Cultivarium JERBOA Toolkit
(Kit #
1000000275
)
Depositing Lab: Nili Ostrov, Charlie Gilbert, Cultivarium Tools
The JERBOA Toolkit (Jumping Element libRaries with Broad hOst rAnge) is a Golden Gate genetic toolkit developed to help researchers identify functional transposons and establish genetic tractability in diverse non-model bacteria. The kit enables high-throughput, pooled library screening of thousands of transposon configurations (varying transposases, antibiotic resistance markers, and promoter-RBS elements driving their expression) to determine the optimal construct for a specific host.
This kit will be sent as bacterial glycerol stocks in 96-well plate format.
Original Publication
A scalable transposon mutagenesis system for non-model bacteria Gilbert C, Leung J, Crits-Christoph A, Kang SC, Esmurria A, Fenn K, Brumwell SL, Martin-Moldes Z, Mendler K, Barnum TP, Nguyen M-A, Lee HH, Ostrov N. bioRxiv 2025.12.22.696024. doi: 10.64898/2025.12.22.696024. Article (Link opens in a new window).
Description
Establishing genome-wide transposon mutagenesis in non-model microbes is often hindered by unpredictable host factors affecting transposase activity and selectable marker performance. The JERBOA Toolkit addresses this by providing a standardized, scalable framework for transposon variant screening.
The toolkit consists of the following modular parts that can be assembled to create transposon variant libraries:
- Transposase systems: both hyperactive Himar1 (mariner) and Tn5 transposases.
- Selectable markers: six commonly-used bacterial antibiotic resistance (ABR) genes (KAN, GEN, ERM, TET, CAM, and SPEC).
- Promoter variants: 18 diverse promoter-5' UTR sequences (including 16 broad-host-range promoters, a "dummy" negative control promoter, and standard resistance marker gene/transposase gene promoters) to independently drive the transposase and ABR marker.
- Combinatorial scale: the system supports up to 3,888 unique configurations (2 transposases × 6 markers × 18 transposase promoters × 18 ABR promoters).
Each unique final transposon variant plasmid possesses a unique DNA barcode within the transposon, allowing for quantification of transposition efficiency via Tn-seq or Semi-arbitrary (SemiArb) PCR. This approach has been validated across 92 non-model bacteria, identifying functional tools for species with no previously described genetic systems.
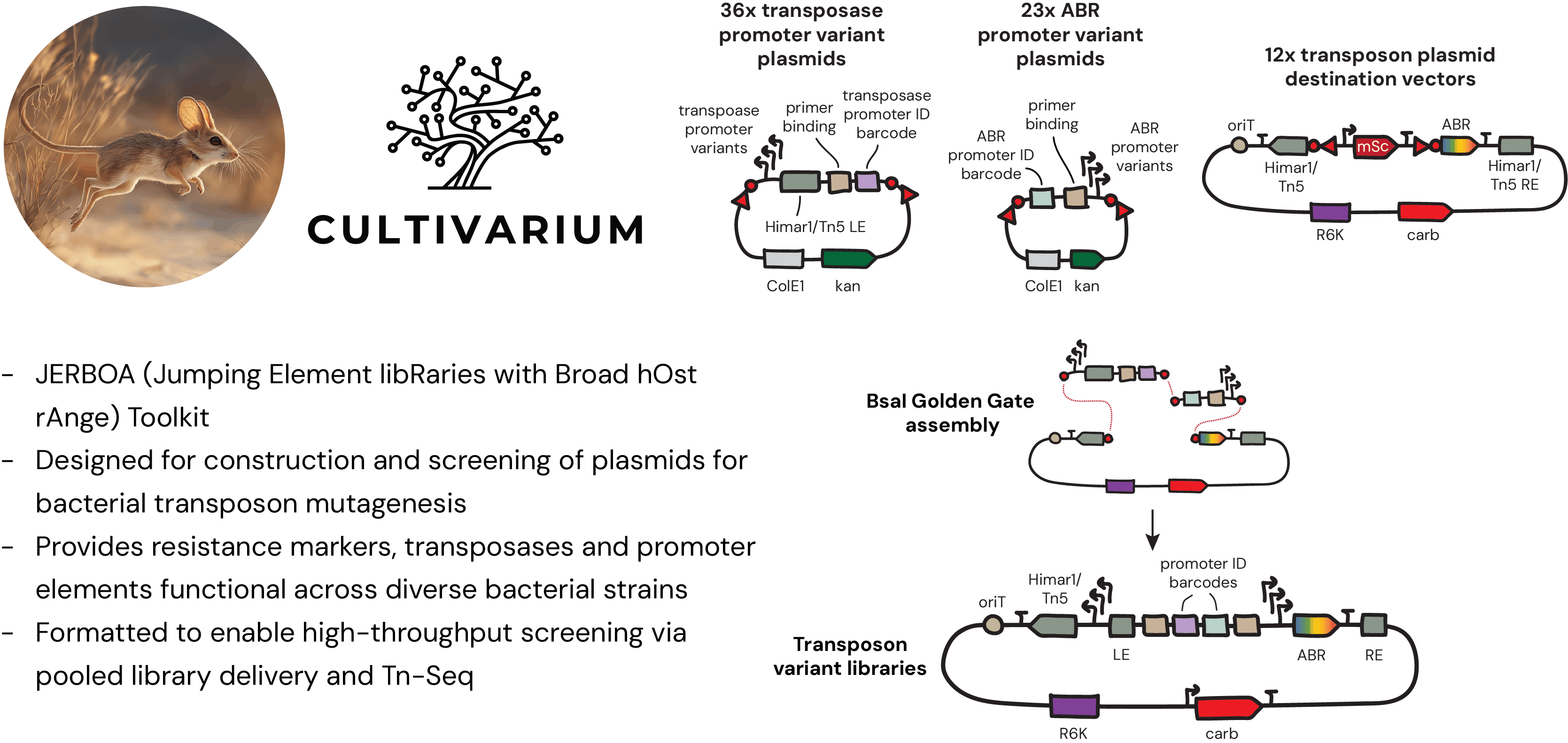
Figure 1: Overview of the JERBOA Toolkit. The toolkit is designed for the modular construction and high-throughput screening of barcoded transposon libraries. The toolkit provides three categories of plasmids: 36 transposase promoter part plasmids, each carrying a bacterial promoter sequence paired with either a Himar1 or Tn5 left mosaic end (LE) and a unique promoter ID barcode; 23 ABR gene promoter part plasmids, each carrying a bacterial promoter sequence and unique ID barcode; and 12 transposon destination vectors featuring either a Himar1 or Tn5 transposase and one of six ABR genes. In-built BsaI restriction enzyme cut sites and overhangs (red triangle and red circle) enable Bsal Golden Gate assembly, and assembly of pooled libraries of up to 3,888 unique transposon variants. Each final construct contains unique barcodes within the transposon for identification via Tn-Seq as well as an RK2/RP4 origin of transfer (oriT) for conjugative delivery, a conditional R6Kγ origin, and a carbenicillin (carb) marker for selection in the E. coli donor.
Kit Documentation
Cultivarium JERBOA Toolkit plasmid sequences (ZIP, 171 KB)How to Cite this Kit
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which they were created, and include Addgene in the Materials and Methods of your future publications.
For your Materials and Methods section:
"The Cultivarium JERBOA Toolkit was a gift from Nili Ostrov, Charlie Gilbert, and Cultivarium Tools (Addgene kit #1000000275)."
For your Reference section:
A scalable transposon mutagenesis system for non-model bacteria Gilbert C, Leung J, Crits-Christoph A, Kang SC, Esmurria A, Fenn K, Brumwell SL, Martin-Moldes Z, Mendler K, Barnum TP, Nguyen M-A, Lee HH, Ostrov N. bioRxiv 2025.12.22.696024. doi: 10.64898/2025.12.22.696024. Article (Link opens in a new window)
Cultivarium JERBOA Toolkit - #1000000275
- Resistance Color Key
Each circle corresponds to a specific antibiotic resistance in the kit plate map wells.
- Inventory
Searchable and sortable table of all plasmids in kit. The Well column lists the plasmid well location in its plate. The Plasmid column links to a plasmid's individual web page.
- Kit Plate Map
96-well plate map for plasmid layout. Hovering over a well reveals the plasmid name, while clicking on a well opens the plasmid page.
Resistance Color Key
| Chloramphenicol | |
| Ampicillin | |
| Kanamycin |
Inventory
| Well | Plasmid | Resistance |
|---|---|---|
| A / 1 | pGLU_63 |
|
| A / 2 | pGLU_64 |
|
| A / 3 | pGLU_66 |
|
| A / 4 | pGLU_67 |
|
| A / 5 | pGLU_68 |
|
| A / 6 | pGLU_69 |
|
| A / 7 | pGLU_71 |
|
| A / 8 | pGLU_72 |
|
| A / 9 | pGLU_73 |
|
| A / 10 | pGLU_74 |
|
| A / 11 | pGLU_75 |
|
| A / 12 | pGLU_76 |
|
| B / 1 | pGLU_77 |
|
| B / 2 | pGLU_79 |
|
| B / 3 | pGLU_80 |
|
| B / 4 | pGLU_81 |
|
| B / 5 | pGLU_82 |
|
| B / 6 | pGLU_103 |
|
| B / 7 | pGLU_104 |
|
| B / 8 | pGLU_105 |
|
| B / 9 | pGLU_107 |
|
| B / 10 | pGLU_108 |
|
| B / 11 | pGLU_109 |
|
| B / 12 | pGLU_110 |
|
| C / 1 | pGLU_111 |
|
| C / 2 | pGLU_112 |
|
| C / 3 | pGLU_113 |
|
| C / 4 | pGLU_115 |
|
| C / 5 | pGLU_116 |
|
| C / 6 | pGLU_117 |
|
| C / 7 | pGLU_118 |
|
| C / 8 | pGLU_119 |
|
| C / 9 | pGLU_85 |
|
| C / 10 | pGLU_86 |
|
| C / 11 | pGLU_87 |
|
| C / 12 | pGLU_89 |
|
| D / 1 | pGLU_90 |
|
| D / 2 | pGLU_91 |
|
| D / 3 | pGLU_92 |
|
| D / 4 | pGLU_93 |
|
| D / 5 | pGLU_94 |
|
| D / 6 | pGLU_95 |
|
| D / 7 | pGLU_97 |
|
| D / 8 | pGLU_98 |
|
| D / 9 | pGLU_99 |
|
| D / 10 | pGLU_100 |
|
| D / 11 | pGLU_101 |
|
| D / 12 | pGLU_7 |
|
| E / 1 | pGLU_8 |
|
| E / 2 | pGLU_9 |
|
| E / 3 | pGLU_10 |
|
| E / 4 | pGLU_11 |
|
| E / 5 | pGLU_12 |
|
| E / 6 | pGLU_128 |
|
| E / 7 | pGLU_129 |
|
| E / 8 | pGLU_130 |
|
| E / 9 | pGLU_131 |
|
| E / 10 | pGLU_132 |
|
| E / 11 | pGLU_133 |
|
| E / 12 | pGLU_62 |
|
| F / 1 | pGLU_65 |
|
| F / 2 | pGLU_83 |
|
| F / 3 | pGLU_120 |
|
| F / 4 | pGLU_70 |
|
| F / 5 | pGLU_78 |
|
| F / 6 | pGLU_84 |
|
| F / 7 | pGLU_106 |
|
| F / 8 | pGLU_114 |
|
| F / 9 | pGLU_88 |
|
| F / 10 | pGLU_96 |
|
| F / 11 | pGLU_102 |
|


