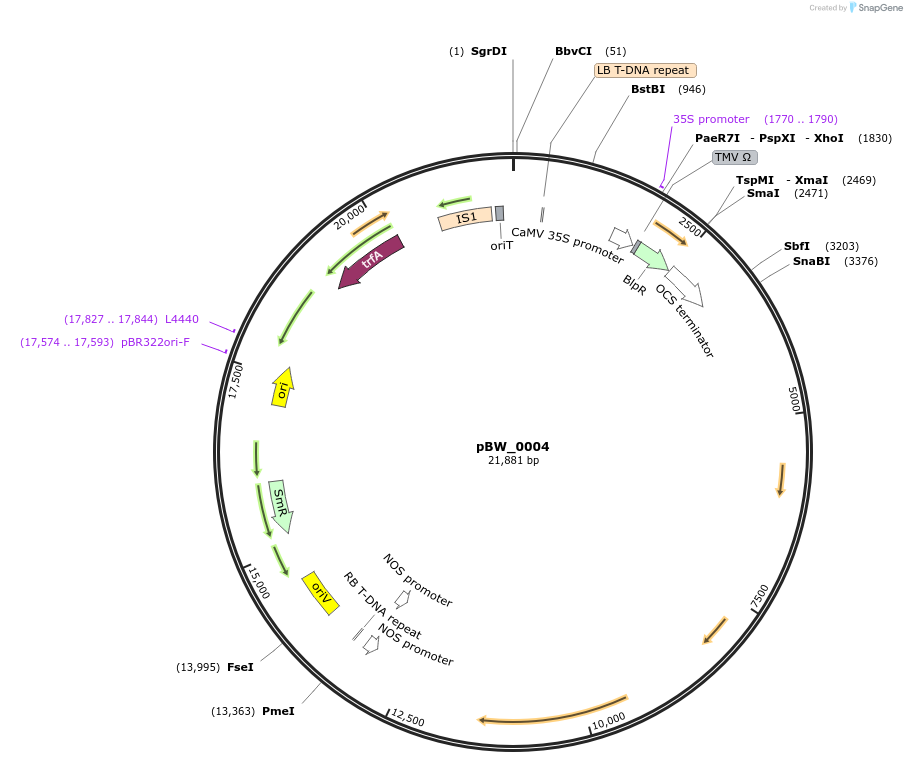
pBW_0004
Plasmid#102836PurposeBinary vector containing Sr22 PI190945 allele driven by Sr33 promoter, Sr33 terminatorDepositorAvailable SinceFeb. 5, 2021AvailabilityAcademic Institutions and Nonprofits only