We narrowed to 11,516 results for: Kars
-
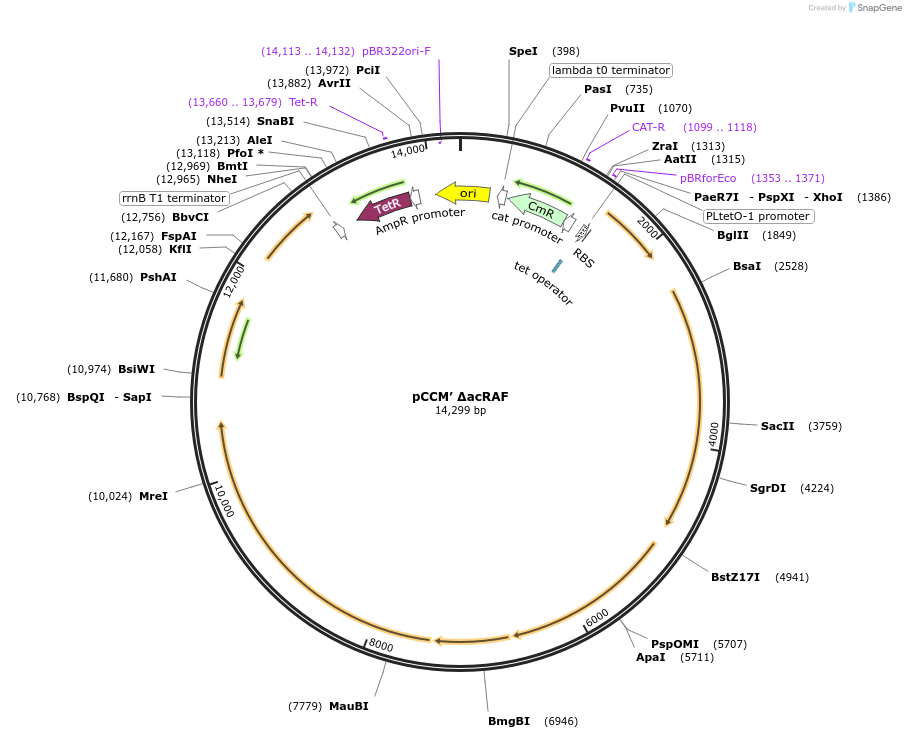
Plasmid#162710PurposeSecond CCM operon lacking acRAF; Deletion of putative rubisco chaperone, acRAFDepositorInsertSecond CCM operon cloned from H. neapolitanus
ExpressionBacterialMutationpCCM' with a deletion of putative rubisco ch…PromoterPLtet0-1 promoterAvailable SinceFeb. 9, 2021AvailabilityAcademic Institutions and Nonprofits only -
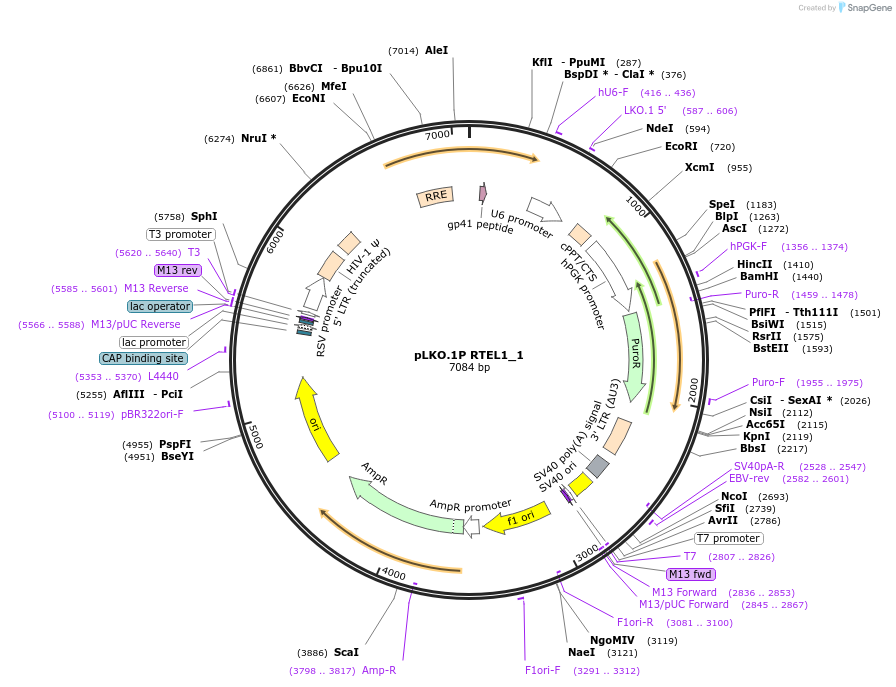
pLKO.1P RTEL1_1
Plasmid#160799PurposeSuppress RTEL1DepositorInsertshRTEL1_1
UseLentiviralAvailable SinceJan. 4, 2021AvailabilityAcademic Institutions and Nonprofits only -
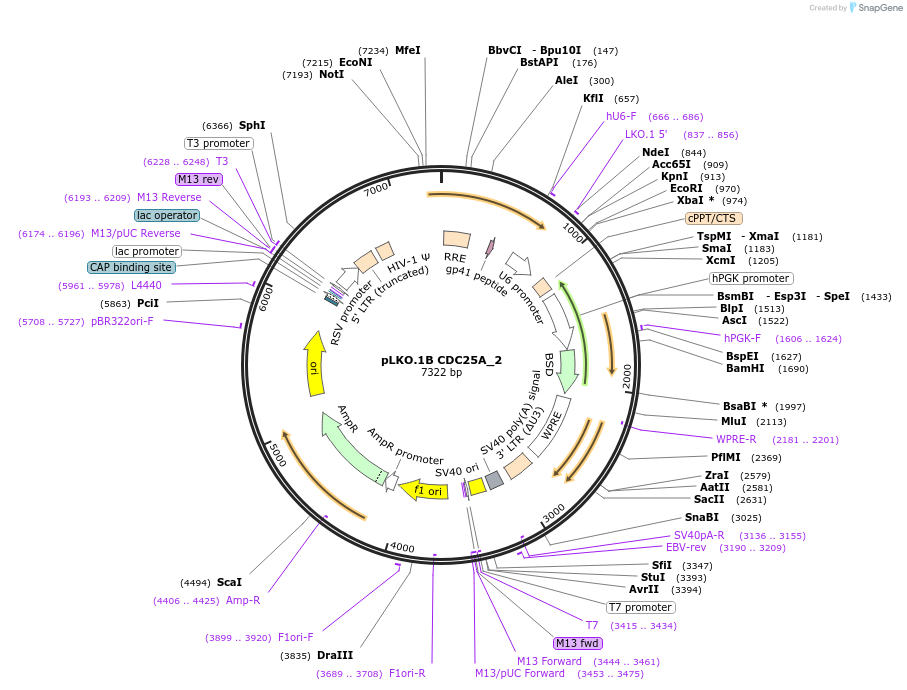
pLKO.1B CDC25A_2
Plasmid#160768PurposeSuppress CDC25ADepositorInsertshCDC25A_2
UseLentiviralAvailable SinceJan. 4, 2021AvailabilityAcademic Institutions and Nonprofits only -
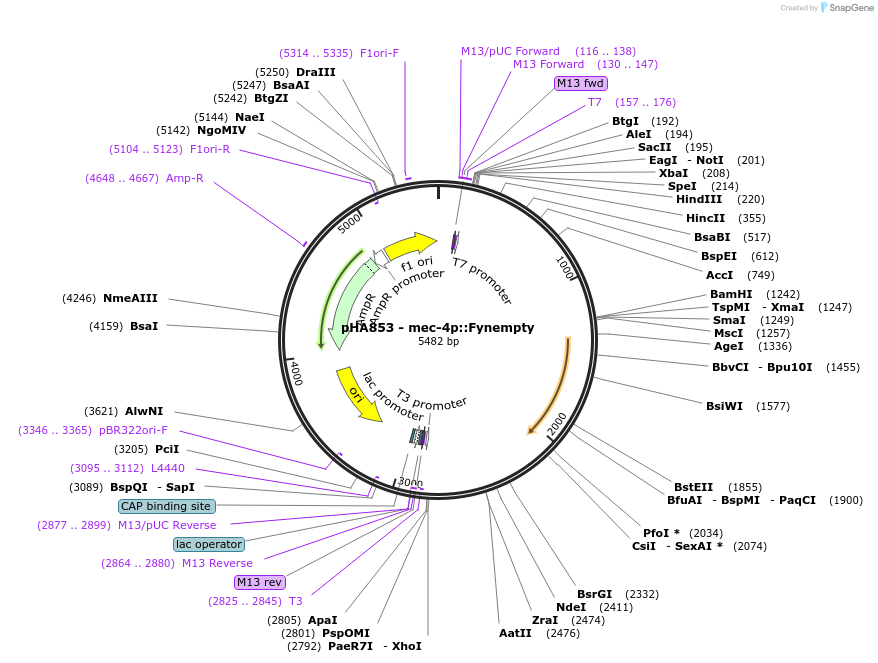
pHA853 - mec-4p::Fynempty
Plasmid#139210PurposeControl vector for Fyn kinase expression in mec-4 neuronsDepositorAvailable SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
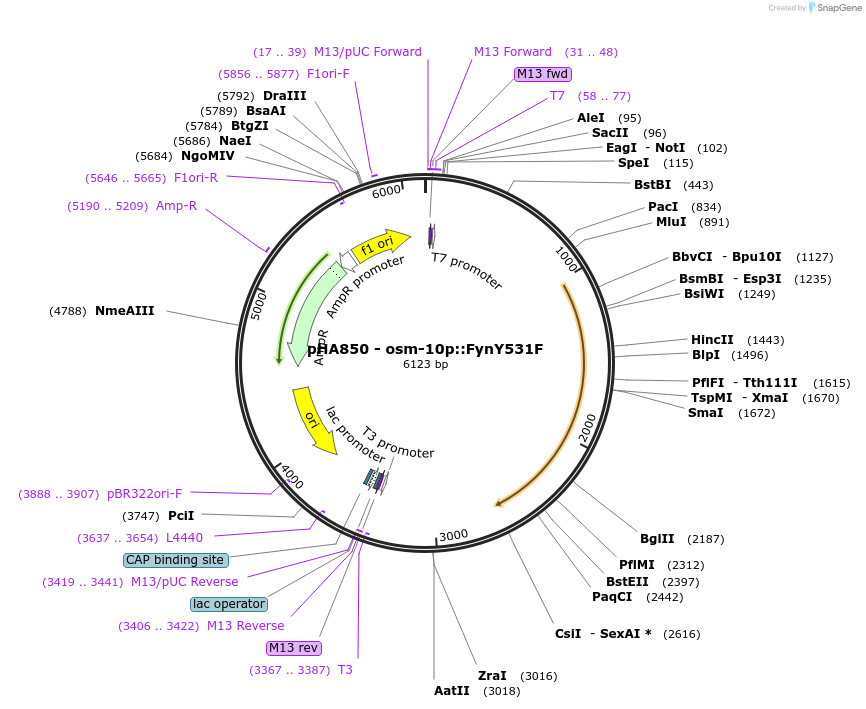
pHA850 - osm-10p::FynY531F
Plasmid#139207PurposeExpress activated Fyn kinase in osm-10 neuronsDepositorAvailable SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
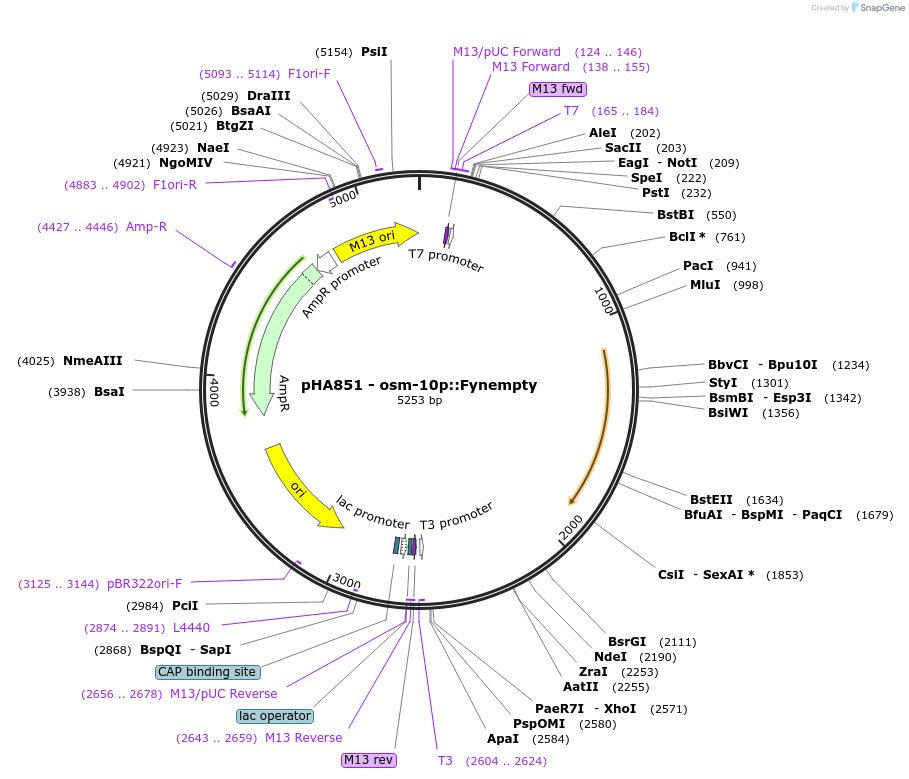
pHA851 - osm-10p::Fynempty
Plasmid#139208PurposeControl vector for Fyn kinase expression in osm-10 neuronsDepositorAvailable SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
FUS_LC_withR
Plasmid#139128Purposeexpress FUS LC with the same number and spacing of arginines as hnRNPA2 LCDepositorAvailable SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
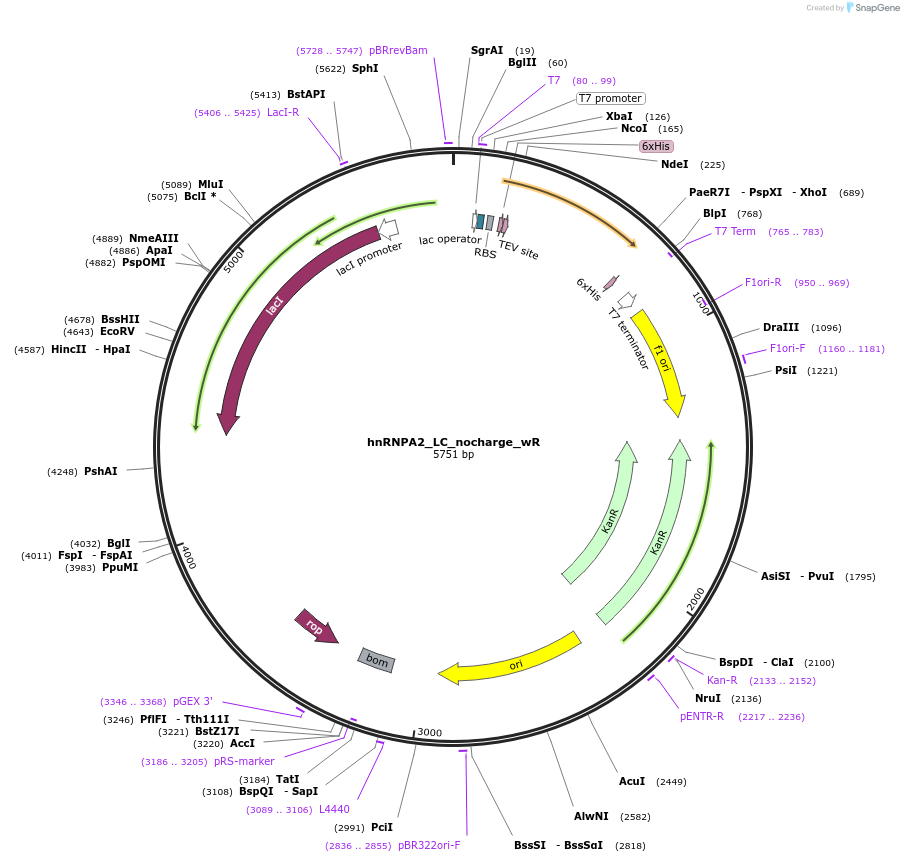
hnRNPA2_LC_nocharge_wR
Plasmid#139119Purposeexpress hnRNPA2 LC with no charged residues with R added backDepositorAvailable SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
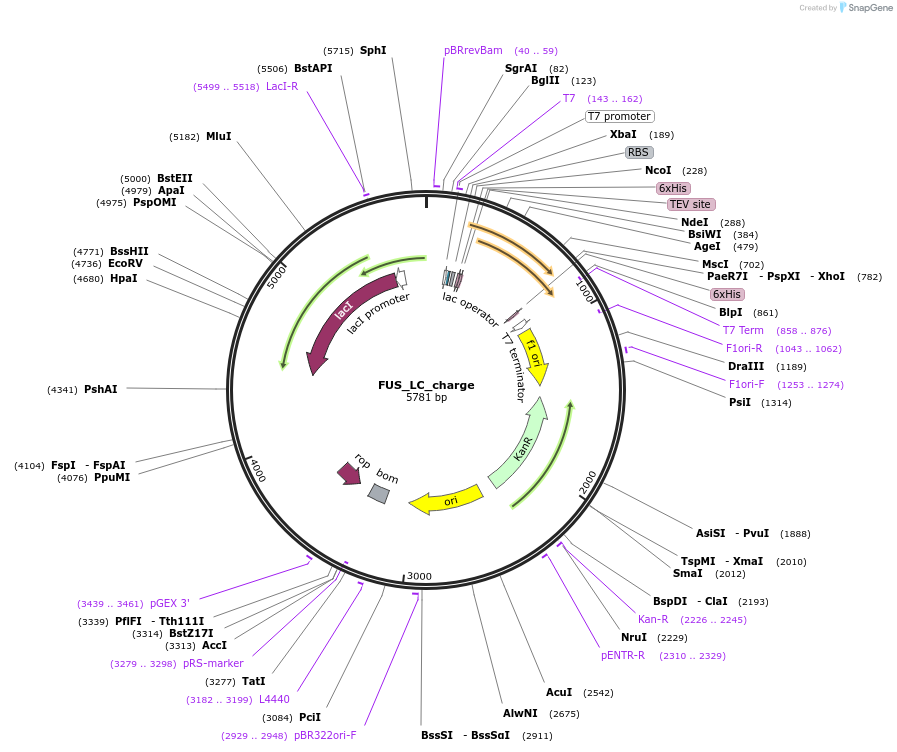
FUS_LC_charge
Plasmid#139126Purposeexpress FUS LC with an hnRNPA2 LC-like charge patterningDepositorInsertFUS (FUS Human)
ExpressionBacterialMutationAdd R, K, D to make FUS LC have similar charge pa…Available SinceDec. 7, 2020AvailabilityAcademic Institutions and Nonprofits only -
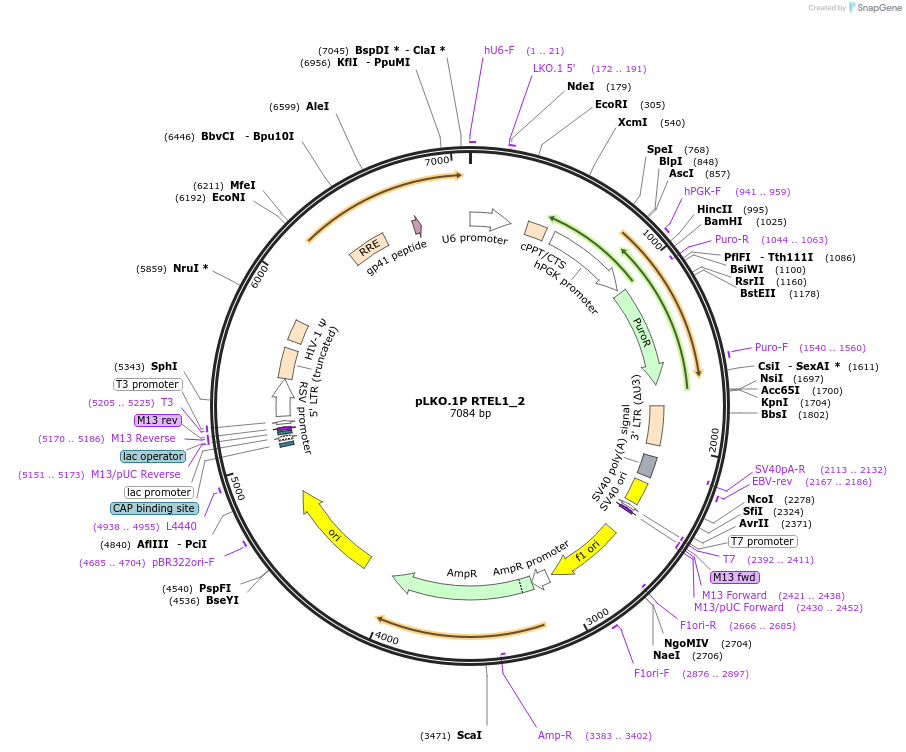
pLKO.1P RTEL1_2
Plasmid#160800PurposeSuppress RTEL1DepositorInsertshRTEL1_2
UseLentiviralAvailable SinceOct. 30, 2020AvailabilityAcademic Institutions and Nonprofits only -
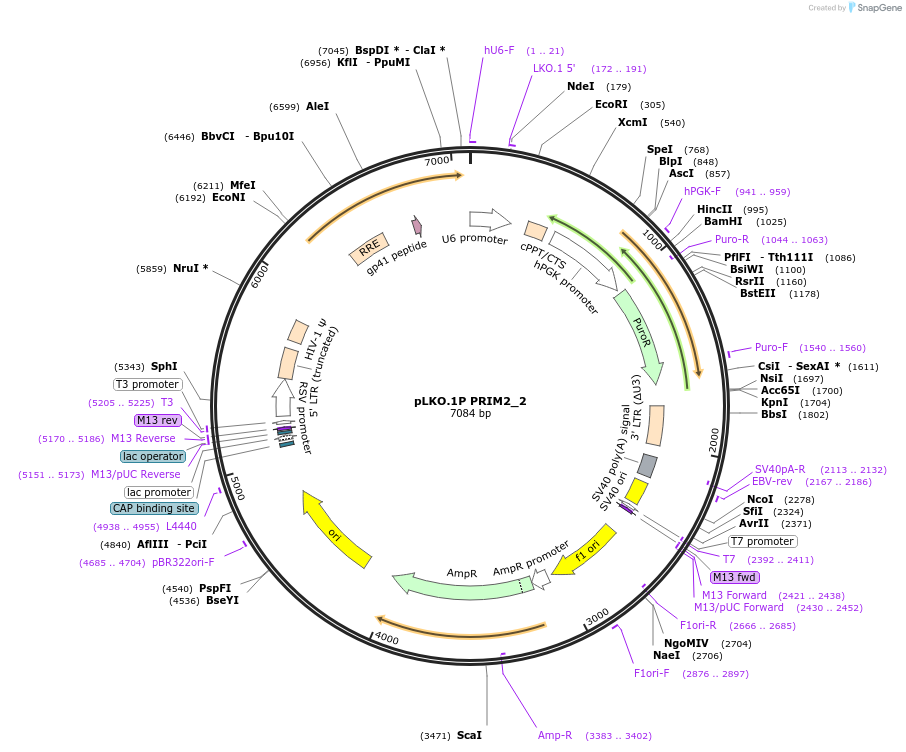
pLKO.1P PRIM2_2
Plasmid#160796PurposeSuppress PRIM2DepositorInsertshPRIM2_2
UseLentiviralAvailable SinceOct. 30, 2020AvailabilityAcademic Institutions and Nonprofits only -
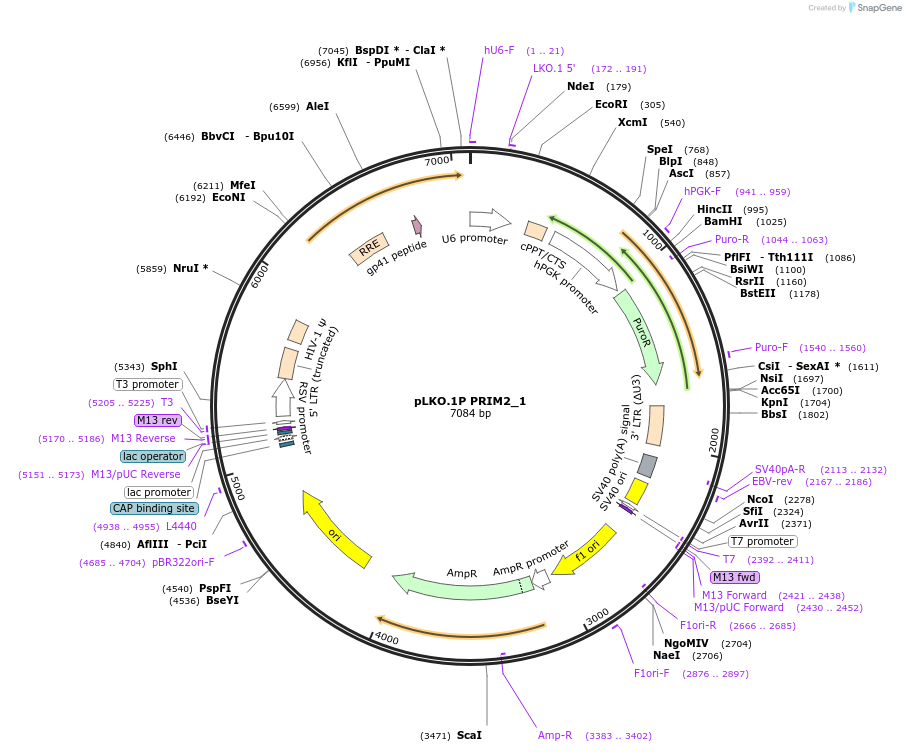
pLKO.1P PRIM2_1
Plasmid#160795PurposeSuppress PRIM2DepositorInsertshPRIM2_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
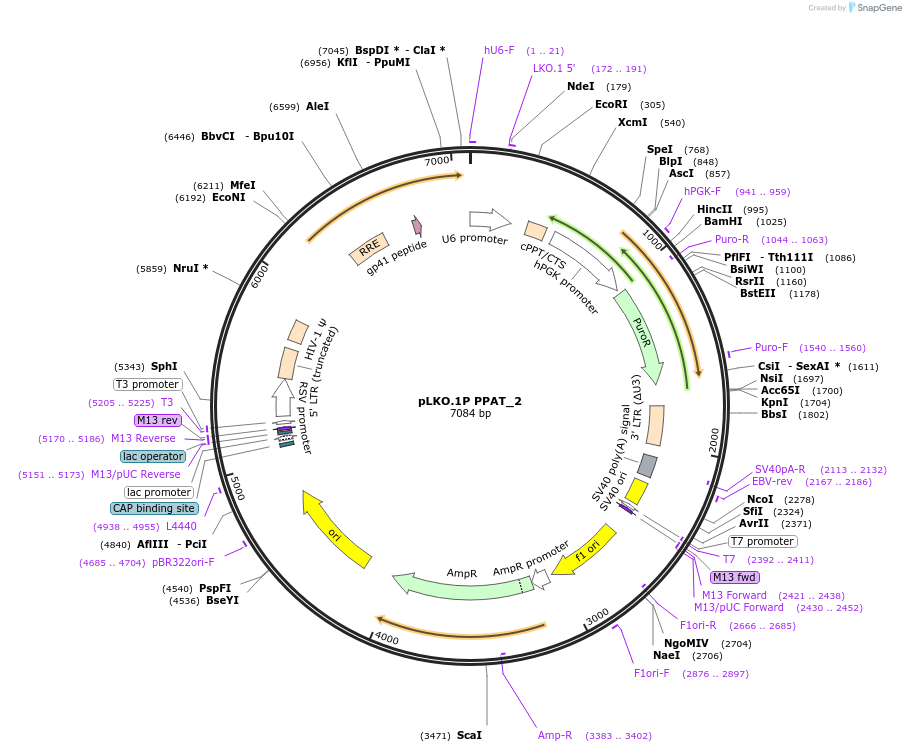
pLKO.1P PPAT_2
Plasmid#160794PurposeSuppress PPATDepositorInsertshPPAT_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
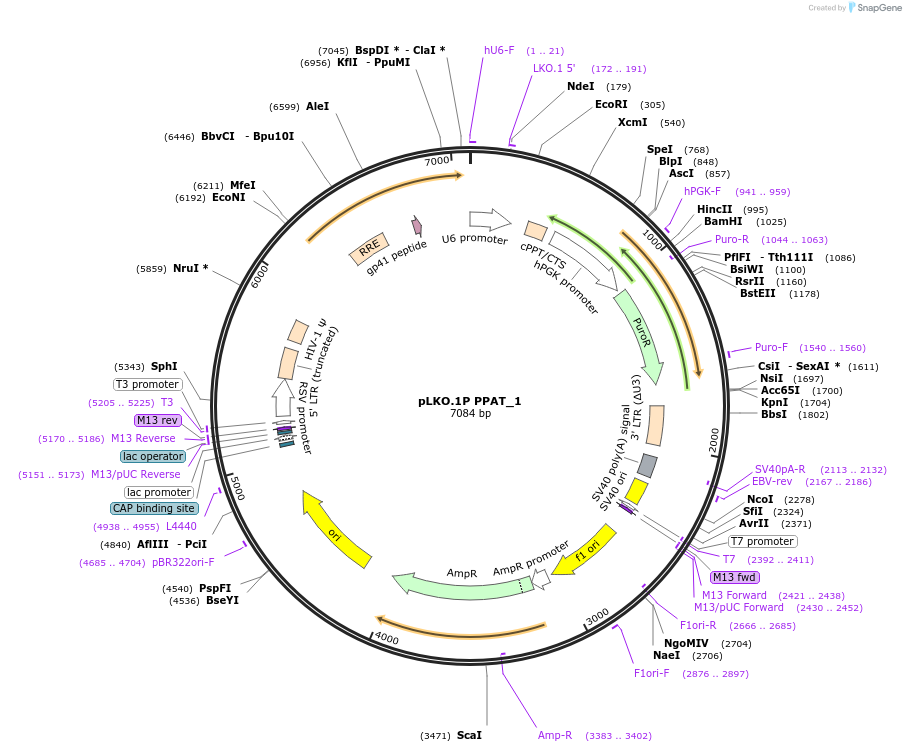
pLKO.1P PPAT_1
Plasmid#160793PurposeSuppress PPATDepositorInsertshPPAT_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
pLKO.1P NTHL1_2
Plasmid#160788PurposeSuppress NTHL1DepositorInsertshNTHL1_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
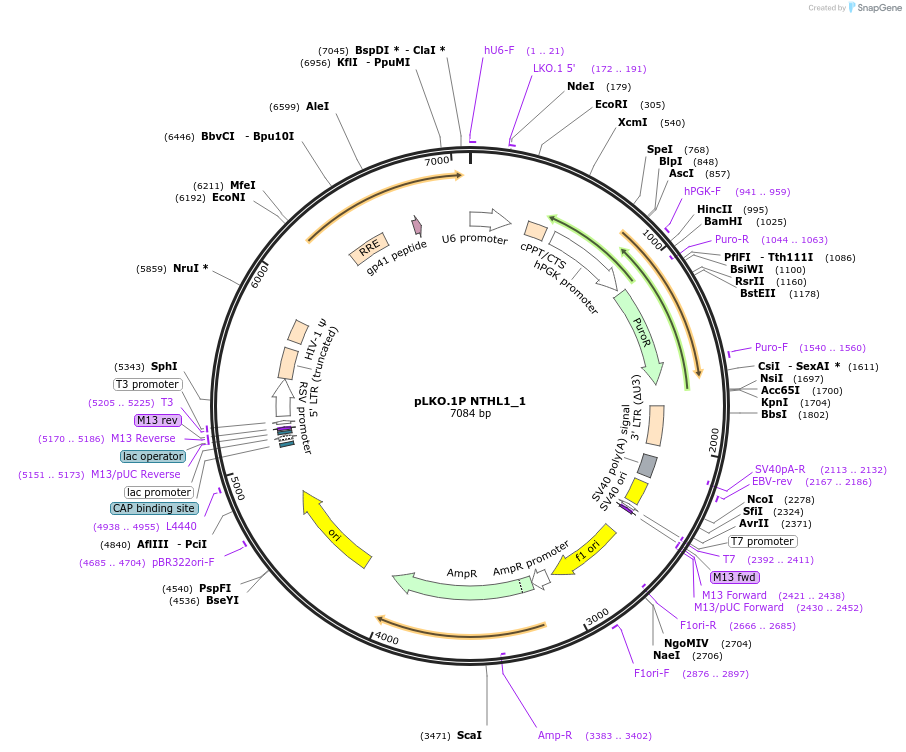
pLKO.1P NTHL1_1
Plasmid#160787PurposeSuppress NTHL1DepositorInsertshNTHL1_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
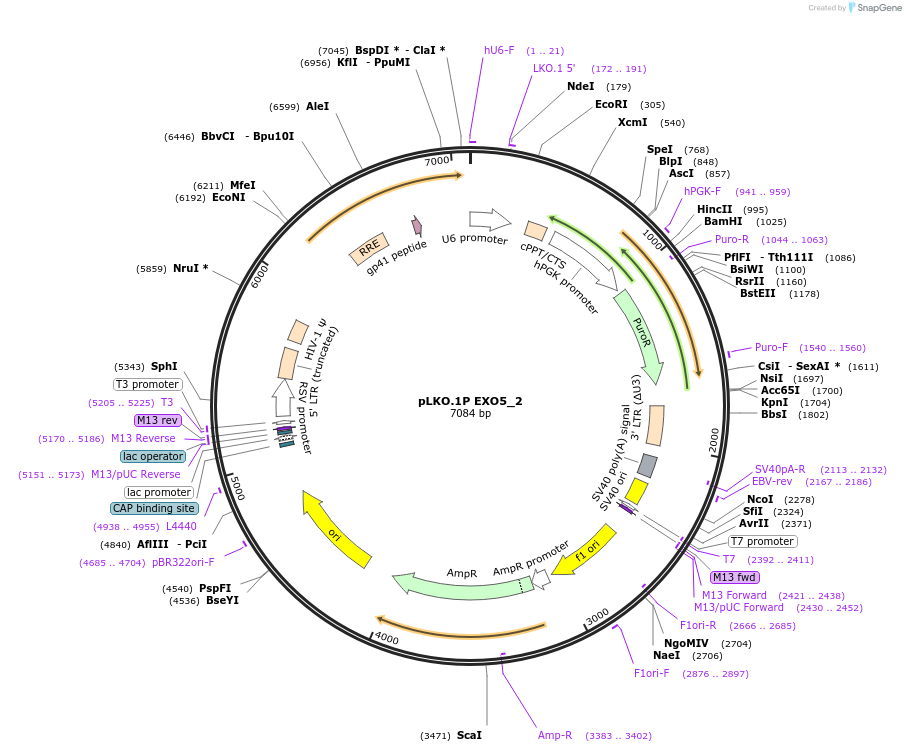
pLKO.1P EXO5_2
Plasmid#160784PurposeSuppress EXO5DepositorInsertshEXO5_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
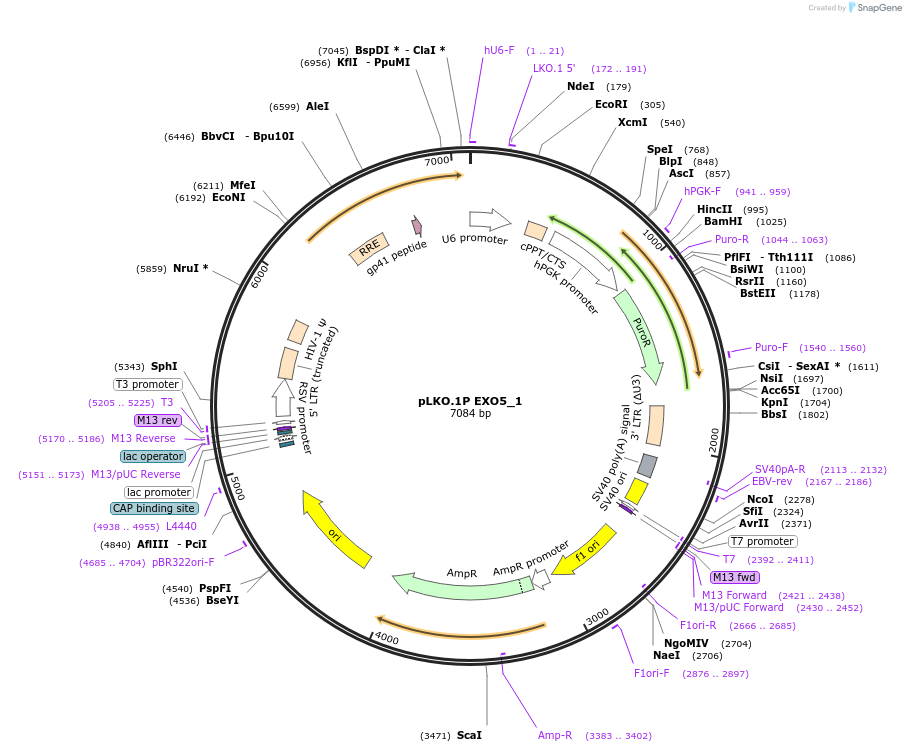
pLKO.1P EXO5_1
Plasmid#160783PurposeSuppress EXO5DepositorInsertshEXO5_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
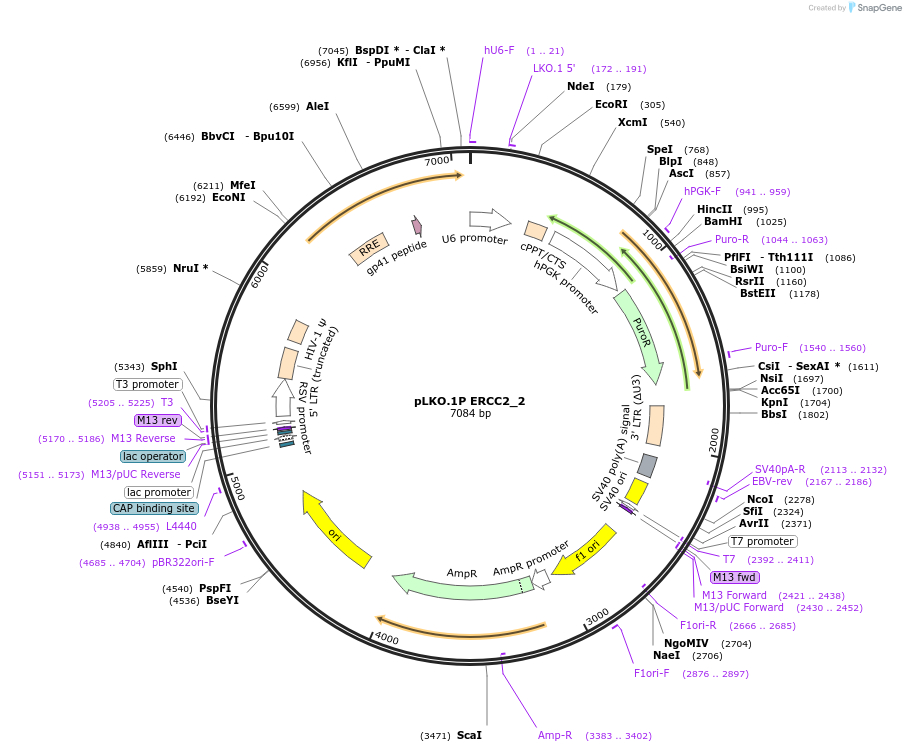
pLKO.1P ERCC2_2
Plasmid#160782PurposeSuppress ERCC2DepositorInsertshERCC2_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
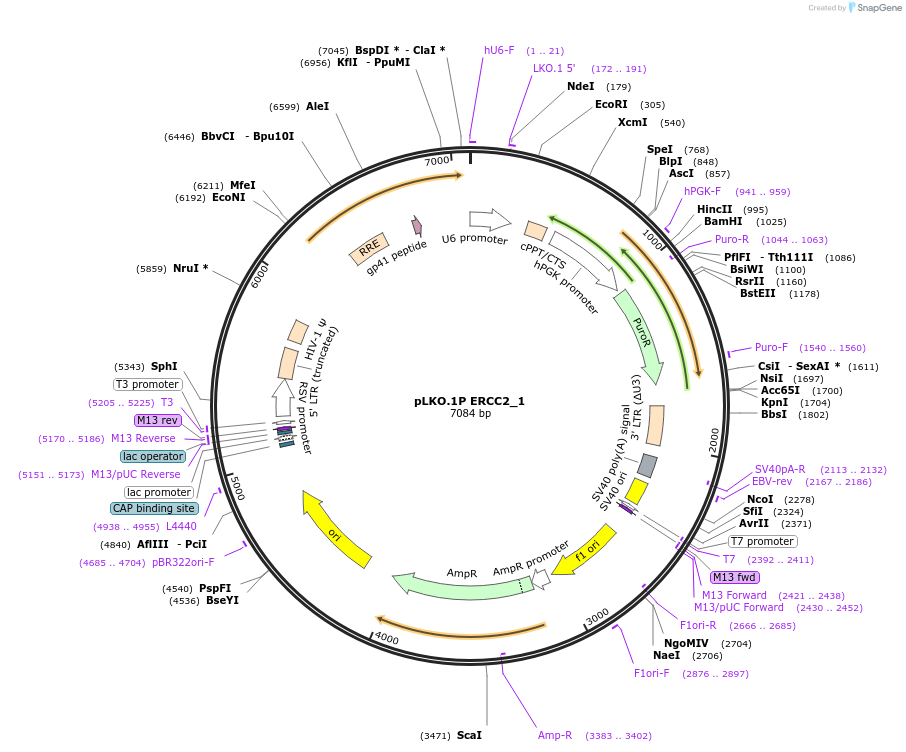
pLKO.1P ERCC2_1
Plasmid#160781PurposeSuppress ERCC2DepositorInsertshERCC2_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
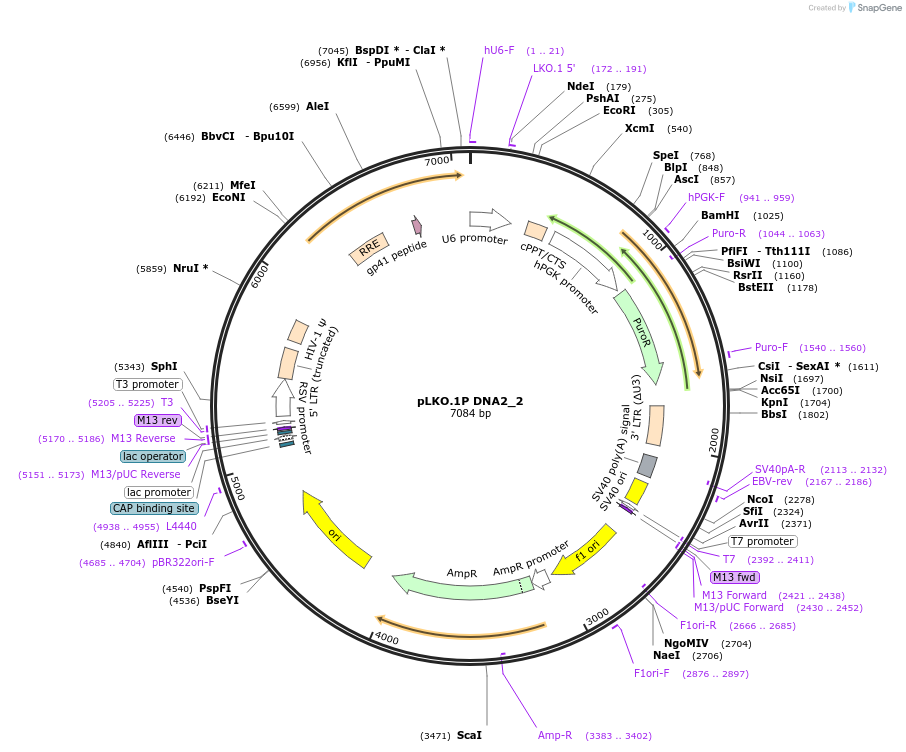
pLKO.1P DNA2_2
Plasmid#160780PurposeSuppress DNA2DepositorInsertshDNA2_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
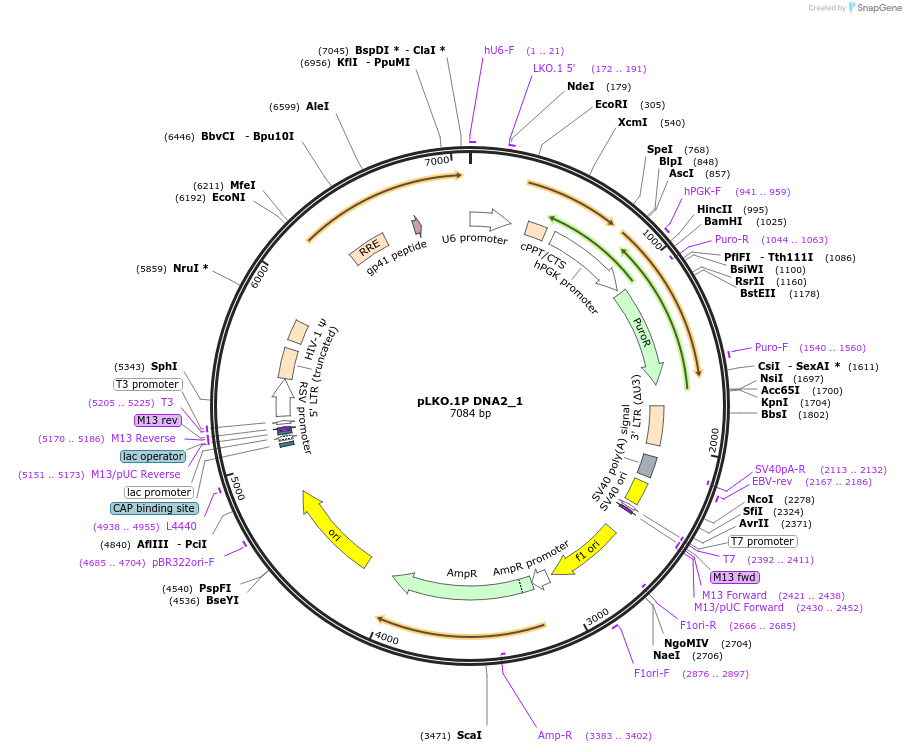
pLKO.1P DNA2_1
Plasmid#160779PurposeSuppress DNA2DepositorInsertshDNA2_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
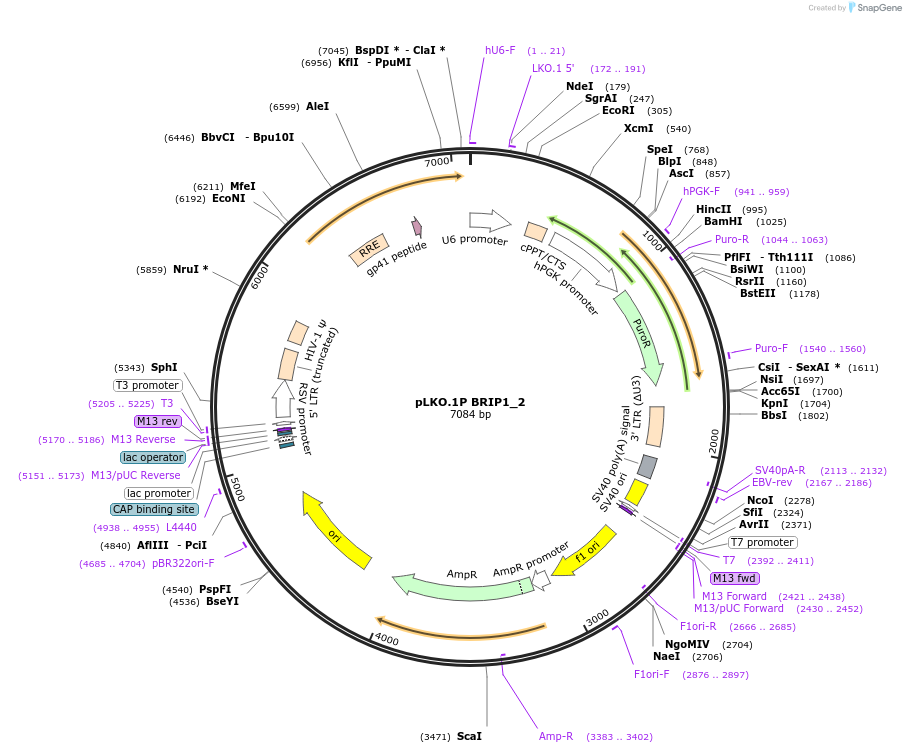
pLKO.1P BRIP1_2
Plasmid#160772PurposeSuppress BRIP1DepositorInsertshBRIP1_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
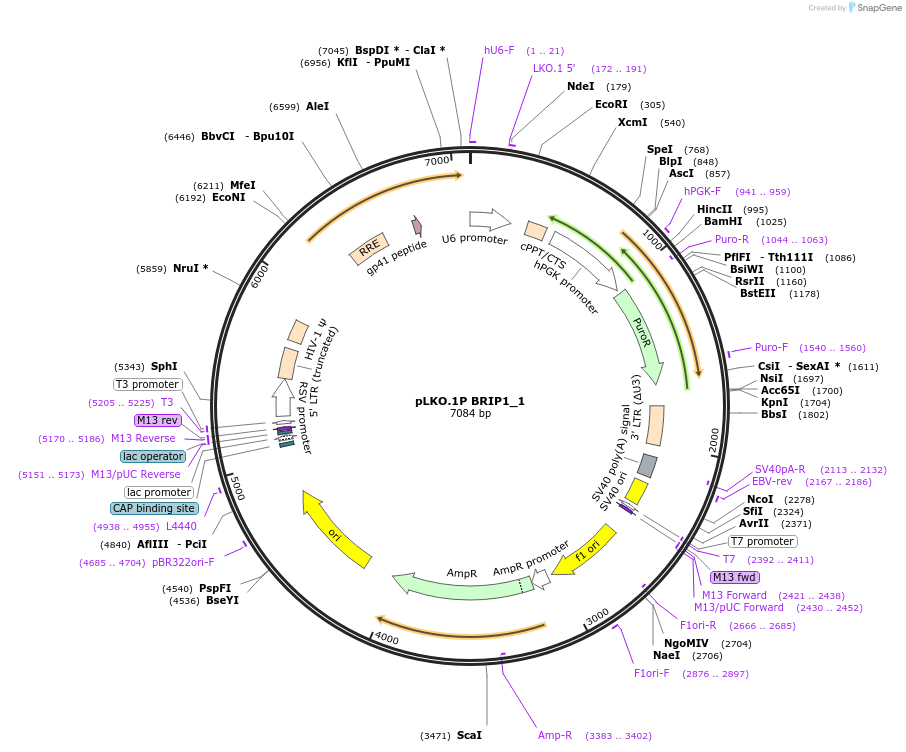
pLKO.1P BRIP1_1
Plasmid#160771PurposeSuppress BRIP1DepositorInsertshBRIP1_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
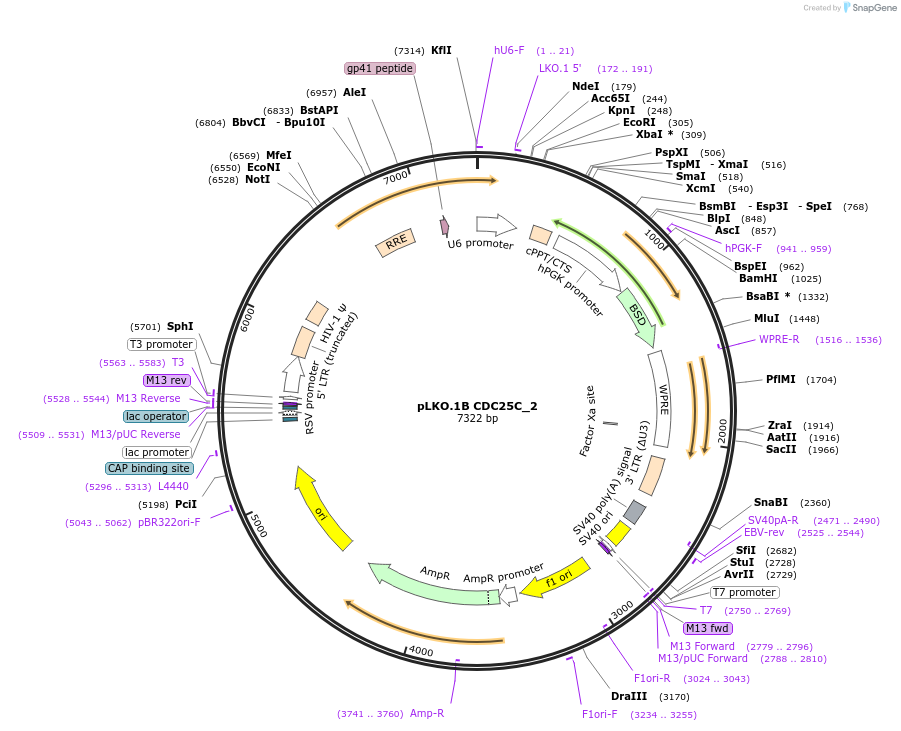
pLKO.1B CDC25C_2
Plasmid#160770PurposeSuppress CDC25CDepositorInsertshCDC25C_2
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
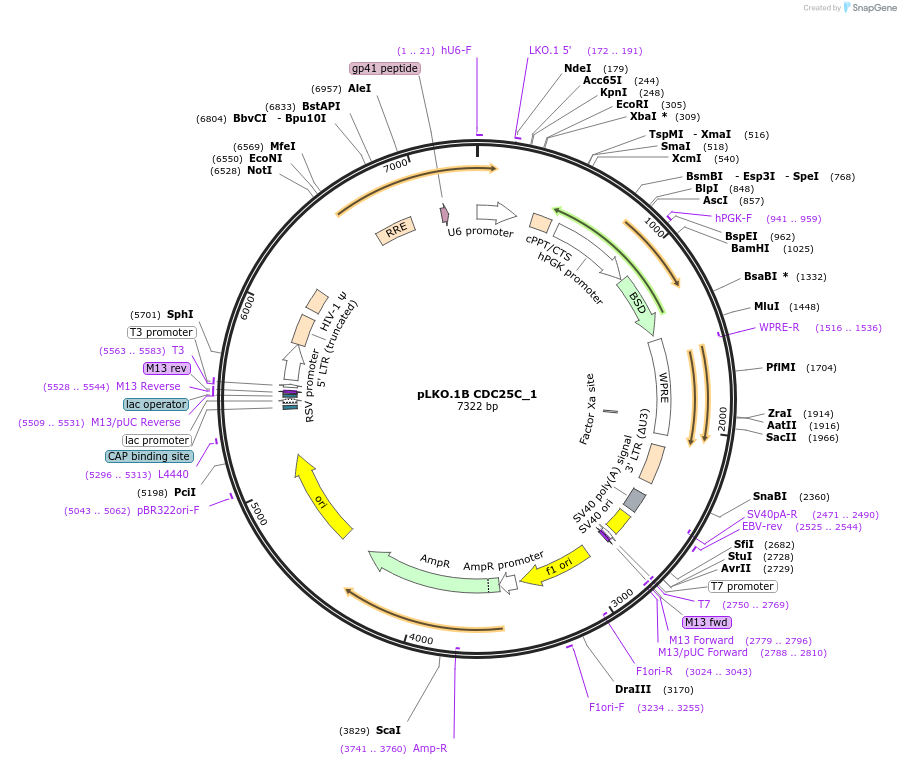
pLKO.1B CDC25C_1
Plasmid#160769PurposeSuppress CDC25CDepositorInsertshCDC25C_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
pLKO.1B CDC25A_1
Plasmid#160767PurposeSuppress CDC25ADepositorInsertshCDC25A_1
UseLentiviralAvailable SinceOct. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
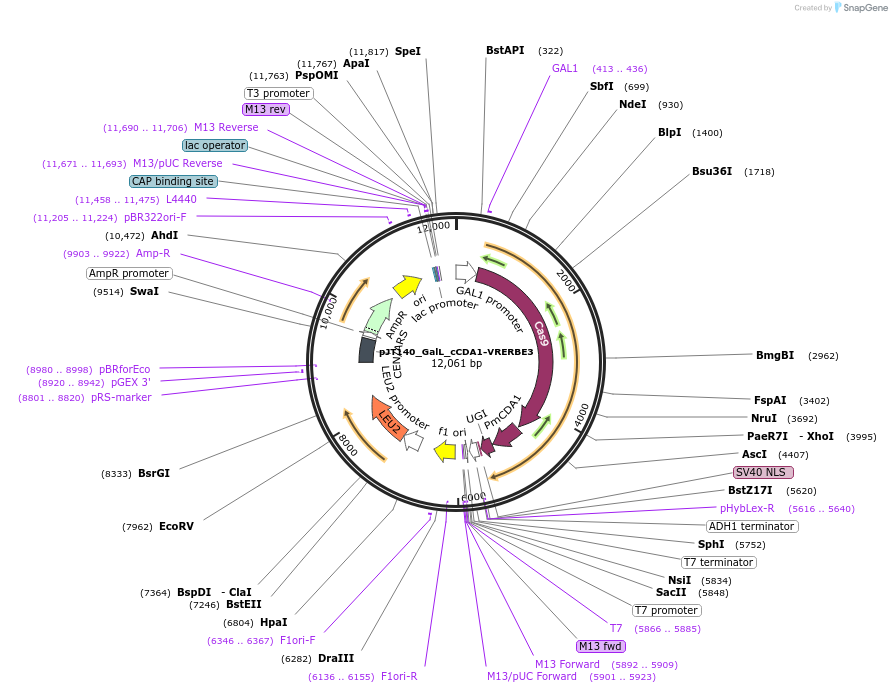
pJT140_GalL_cCDA1-VRERBE3
Plasmid#145103PurposeExpressing base editor cCDA1-VRERBE3 in yeast cellsDepositorInsertcCDA1-VRERBE3
UseCRISPRExpressionYeastMutationspCas9(D10A/D1135V/G1218R/R1335E/T1337R)PromoterGalLAvailable SinceSept. 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
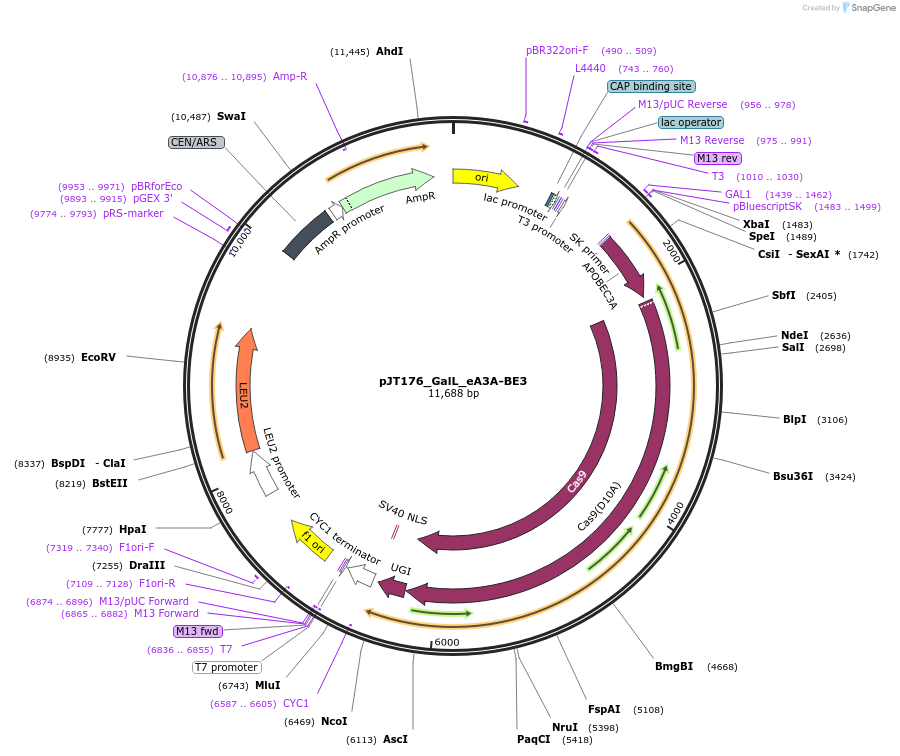
pJT176_GalL_eA3A-BE3
Plasmid#145125PurposeExpressing base editor eA3A-BE3 in yeast cellsDepositorInsertA3A(N57G)-BE3
UseCRISPRExpressionYeastMutationA3A(N57G); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
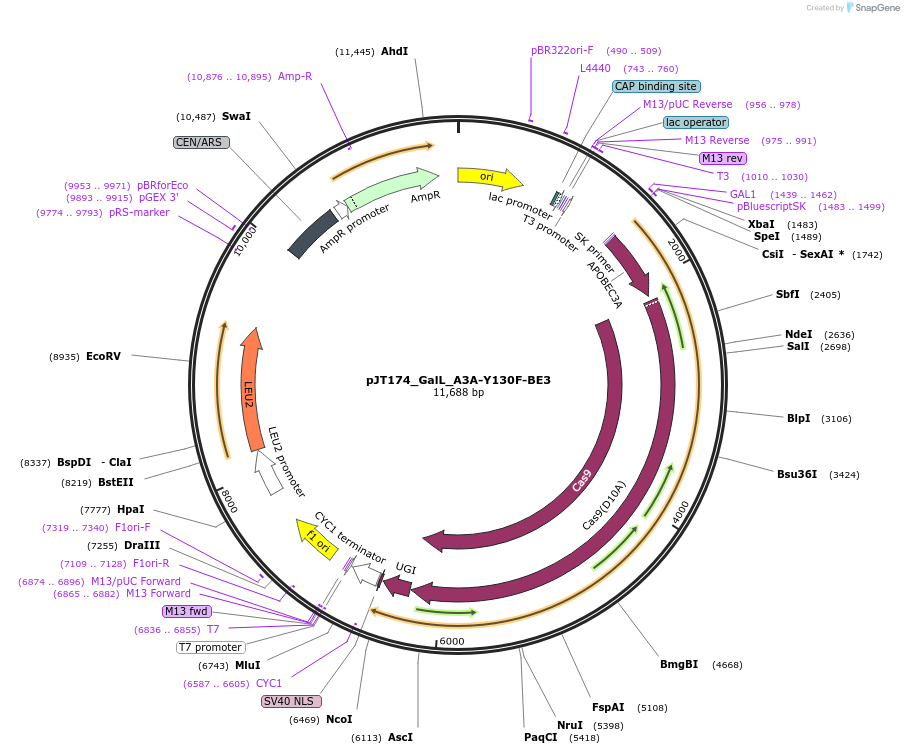
pJT174_GalL_A3A-Y130F-BE3
Plasmid#145123PurposeExpressing base editor A3A-Y130F-BE3 in yeast cellsDepositorInsertA3A(Y130F)-BE3
UseCRISPRExpressionYeastMutationA3A(Y130F); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
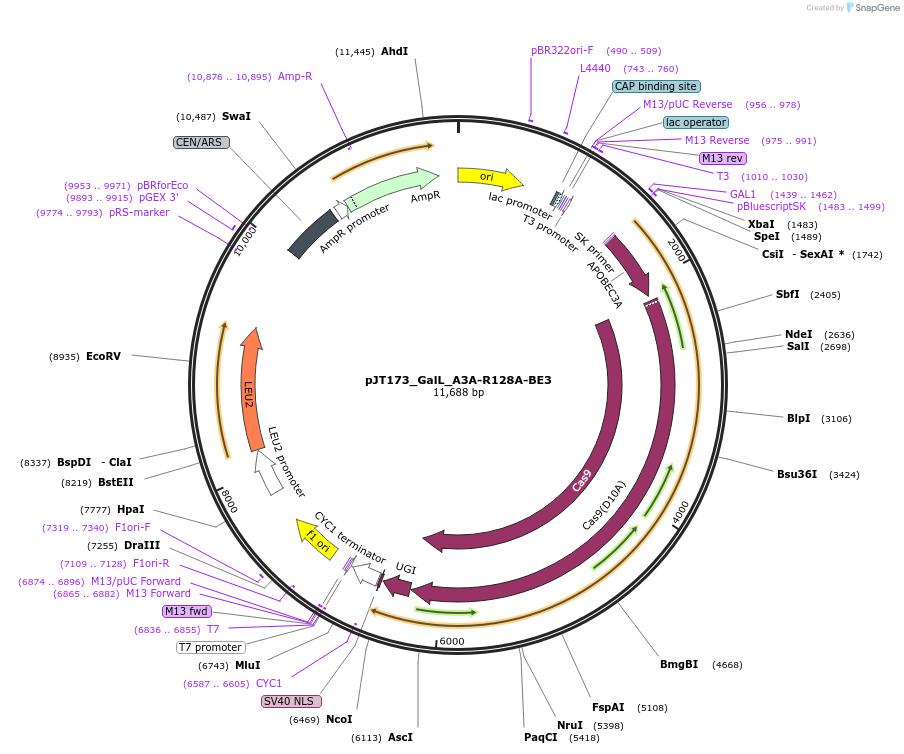
pJT173_GalL_A3A-R128A-BE3
Plasmid#145122PurposeExpressing base editor A3A-R128A-BE3 in yeast cellsDepositorInsertA3A(R128A)-BE3
UseCRISPRExpressionYeastMutationA3A(R128A); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
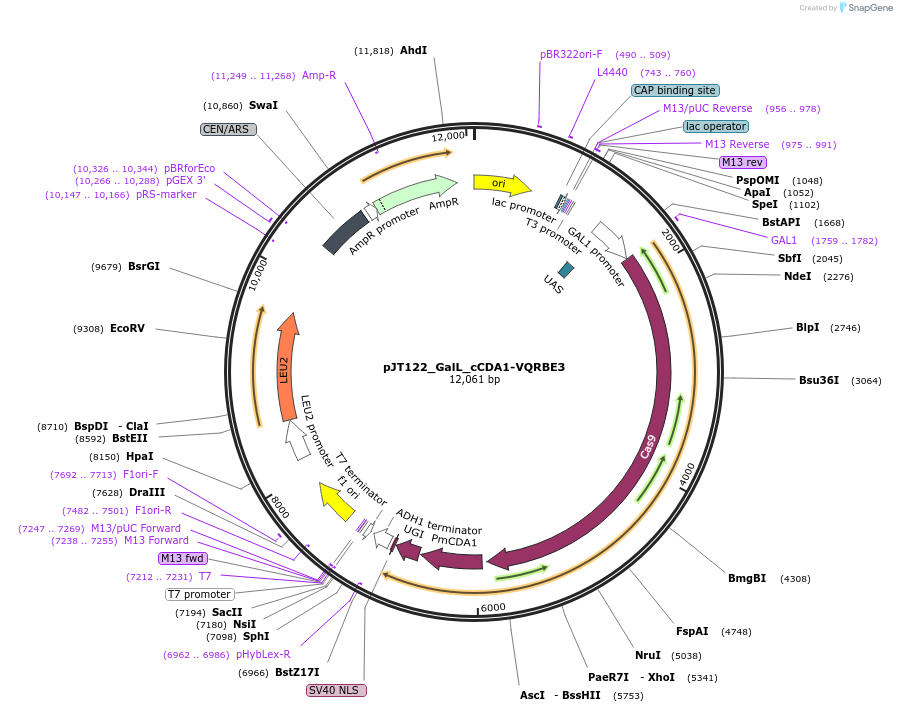
pJT122_GalL_cCDA1-VQRBE3
Plasmid#145087PurposeExpressing base editor cCDA1-VQRBE3 in yeast cellsDepositorInsertcCDA1-VQRBE3
UseCRISPRExpressionYeastMutationspCas9(D10A, D1135V, R1335Q, T1337R)PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
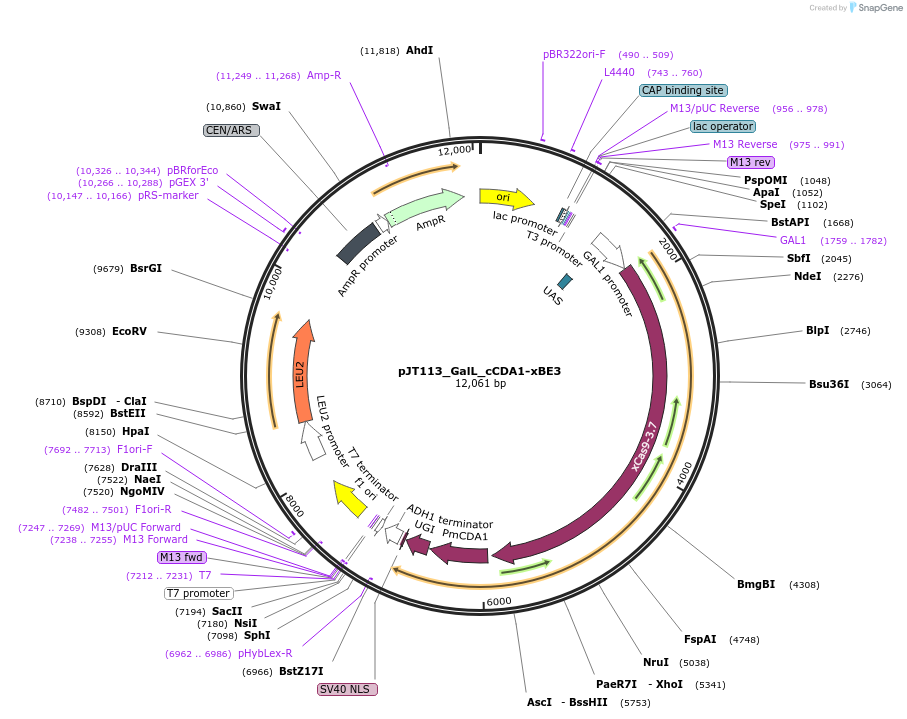
pJT113_GalL_cCDA1-xBE3
Plasmid#145079PurposeExpressing base editor cCDA1-xBE3 in yeast cellsDepositorInsertcCDA1-xBE3
UseCRISPRExpressionYeastMutationspCas9(D10A, A262T, R324L, S409I, E480K, E543D, M…PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
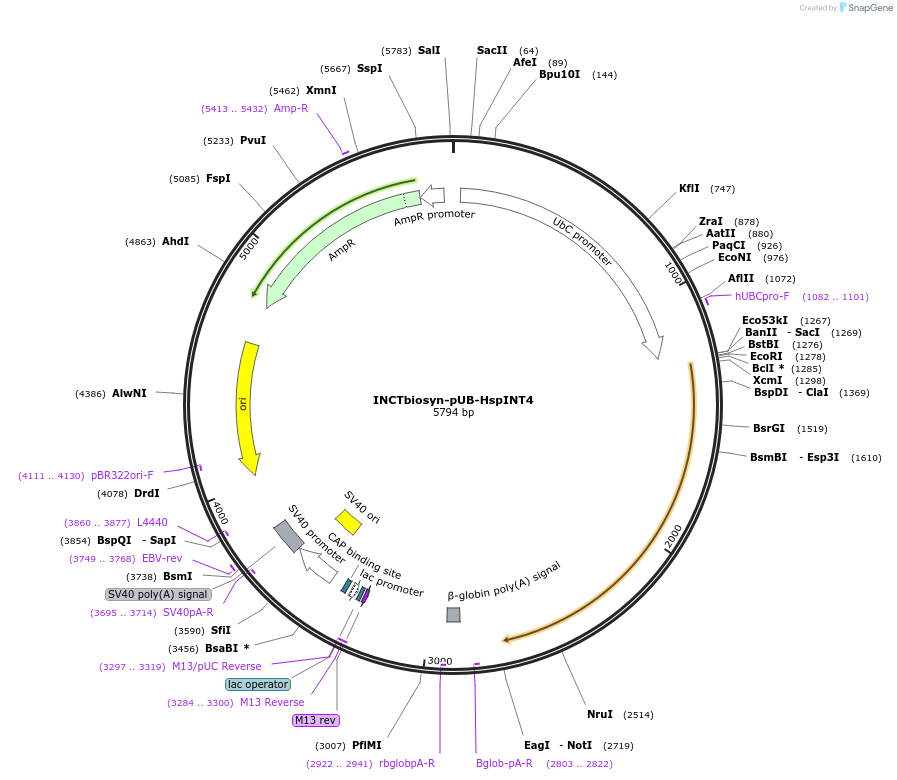
INCTbiosyn-pUB-HspINT4
Plasmid#127513PurposePlasmid encodes H. sapiens codon optimized Integrase 4.DepositorInsertIntegrase 4 coding sequence codon optimized for H. sapiens expression.
ExpressionMammalianAvailable SinceMay 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
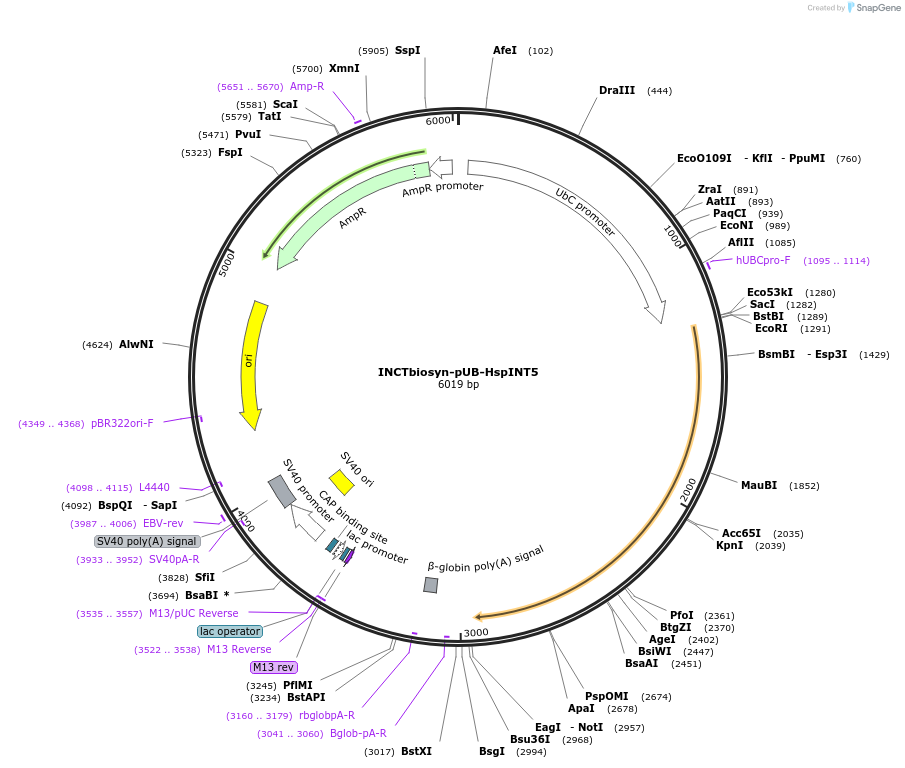
INCTbiosyn-pUB-HspINT5
Plasmid#127514PurposePlasmid encodes H. sapiens codon optimized Integrase 5.DepositorInsertIntegrase 5 coding sequence codon optimized for H. sapiens expression.
ExpressionMammalianAvailable SinceMay 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
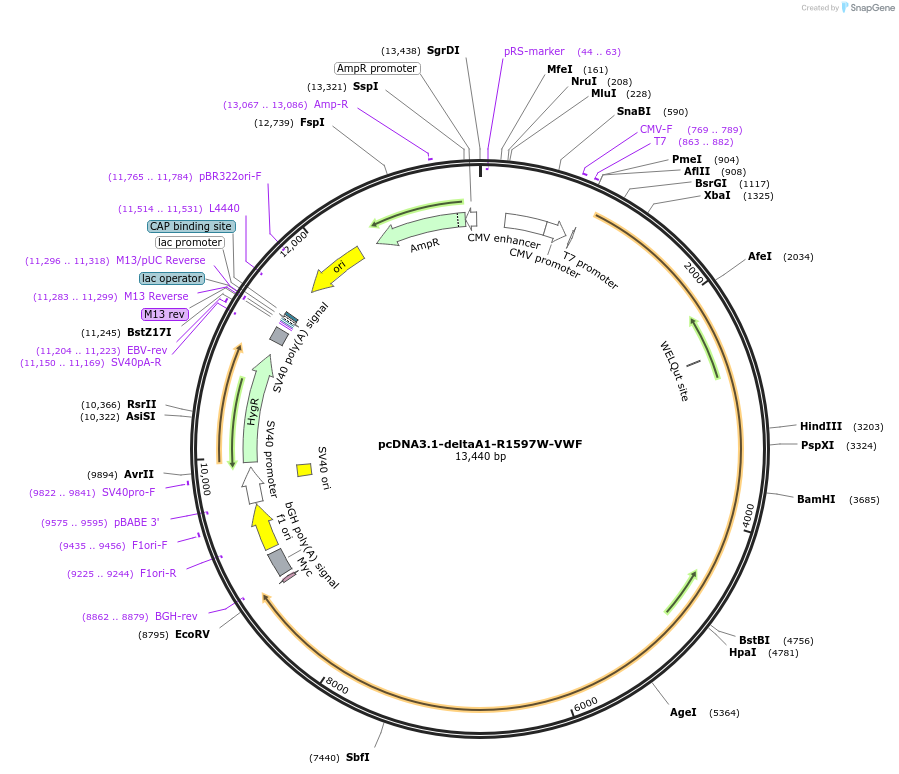
pcDNA3.1-deltaA1-R1597W-VWF
Plasmid#124826PurposeExpresses multimeric human VWF protein without A1 domain and a R1597W mutation in A2 domain. This protein has C-terminal c-Myc tag.DepositorInsertVWF without A1 domain and R1597W mutation A2 domain (VWF Human)
ExpressionMammalianPromoterCMVAvailable SinceMay 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
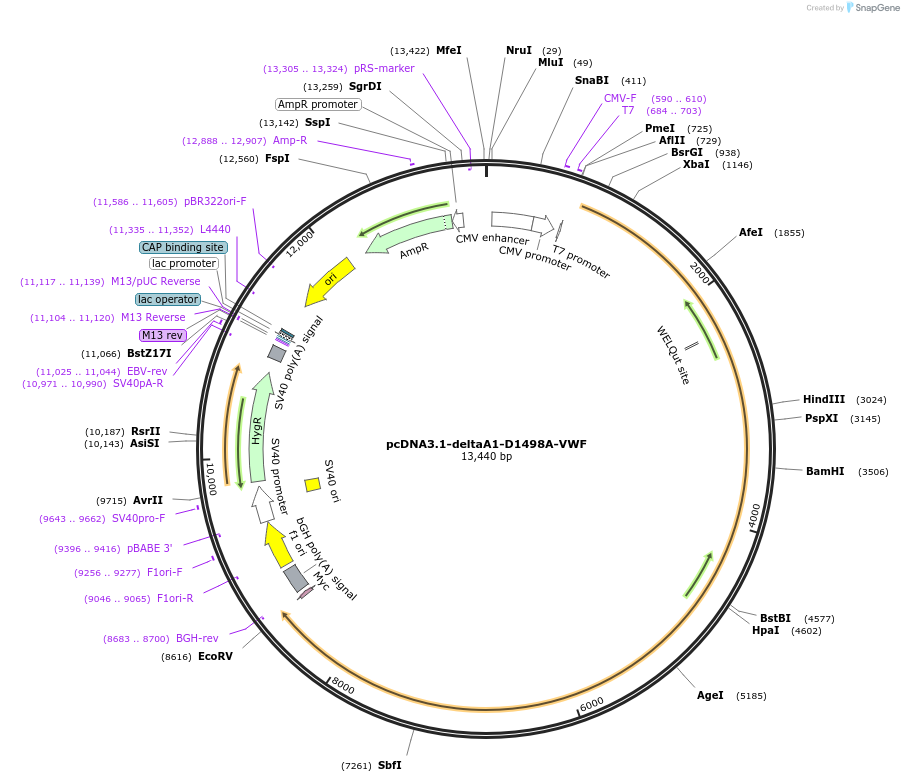
pcDNA3.1-deltaA1-D1498A-VWF
Plasmid#124825PurposeExpresses multimeric human VWF protein without A1 domain and a D1498A mutation in A2 domain. This protein has C-terminal c-Myc tag.DepositorInsertVWF without A1 domain and D1498A mutation A2 domain (VWF Human)
ExpressionMammalianPromoterCMVAvailable SinceMay 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
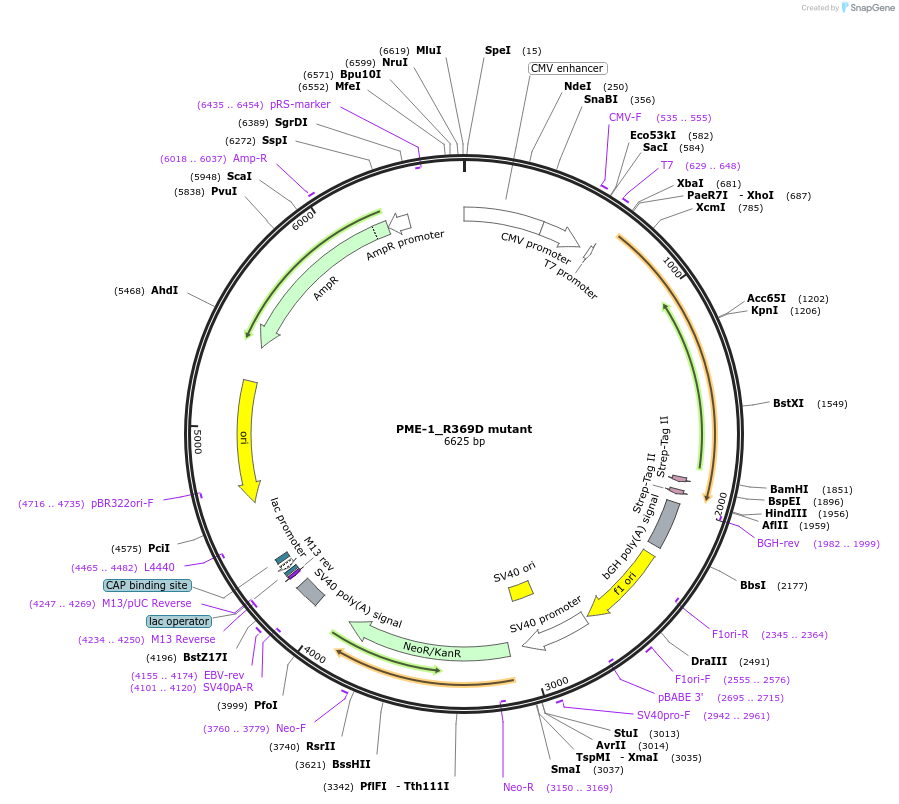
PME-1_R369D mutant
Plasmid#119294PurposeMammalian overexpression of StrepII-tagged full length PPME1 with mutation in the PP2Ac binding motifDepositorAvailable SinceJan. 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
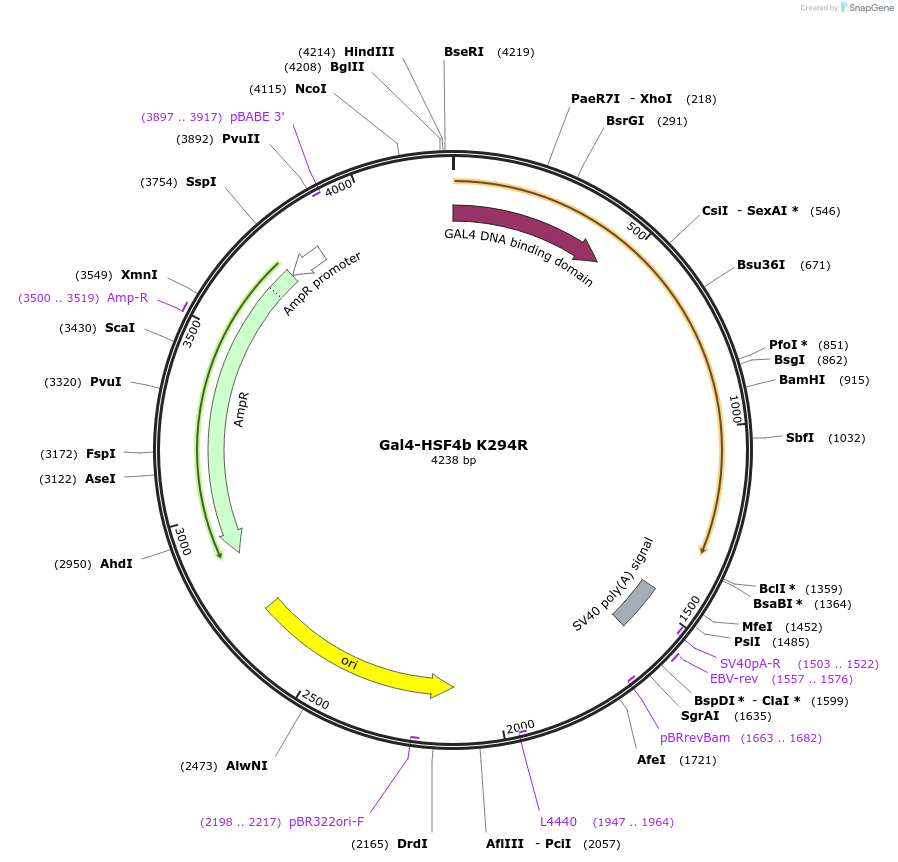
Gal4-HSF4b K294R
Plasmid#118359PurposeThis plasmid expresses amino-acids 200-493 of hHSF4b K294R mutant fused to the GAL-4 DNA binding domain. This construct cannot undergo sumoylation on K294.DepositorAvailable SinceNov. 20, 2018AvailabilityAcademic Institutions and Nonprofits only -
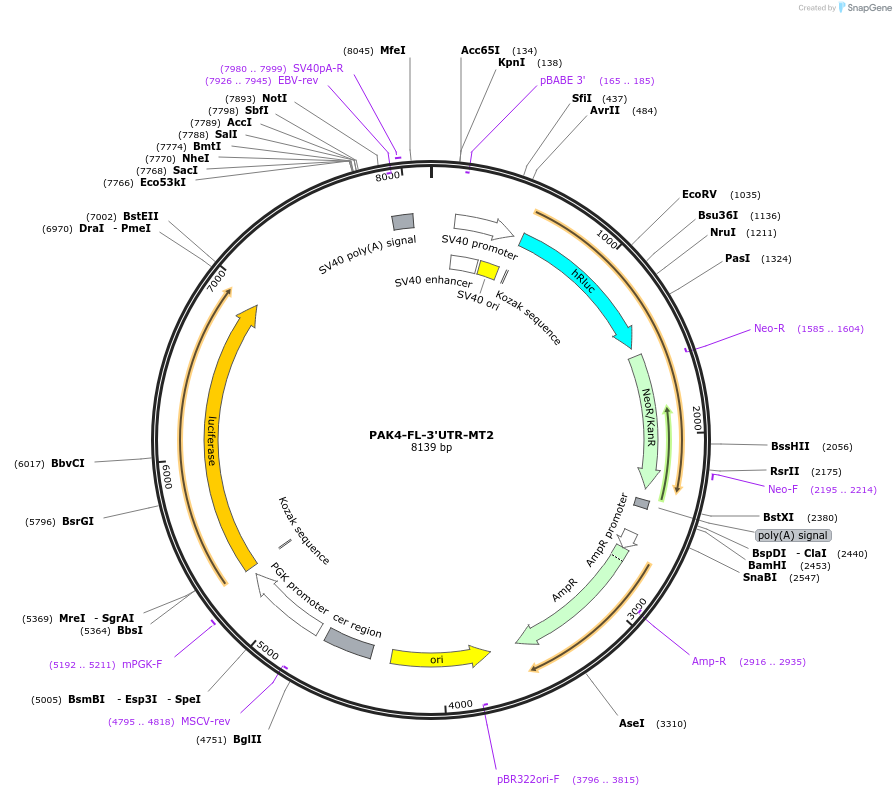
PAK4-FL-3'UTR-MT2
Plasmid#113093PurposeLuciferase reporter containing PAK4-FL- 3' UTR with the second predicted MiR-199a-3p binding site mutatedDepositorInsertp21 Activated kinase 4 (PAK4 Human)
UseLuciferaseExpressionMammalianMutationSecond predicted miR-199a-3p binding site mutated…PromoterPGKAvailable SinceAug. 17, 2018AvailabilityAcademic Institutions and Nonprofits only