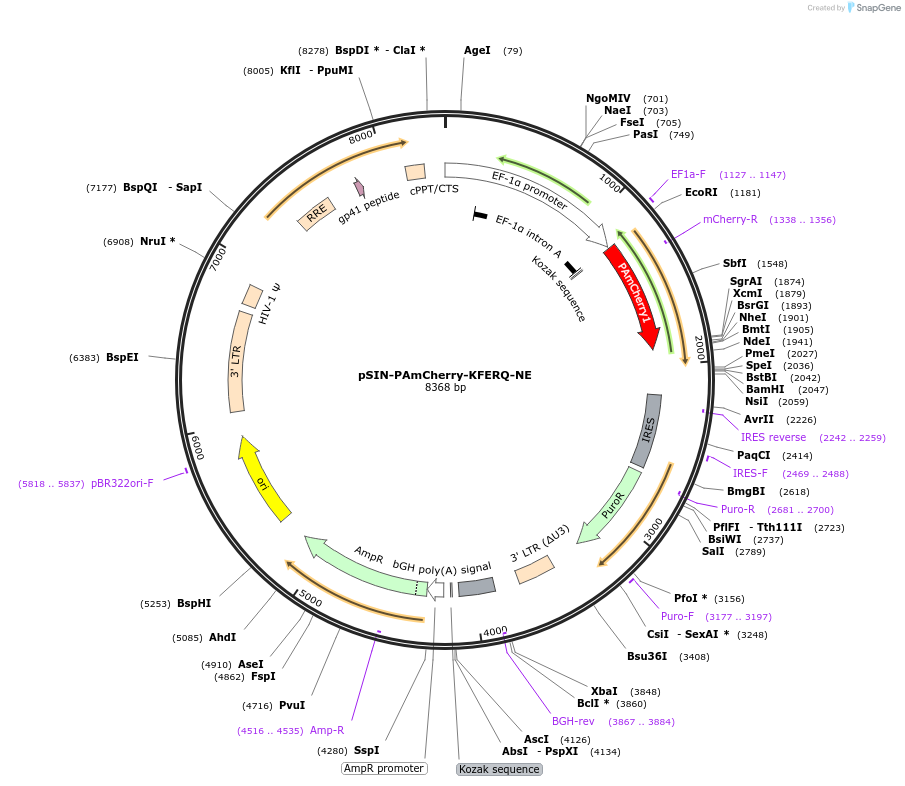
-
PurposeLentiviral overexpression of PA-mCherry-KFERQ fusion
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 102365 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepSin4-EF2-IRES-Pur
-
Backbone manufacturerSINF-EF-G (from Dr. Robert Hawley, George Washington University) was modified to make the lentiviral backbone.
- Backbone size w/o insert (bp) 7500
-
Vector typeMammalian Expression, Lentiviral
-
Selectable markersPuromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Copy numberHigh Copy
Gene/Insert 1
-
Gene/Insert namePA-mCherry
-
SpeciesSynthetic
- Promoter EF-1a
Cloning Information for Gene/Insert 1
- Cloning method Restriction Enzyme
- 5′ cloning site EcoRI (not destroyed)
- 3′ cloning site NheI (not destroyed)
- 5′ sequencing primer EF-1a Forward
- (Common Sequencing Primers)
Gene/Insert 2
-
Gene/Insert nameKFERQ peptide
-
SpeciesSynthetic
-
Insert Size (bp)825
- Promoter EF-1a
-
Tag
/ Fusion Protein
- NE (C terminal on insert)
Cloning Information for Gene/Insert 2
- Cloning method Restriction Enzyme
- 5′ cloning site NheI (not destroyed)
- 3′ cloning site PmeI (not destroyed)
- 5′ sequencing primer unknown
- 3′ sequencing primer TTAGCTTTCGTTATCATCATAGC
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
"KFERQ" is the consensus peptide sequence found in various native proteins for mediating degradation via Chaperone-Mediated Autophagy (CMA)
There is a "NheI" restriction site which we artificially added between PAmCherry and KFERQ-NE to facilitate cloning. i.e. 5'---PAmCherry-NheI-KFERQ-NE----3'
This plasmid encodes a novel 18-amino-acid protein tag - "NE", which can be detected by a specific mouse monoclonal antibody in Western blotting, immunopreciptation, and immunocytochemistry. (Source of antibody: http://www.versitech.hku.hk/reagents/ne/)
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSIN-PAmCherry-KFERQ-NE was a gift from Shu Leong Ho (Addgene plasmid # 102365 ; http://n2t.net/addgene:102365 ; RRID:Addgene_102365) -
For your References section:
Age-dependent accumulation of oligomeric SNCA/alpha-synuclein from impaired degradation in mutant LRRK2 knockin mouse model of Parkinson disease: role for therapeutic activation of chaperone-mediated autophagy (CMA). Ho PW, Leung CT, Liu H, Pang SY, Lam CS, Xian J, Li L, Kung MH, Ramsden DB, Ho SL. Autophagy. 2019 Apr 14:1-24. doi: 10.1080/15548627.2019.1603545. 10.1080/15548627.2019.1603545 PubMed 30983487