We narrowed to 13,335 results for: ache
-
Plasmid#215246PurposeCDS of Luz gene from N. nambi codon optimized for N.benthamiana. This gene generate the luminiscence of the bioluminiscente pathwayDepositorInsertLuz
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceJune 18, 2024AvailabilityAcademic Institutions and Nonprofits only -
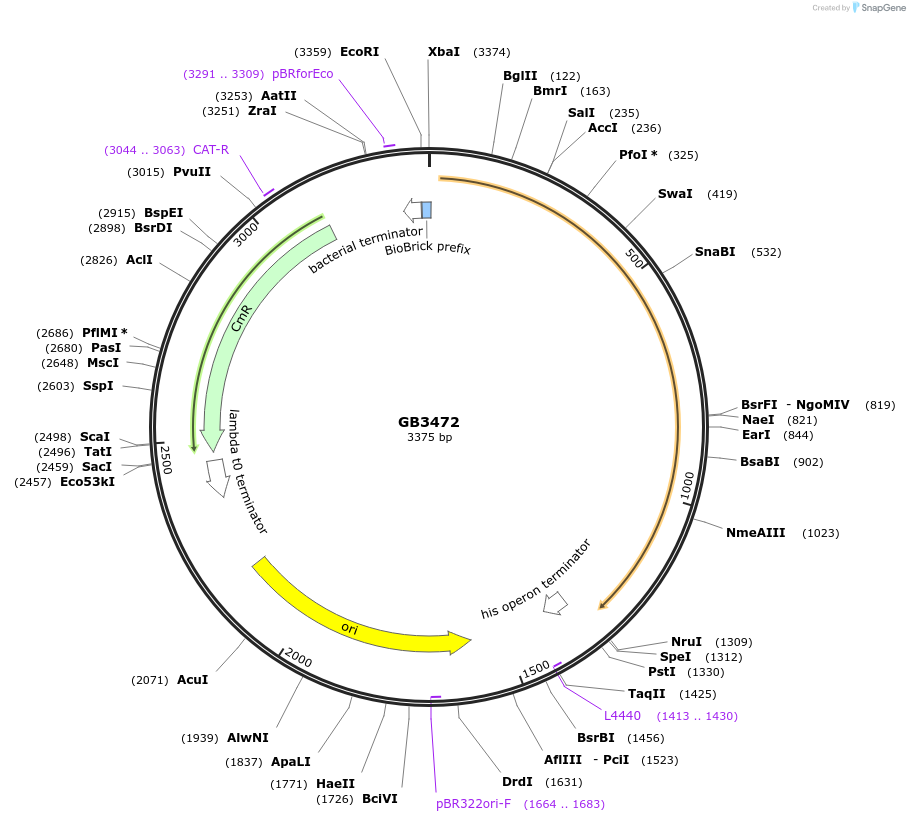
GB3472
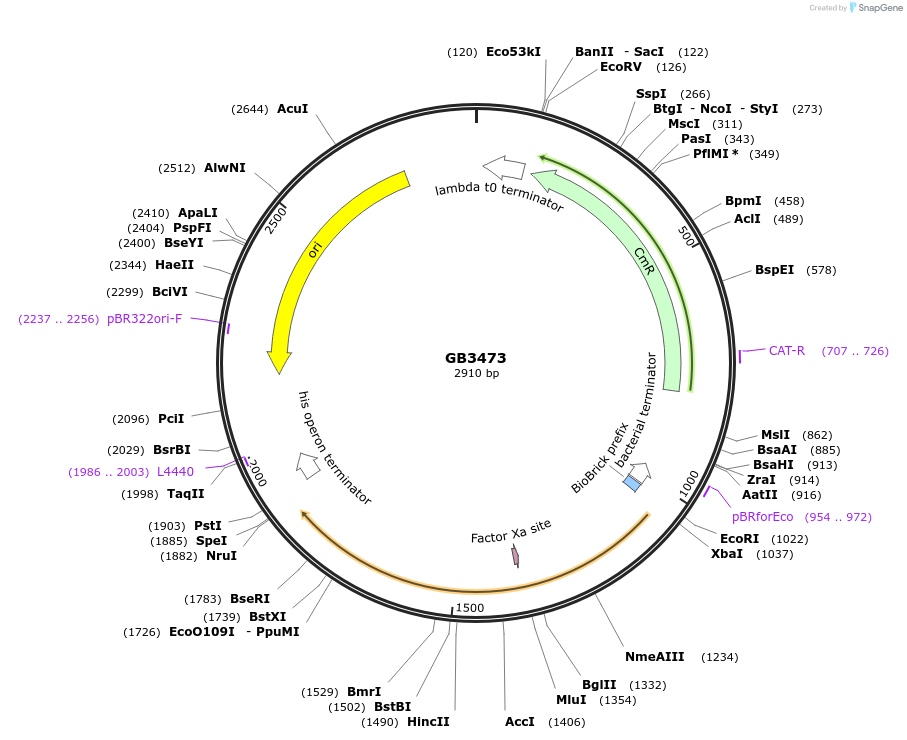
Plasmid#215245PurposeCDS of H3H from N. nambi codon optimized for N.benthamiana. This gene is the second step of the bioluminiscente pathwayDepositorInsertH3H
UseSynthetic BiologyMutationBsaI and BsmBI sites removedAvailable SinceJune 18, 2024AvailabilityAcademic Institutions and Nonprofits only -
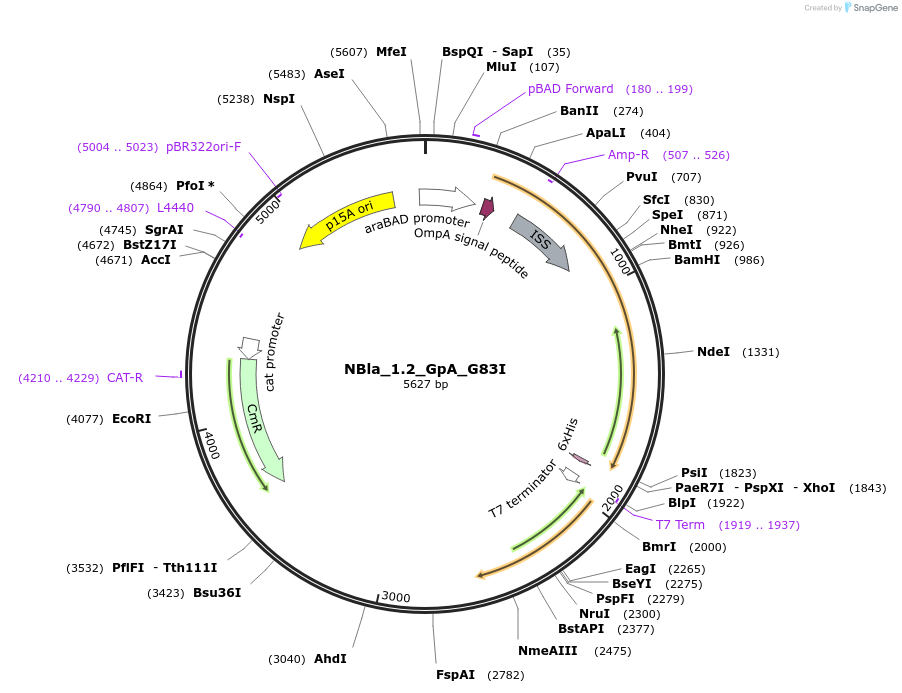
NBla_1.2_GpA_G83I
Plasmid#207844PurposeBLaTM-System GpA G83I negative control in NBLa 1.2 vectorDepositorInsertN-BLa 1.2 fusion protein
TagsFlagExpressionBacterialAvailable SinceJan. 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
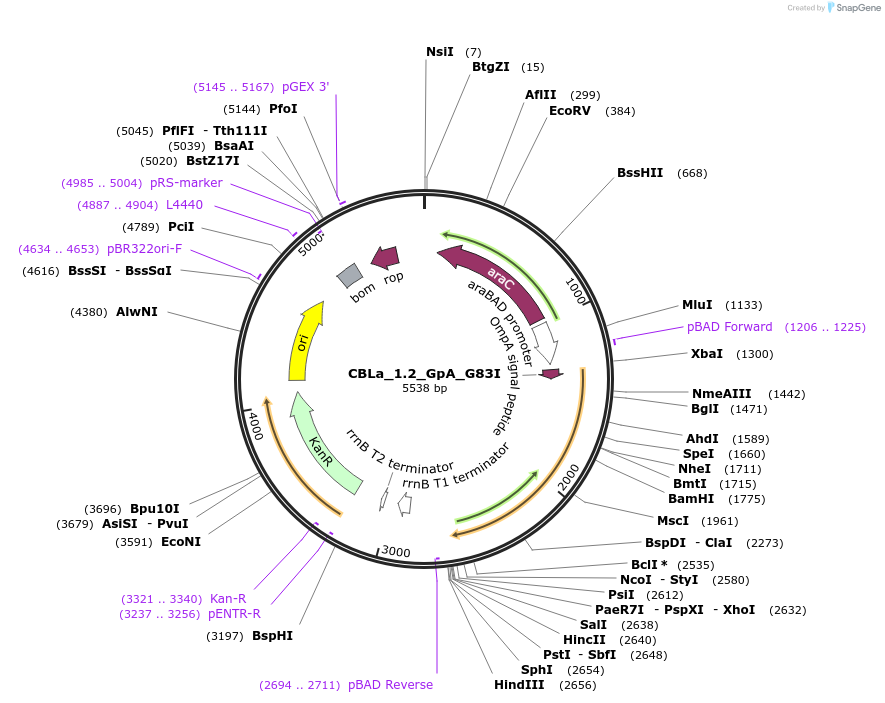
CBLa_1.2_GpA_G83I
Plasmid#207845PurposeBLaTM-System GpA G83I negative control in CBLa 1.2 vectorDepositorInsertN-BLa 1.2 fusion protein
TagsFlagExpressionBacterialAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
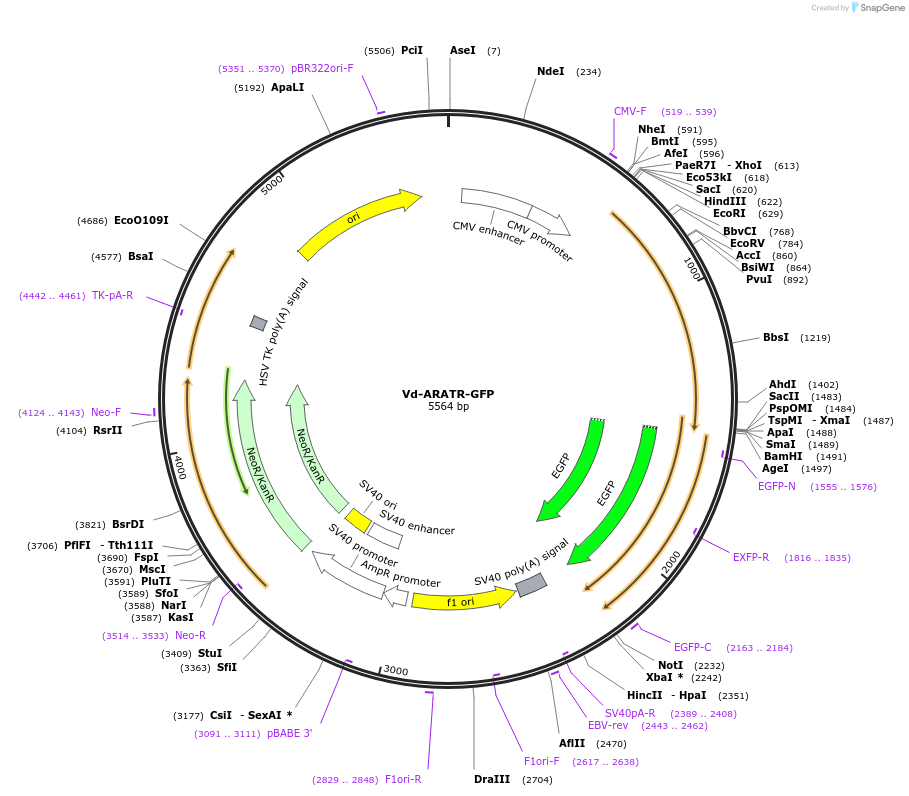
Vd-ARATR-GFP
Plasmid#208903Purposeencodes the arachnotocin receptor (oxytocin/vasopressin-like) from the mite Varroa destructor, tagged with GFPDepositorInsertARATR
TagsEGFPExpressionMammalianAvailable SinceNov. 1, 2023AvailabilityAcademic Institutions and Nonprofits only -
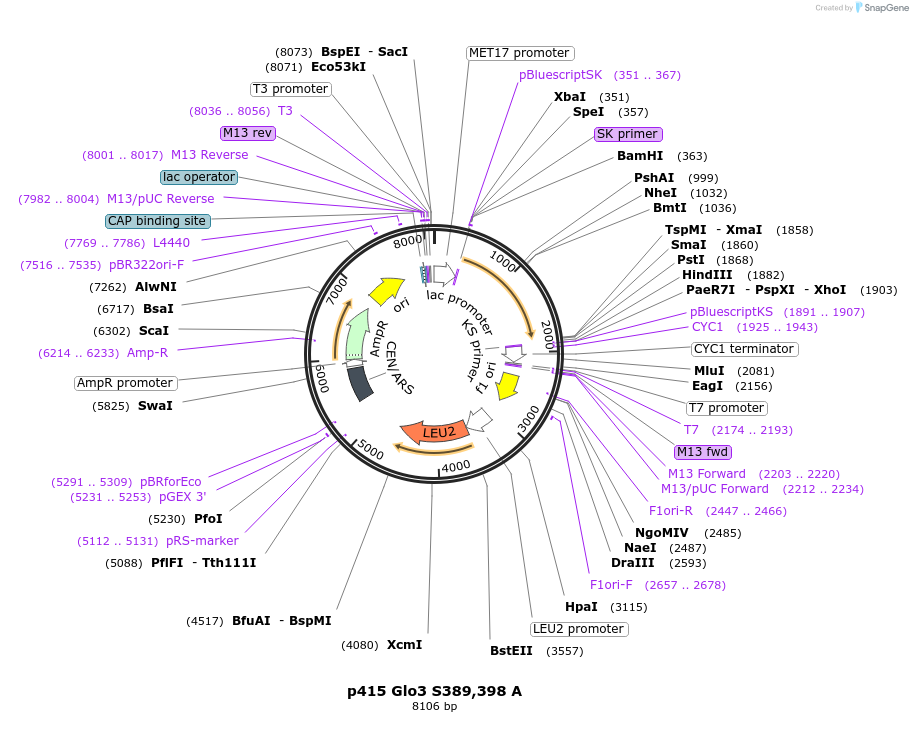
p415 Glo3 S389,398 A
Plasmid#129485PurposeExpression in S. cerevisiaeDepositorInsertGlo3 S389,398 A (GLO3 Budding Yeast)
ExpressionYeastMutationMutation of serines 389 and 398 to alanineAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
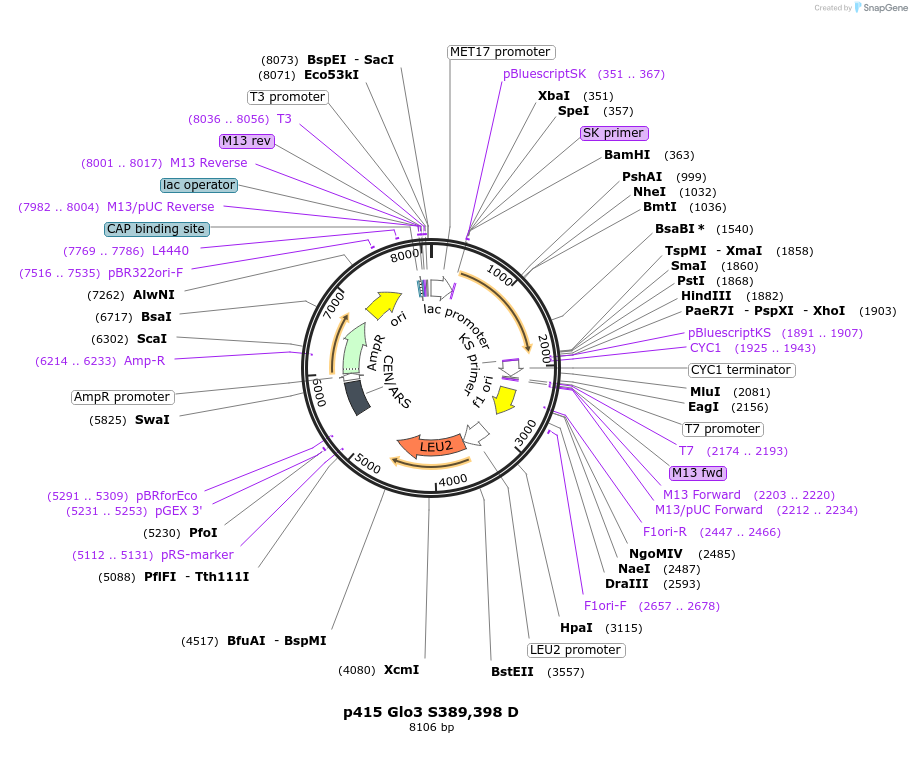
p415 Glo3 S389,398 D
Plasmid#129486PurposeExpression in S. cerevisiaeDepositorInsertGlo3 S389,398 D (GLO3 Budding Yeast)
ExpressionYeastMutationMutation of serines 389 and 398 to AspAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
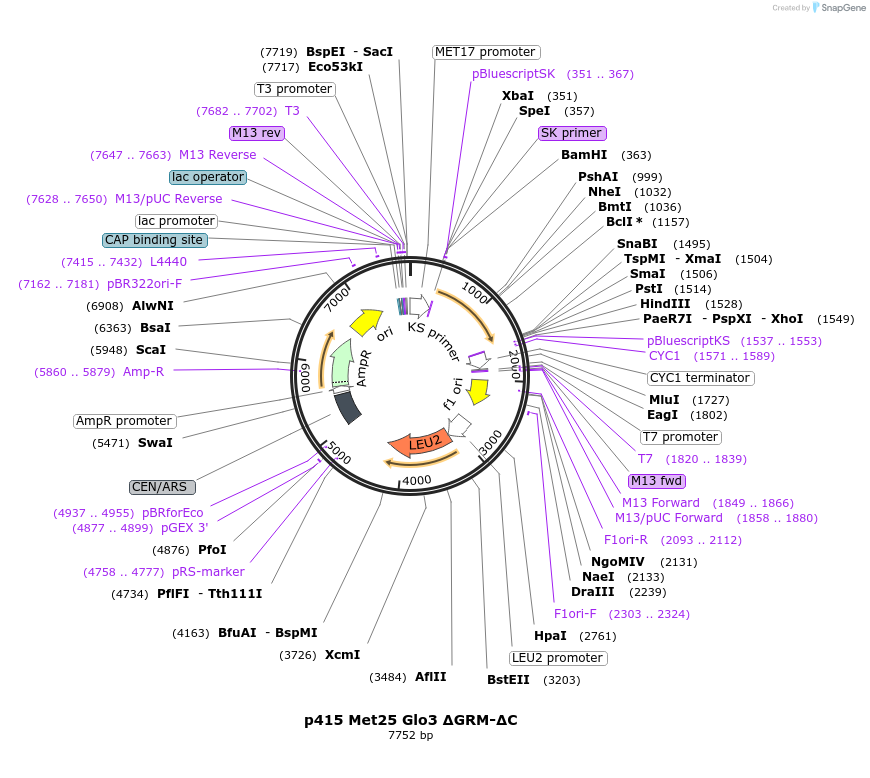
p415 Met25 Glo3 ΔGRM-ΔC
Plasmid#129487PurposeExpression in S. cerevisiaeDepositorInsertGlo3 ΔGRM ΔC (GLO3 Budding Yeast)
ExpressionYeastMutationΔ376-493 (deletion of Glo3 motif & C-terminal…Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
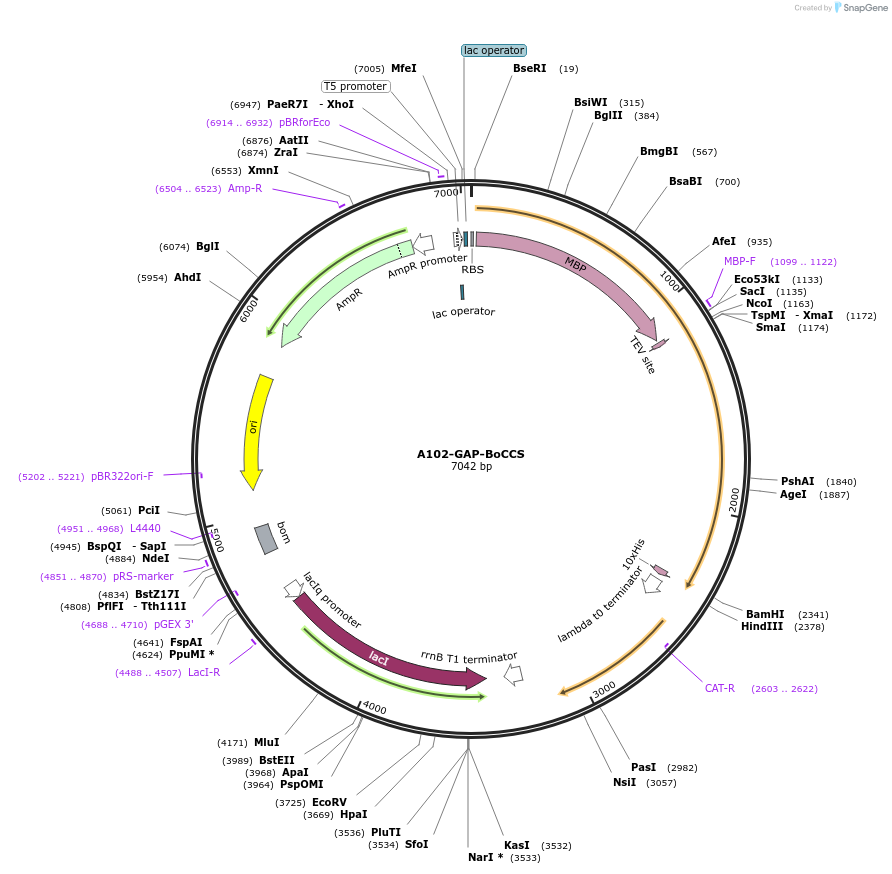
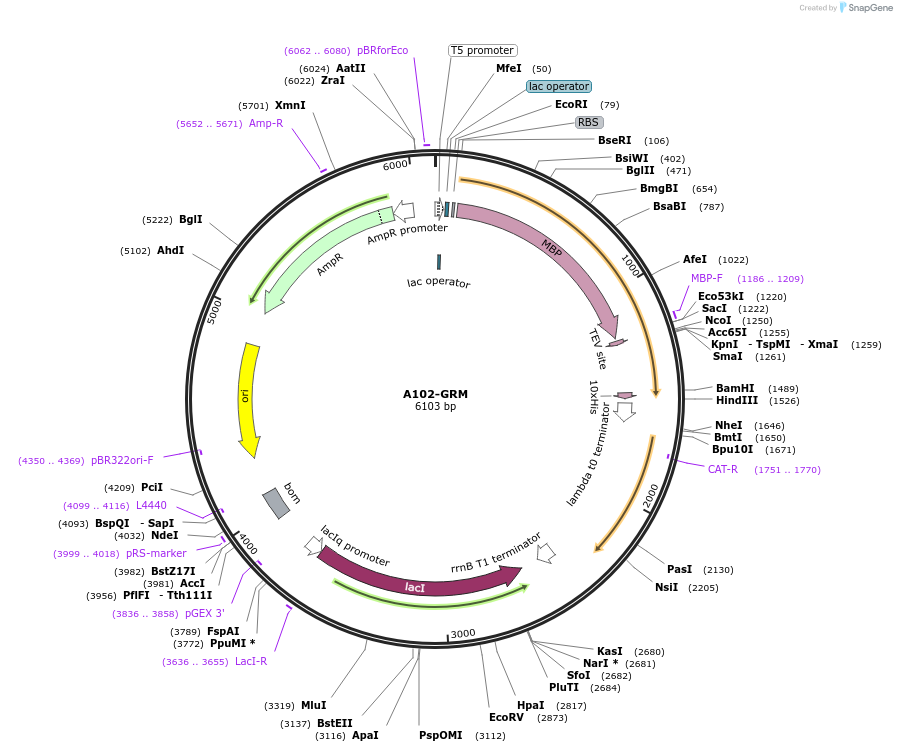
A102-GAP-BoCCS
Plasmid#123287PurposeExpression of truncated Glo3 (1-375) in E.ColiDepositorAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
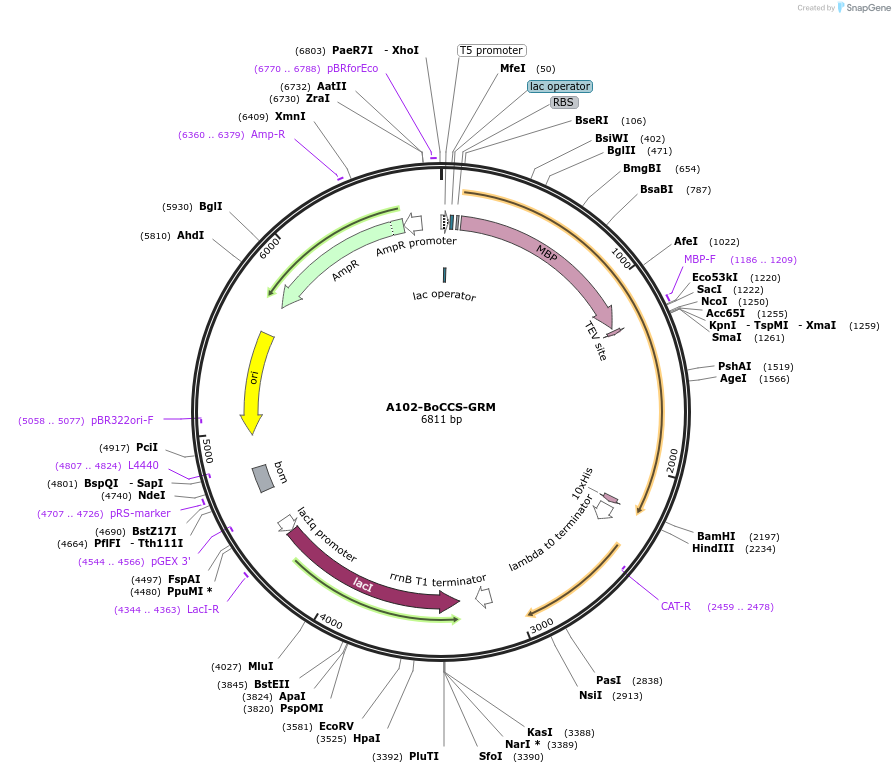
A102-BoCCS-GRM
Plasmid#123288PurposeExpression of truncated Glo3 (137-459) in E.ColiDepositorAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
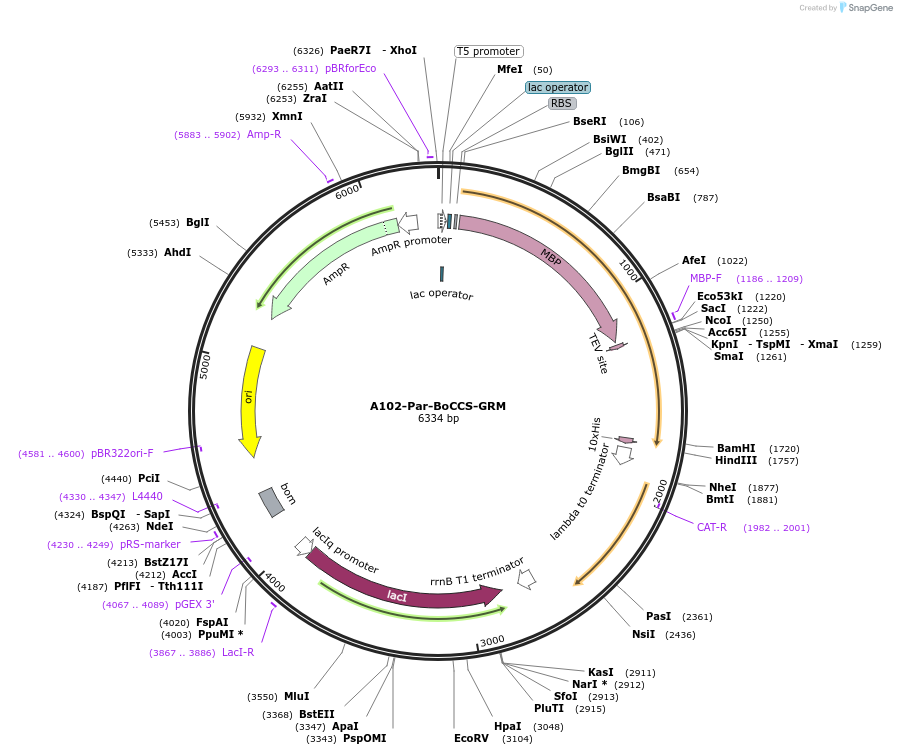
A102-Par-BoCCS-GRM
Plasmid#123289PurposeExpression of truncated Glo3 (296-459) in E.ColiDepositorAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
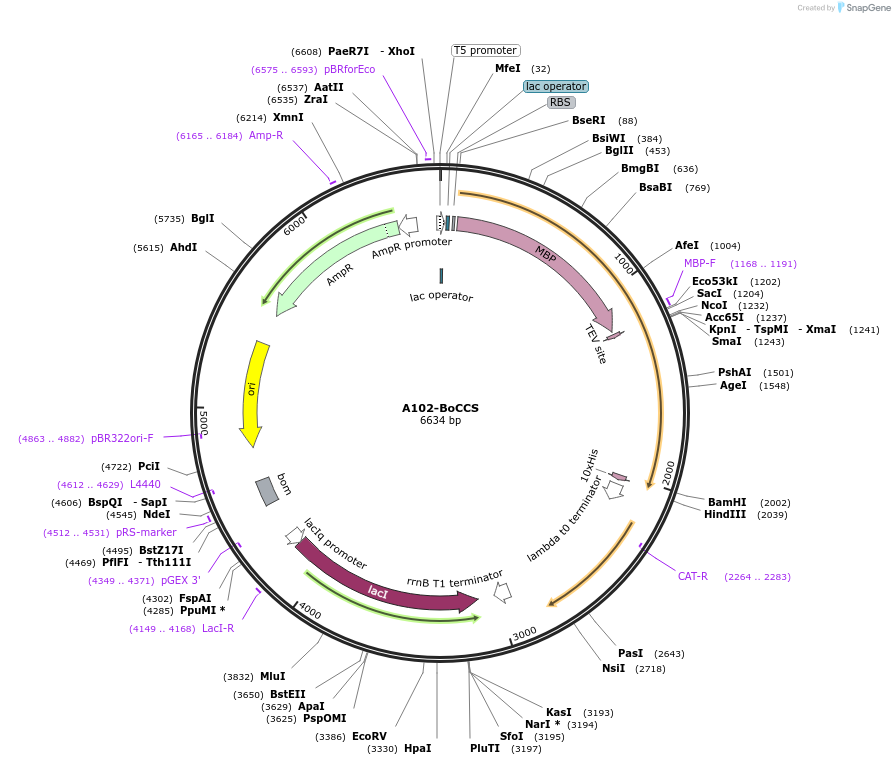
A102-BoCCS
Plasmid#123290PurposeExpression of truncated Glo3 (137-375) in E.ColiDepositorAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
A102-GRM
Plasmid#123291PurposeExpression of truncated Glo3 (373-459) in E.ColiDepositorAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
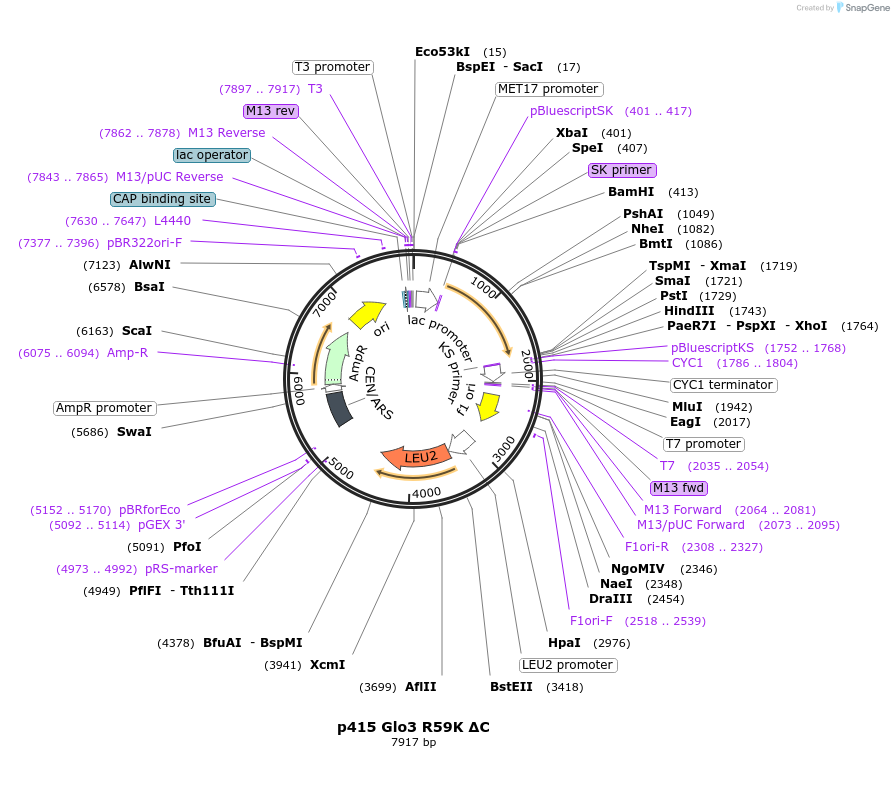
p415 Glo3 R59K ΔC
Plasmid#112653PurposeExpression S.cerevisiaeDepositorInsertGlo3 R59K ΔC
ExpressionYeastMutationR59K, Δ431-493 (deletion of C-terminal amphipathi…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
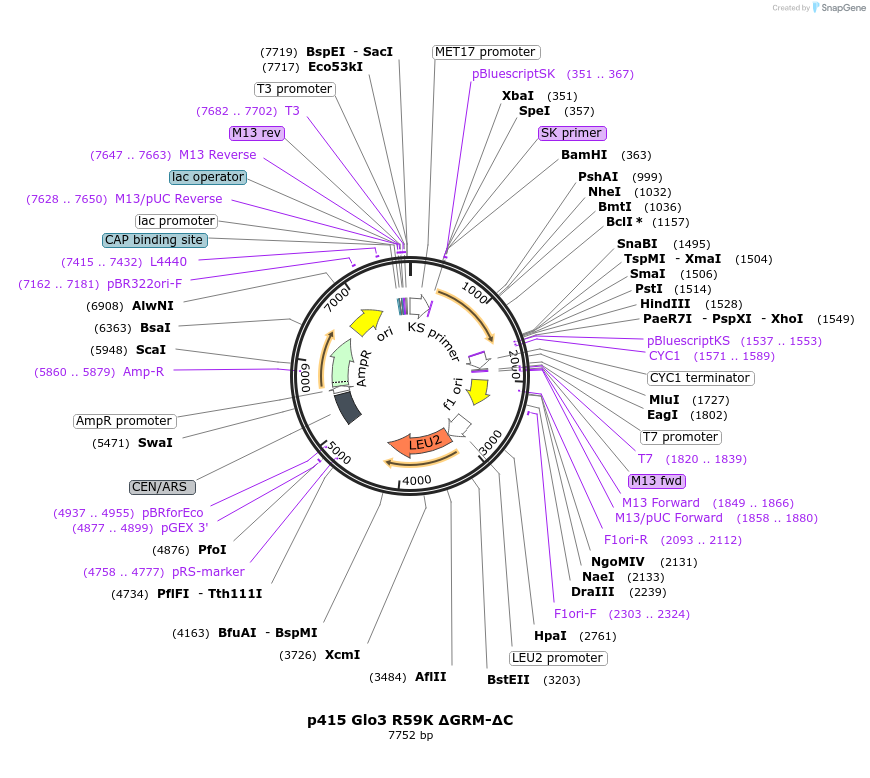
p415 Glo3 R59K ΔGRM-ΔC
Plasmid#112654PurposeExpression S.cerevisiaeDepositorInsertGlo3 R59K ΔGRM ΔC
ExpressionYeastMutationR59K, Δ376-493 (deletion of Glo3 motif & C-te…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
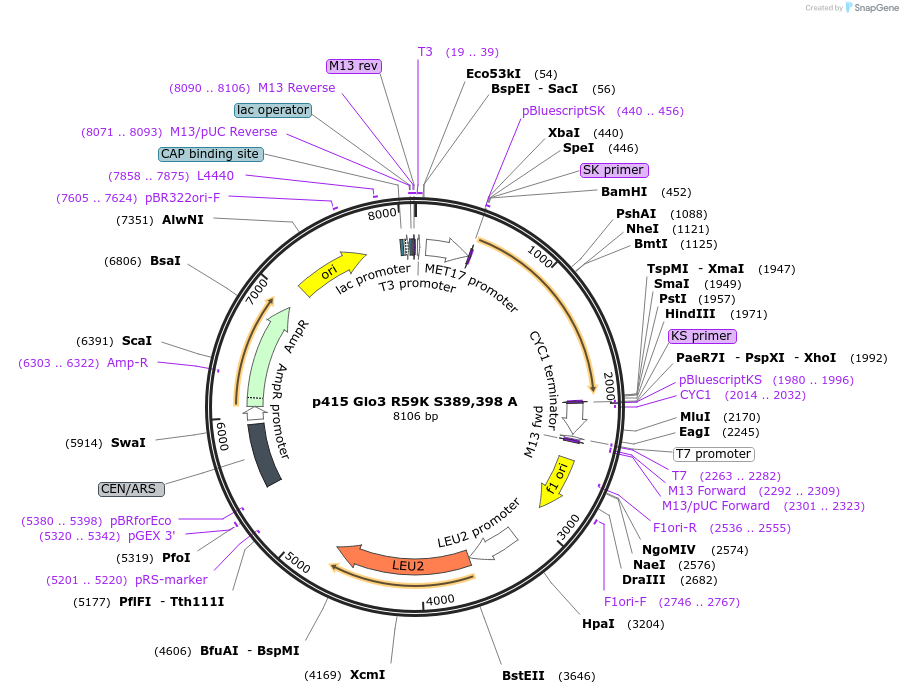
p415 Glo3 R59K S389,398 A
Plasmid#112656PurposeExpression S.cerevisiaeDepositorInsertGlo3 R59K S389,398A
ExpressionYeastMutationR59K, mutations (S389A, S398A) in Glo3 motif, non…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
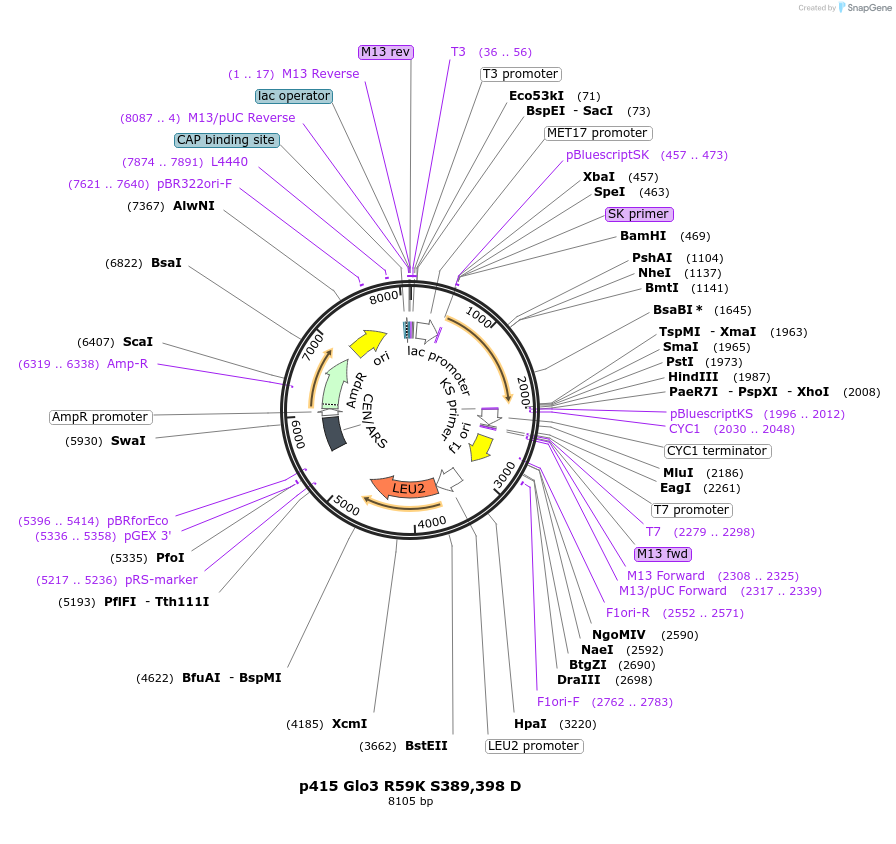
p415 Glo3 R59K S389,398 D
Plasmid#112657PurposeExpression S.cerevisiaeDepositorInsertGlo3 R59K S389,398D
ExpressionYeastMutationR59K, mutations (S389D, S398D) in Glo3 motif, pho…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
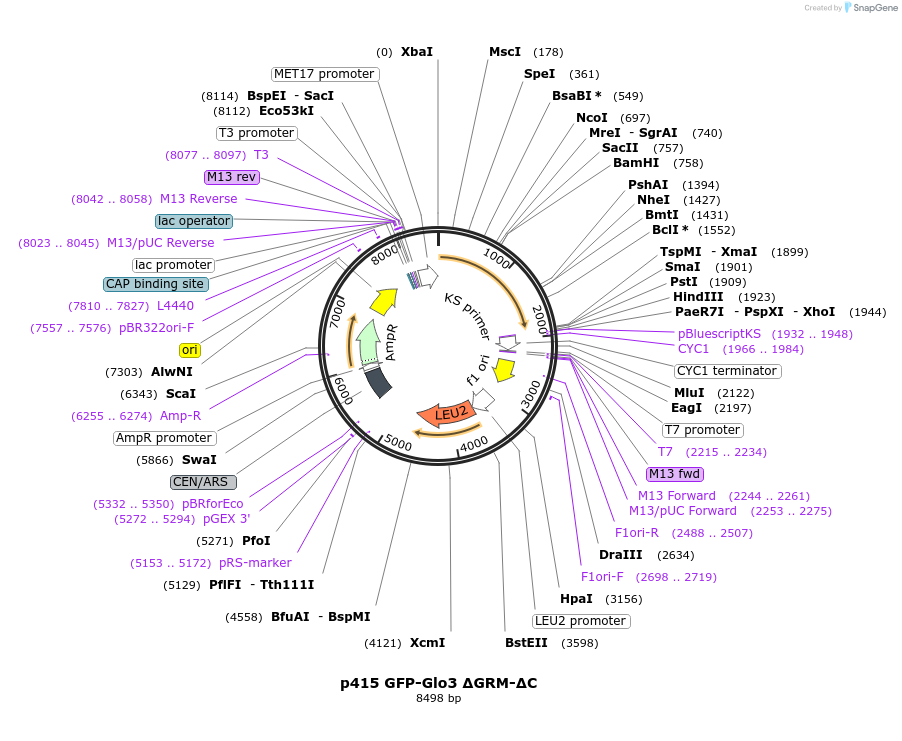
p415 GFP-Glo3 ΔGRM-ΔC
Plasmid#112659PurposeExpression S.cerevisiaeDepositorInsertGFP-Glo3 ΔGRM ΔC
TagssfGFPExpressionYeastMutationΔ376-493 (deletion of Glo3 motif & C-terminal…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
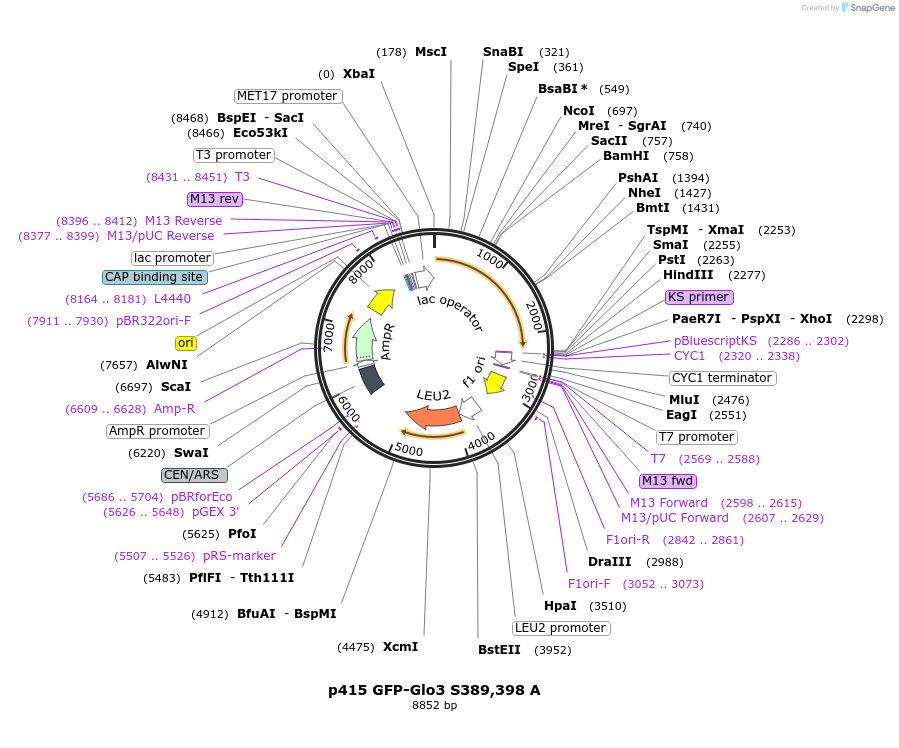
p415 GFP-Glo3 S389,398 A
Plasmid#112660PurposeExpression S.cerevisiaeDepositorInsertGFP- Glo3 S389,398A
TagssfGFPExpressionYeastMutationmutations (S389A, S398A) in Glo3 motif, non-phosp…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
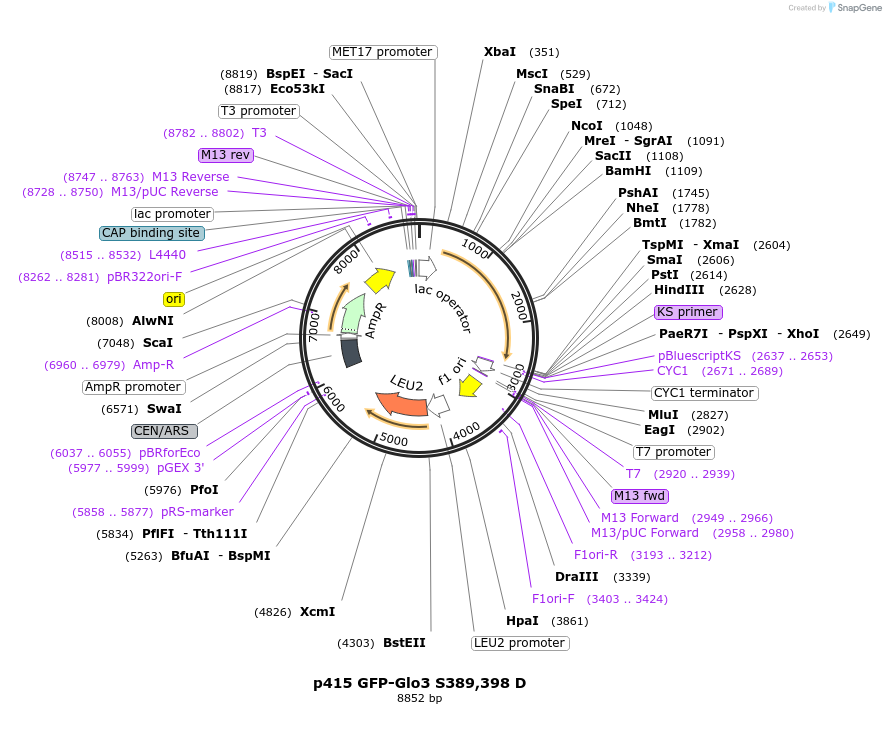
p415 GFP-Glo3 S389,398 D
Plasmid#112661PurposeExpression S.cerevisiaeDepositorInsertGFP-Glo3 S389,398D
TagssfGFPExpressionYeastMutationmutations (S389D, S398D) in Glo3 motif, phosphomi…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only