We narrowed to 211 results for: pichia pastoris
-
Plasmid#227604PurposeInducible coexpression of His-tagged SLC25A46 and Strep-tagged MFN2 in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastPromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only
-
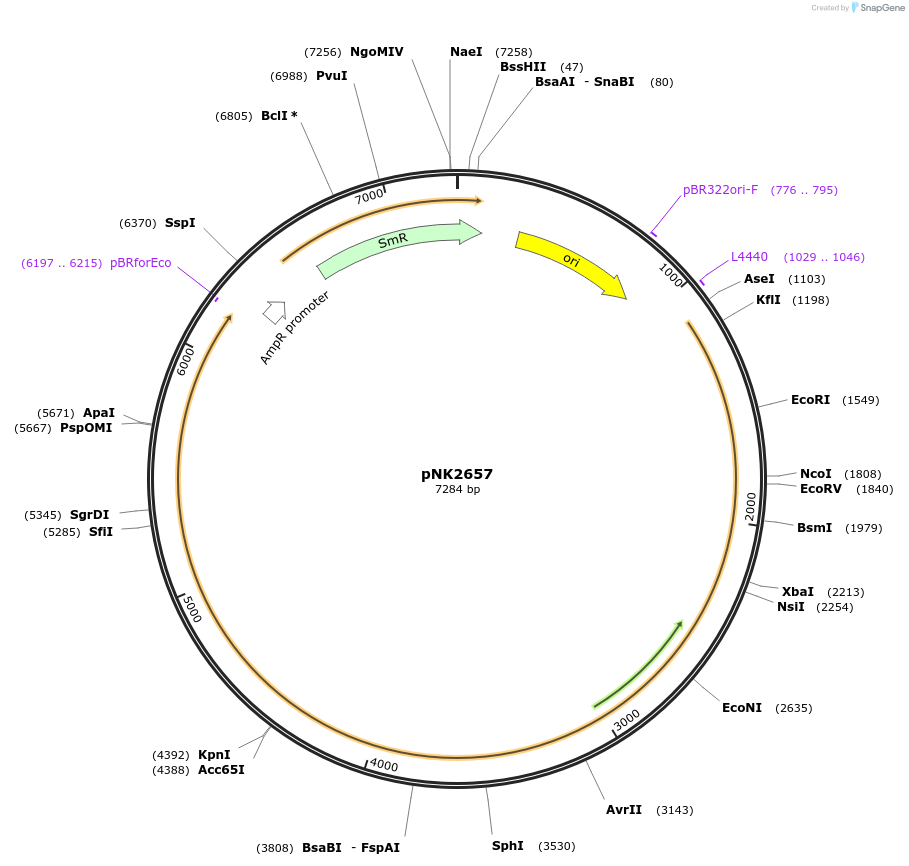
pNK2657
Plasmid#219741PurposeMoClo-compatible Level 0 promoterless vector encoding hispidin-synthase from Mycena citricolor codon-optimised for expression in Nicotiana benthamiana, Pichia pastoris, Homo sapiensDepositorInserthispidin- synthase from Mycena citricolor
UseSynthetic BiologyAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
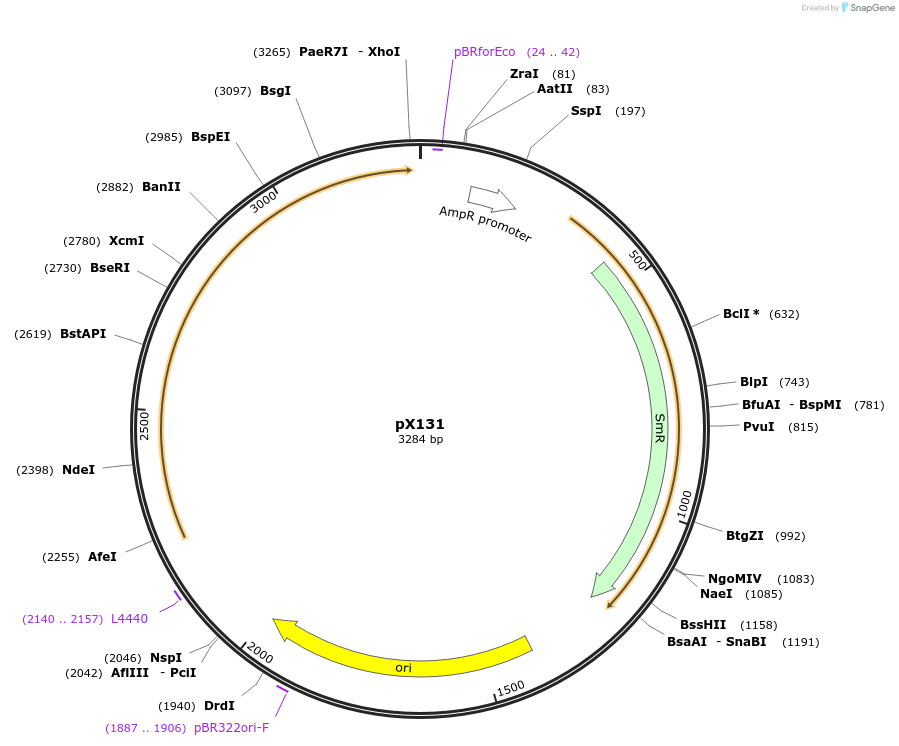
pX131
Plasmid#167149PurposeMoClo-compatible Level 0-SP promoterless vector encoding Aspergillus nidulans 4'-phosphopantetheinyl transferase NpgA codon-optimised for expression in Nicotiana benthamiana and Pichia pastorisDepositorInsert4'-phosphopantetheinyl transferase, npgA
UseSynthetic BiologyAvailable SinceJune 17, 2021AvailabilityAcademic Institutions and Nonprofits only -
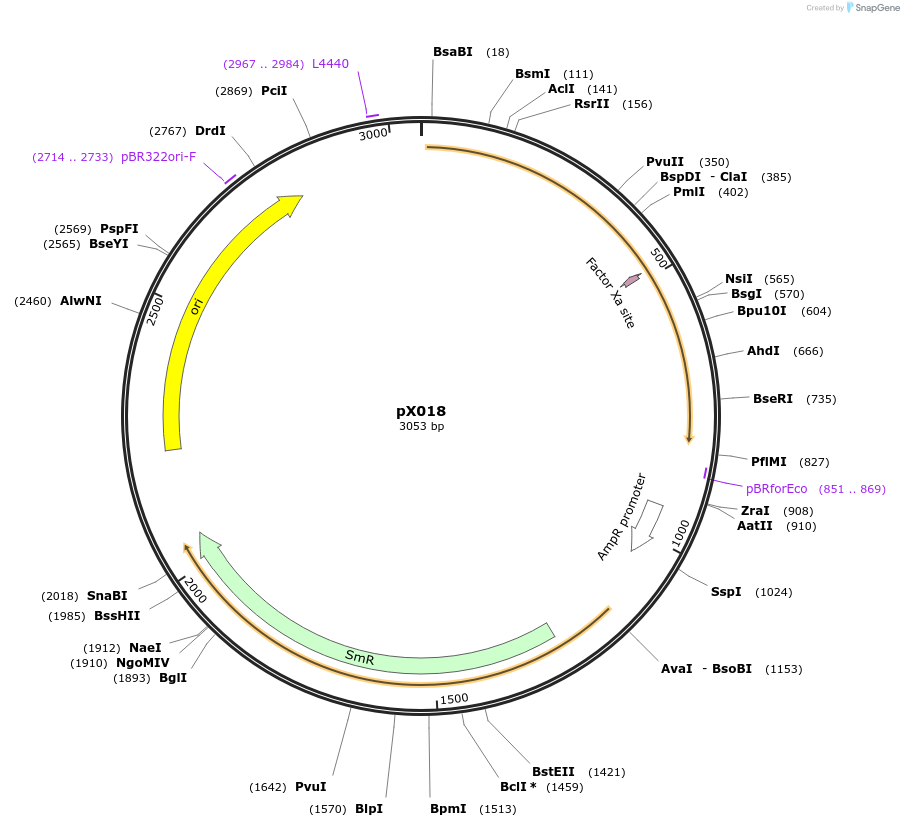
pX018
Plasmid#167144PurposeMoClo-compatible Level 0-SP promoterless vector encoding Neonothopanus nambi luciferase nnLuz codon-optimised for expression in Nicotiana benthamiana and Pichia pastorisDepositorInsertfungal luciferase, nnLuz
UseLuciferase and Synthetic BiologyAvailable SinceJune 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
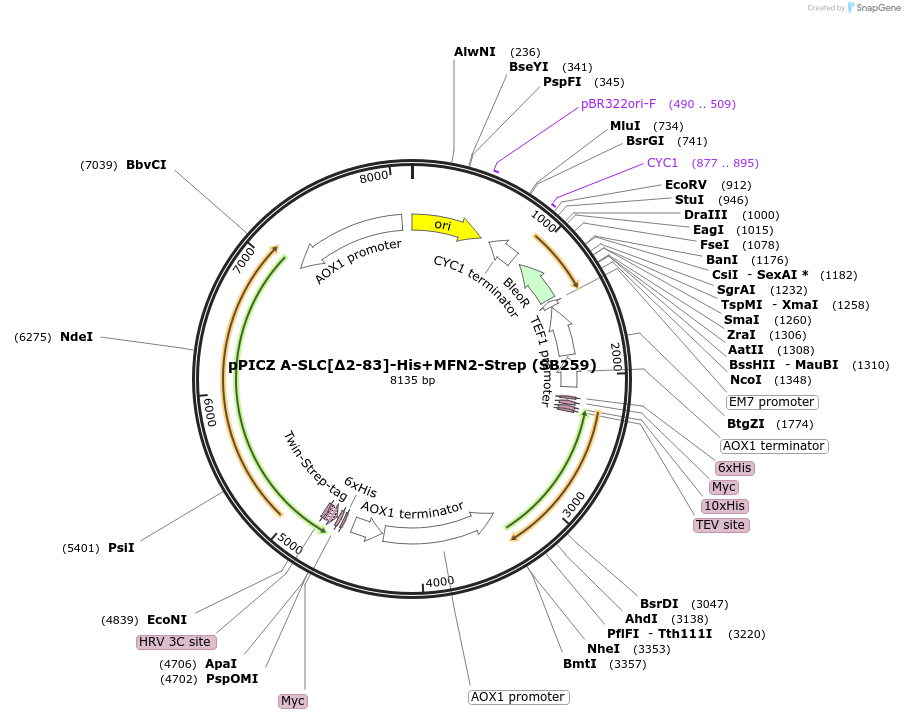
pPICZ A-SLC[∆2-83]-His+MFN2-Strep (SB259)
Plasmid#227608PurposeInducible coexpression of His-tagged SLC25A46 in its N-terminal region (∆2-83) and Strep-tagged MFN2 in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastMutationaa 2-83 deletedPromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
pPICZ A-SLC-His+MFN2[∆TM]-Strep (SB260)
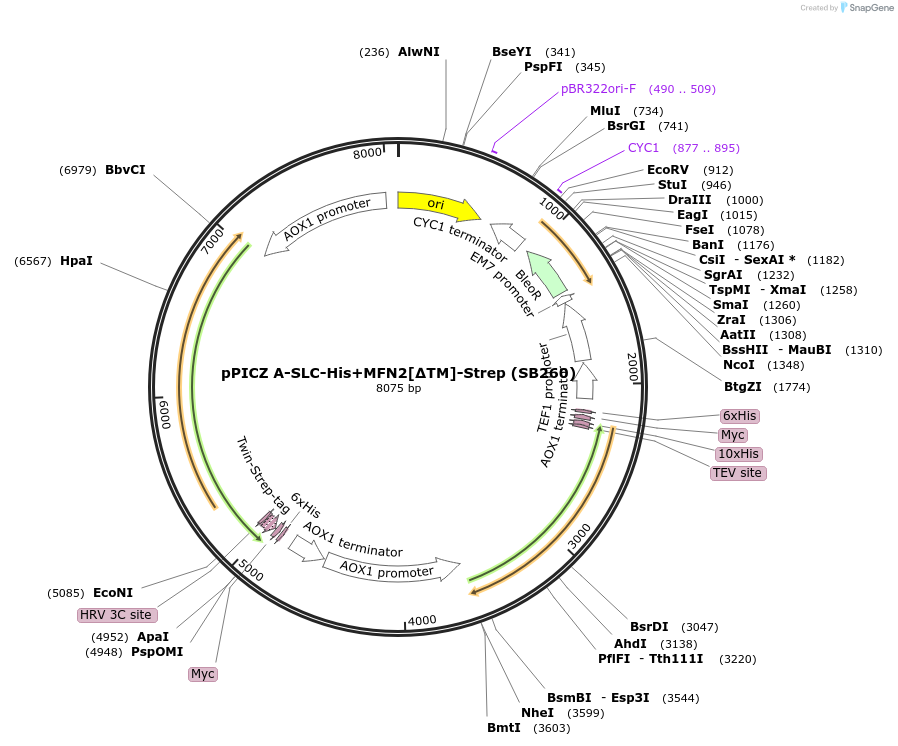
Plasmid#227609PurposeInducible coexpression of His-tagged SLC25A46 and Strep-tagged MFN2 lacking its transmembrane region in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastMutationcontains aa 2-544, 651-757 only [TM deleted]PromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
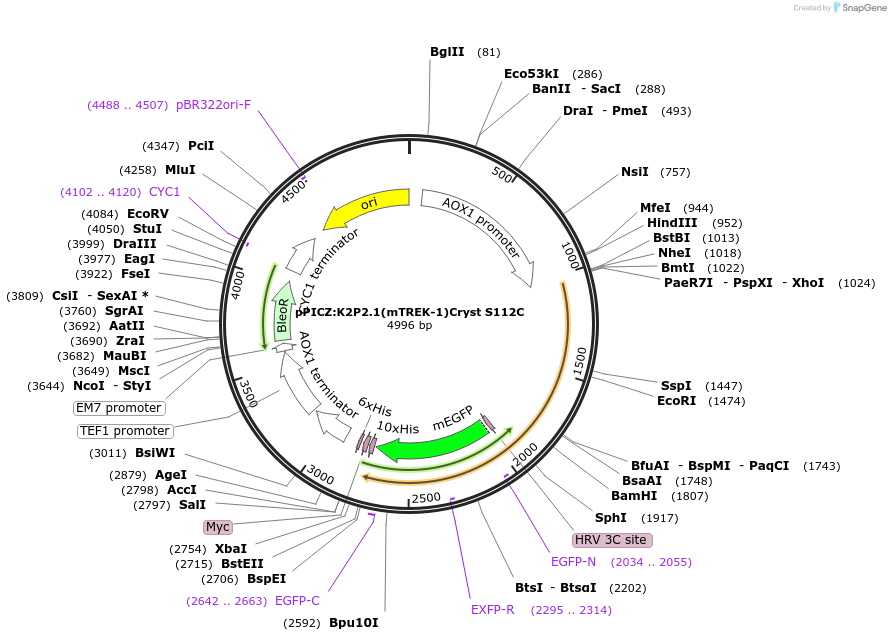
pPICZ:K2P2.1(mTREK-1)Cryst S112C
Plasmid#224780PurposePichia pastoris expression vector for generating mouse TREK-1(21S-322T) + 3C cleavable site + eGFP + 10x His under the AOX1 promoterDepositorInsertPotassium channel subfamily K member 2 (Kcnk2 Mouse)
ExpressionYeastMutationK84R / Q85E / T86K / I88L / A89R / Q90A / A92P / …PromoterAOX1 promoterAvailable SinceSept. 13, 2024AvailabilityAcademic Institutions and Nonprofits only -
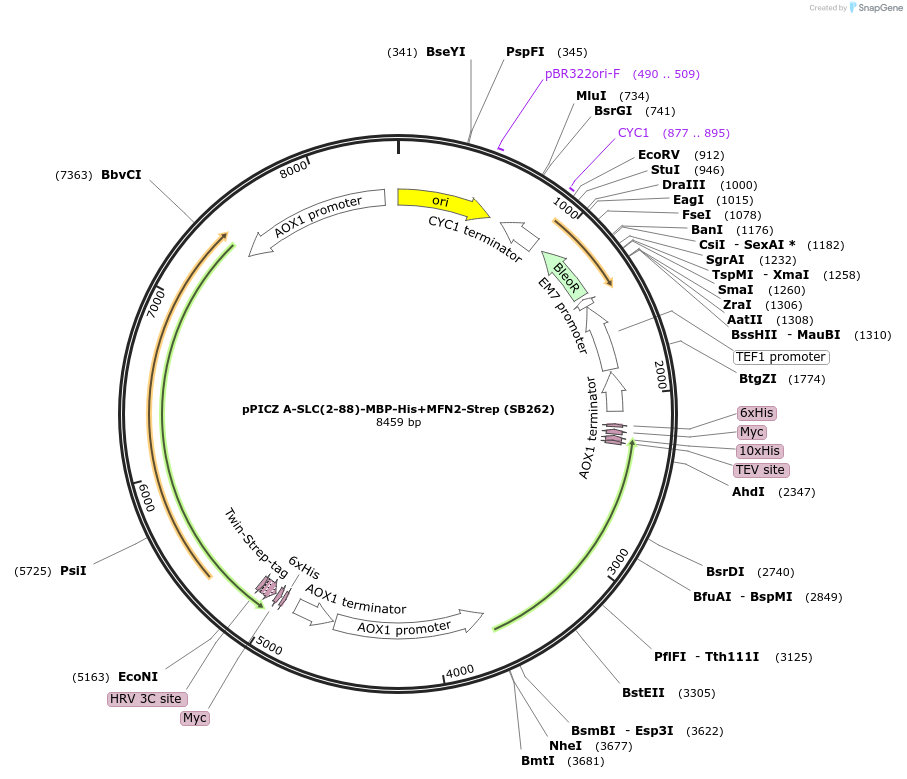
pPICZ A-SLC(2-88)-MBP-His+MFN2-Strep (SB262)
Plasmid#227611PurposeInducible coexpression of His-tagged SLC25A46 truncated to its N-terminal region (2-88) fused to MBP and MFN2-Strep in Pichia pastorisDepositorTagsMBP, 10xHis and Twin-StrepTagExpressionYeastMutationcontains aa 2-88 onlyPromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
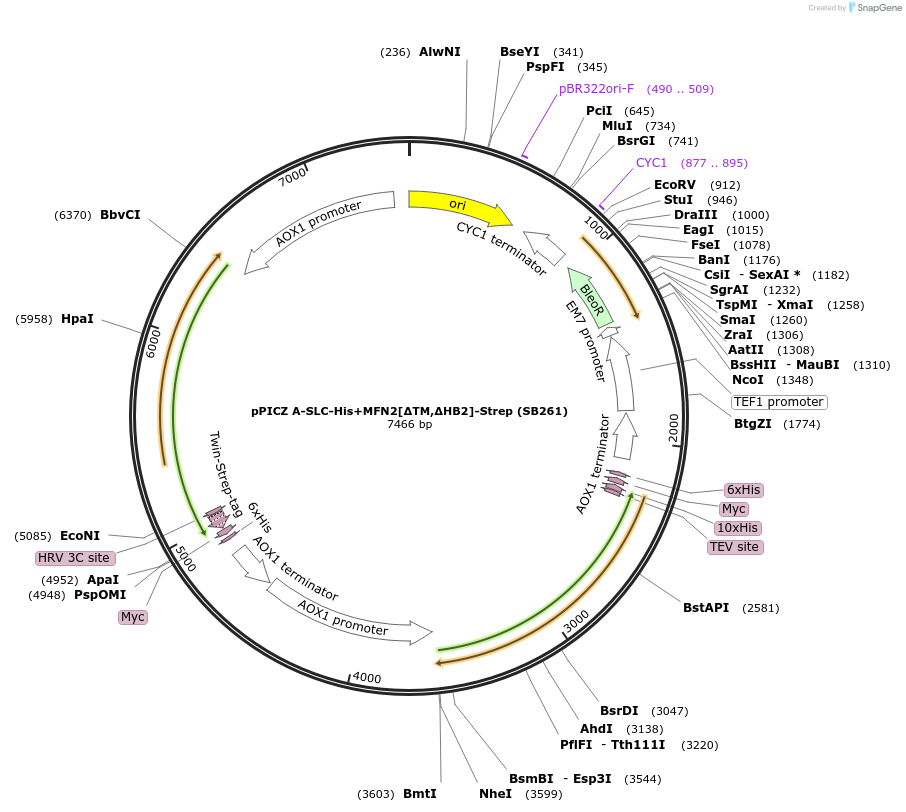
pPICZ A-SLC-His+MFN2[∆TM,∆HB2]-Strep (SB261)
Plasmid#227610PurposeInducible coexpression of His-tagged SLC25A46 and Strep-tagged MFN2 lacking its transmembrane region and helical bundle 2 (HB2) in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastMutationcontains aa 2-400, 706-757 [TM and HB2 deleted]PromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
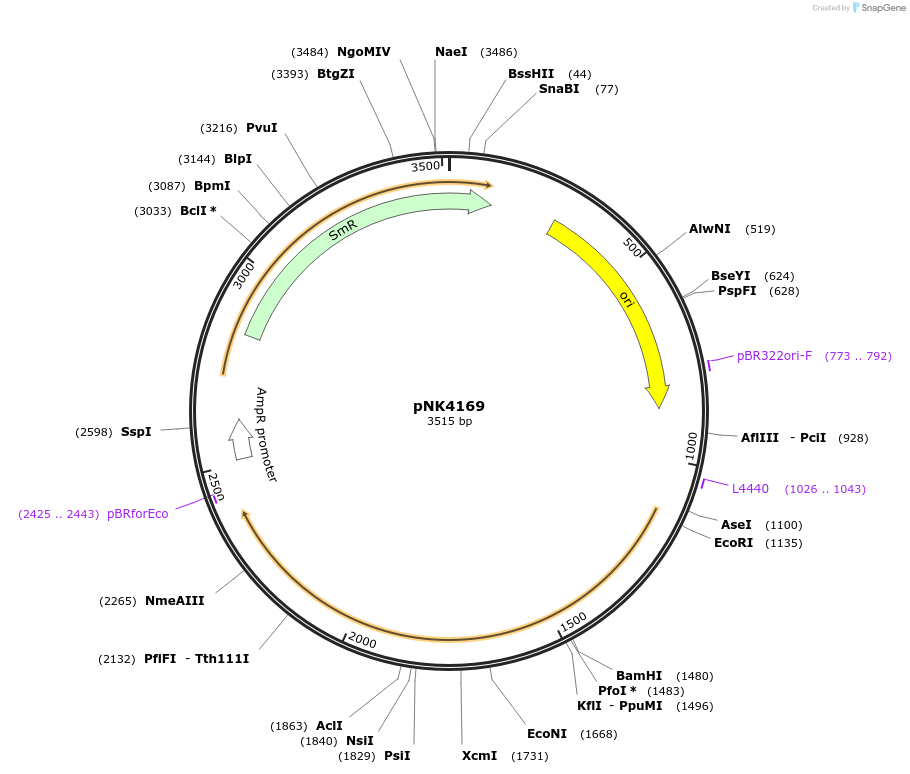
pNK4169
Plasmid#219744PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi hispidin-3-hydroxylase nnH3H_v2 codon-optimised for expression in Pichia pastorisDepositorInsertmutant of fungal hispidin-3-hydroxylase
UseSynthetic BiologyMutationD37E, V181I, S323M, M385KAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
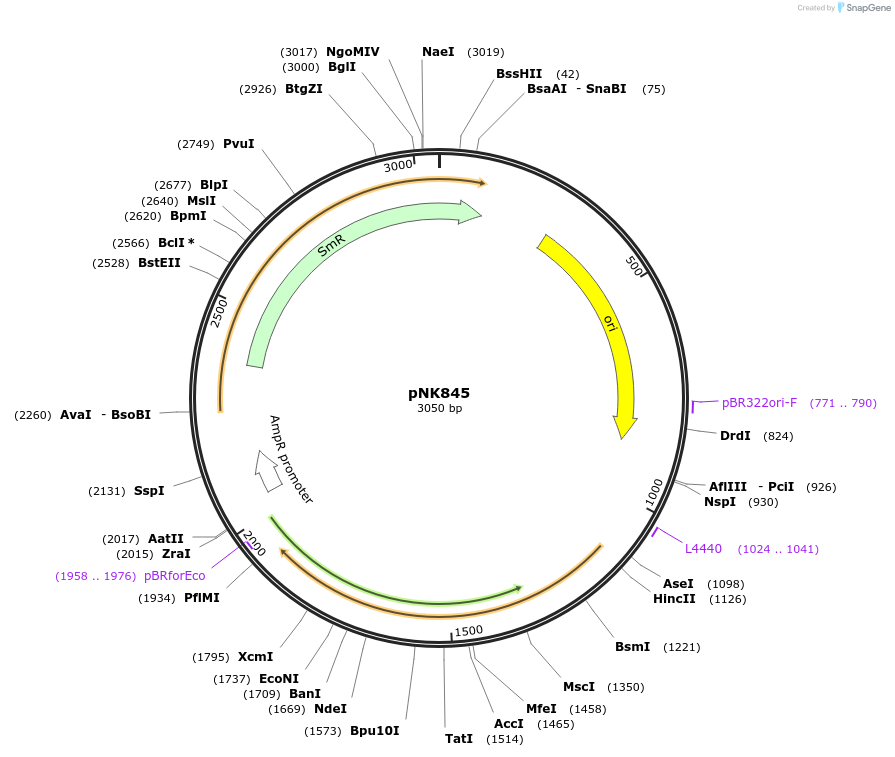
pNK845
Plasmid#219746PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi luciferase nnLuz_v4 codon-optimised for expression in Pichia pastoris, Homo sapiensDepositorInsertmutant of fungal luciferase
UseLuciferase and Synthetic BiologyMutationI3S, N4T, F11L, I63T, T99P, T192S, A199PAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only