-
PurposeBeta-catenin reporter. TCF/LEF sites upstream of a luciferase reporter.
-
Depositing Lab
-
Publication
-
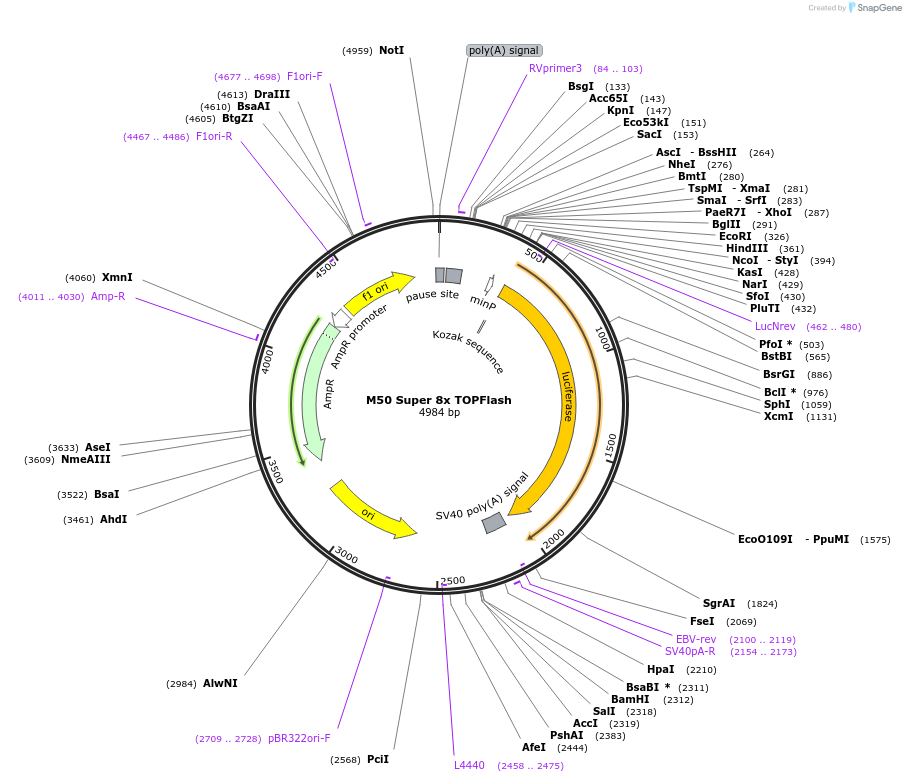
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 12456 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepTA-Luc
-
Backbone manufacturerClontech
- Backbone size w/o insert (bp) 4871
-
Vector typeLuciferase
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameTCF/LEF binding sites
-
Alt namebeta catenin reporter
-
Insert Size (bp)120
-
Entrez GeneCtnnb1 (a.k.a. Bfc, Catnb, Mesc)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site MluI (unknown if destroyed)
- 3′ cloning site MluI (unknown if destroyed)
- 5′ sequencing primer RVPrimer3
- 3′ sequencing primer LucNRev
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe idea for the construct is from Hans Clevers lab, who designed the original TOPflash.
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
This is a luciferase reporter of beta-catenin-mediated transcriptional activation. In HEK cells, maximal activation of this reporter is ~100-fold (activation by Wnt) up to ~1,000-fold (activation by phosphorylation mutants of beta-catenin). The appropriate control plasmid is clone M51, Super8XFOPflash, which has mutant TCF/LEF binding sites.
This construct was made by Ajamete Kaykas in the Moon lab. The backbone is the pTA-Luc vector of Clontech, which provides a minimal TA viral promoter driving expression of the firefly luciferase gene (see company publications for details). 7 TCF/LEF binding sites were cloned into the Mlu1 site of this vector (7 copies of: AGATCAAAGGgggta, with TCF/LEF binding site in CAP letters, and a spacer in lower case, separating each copy of the TCF/LEF site).
Note: This plasmid was published as M50 Super 8x TOPFlash, but the plasmid actually contains 7 TCF/LEF sites.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
M50 Super 8x TOPFlash was a gift from Randall Moon (Addgene plasmid # 12456 ; http://n2t.net/addgene:12456 ; RRID:Addgene_12456) -
For your References section:
Zebrafish prickle, a modulator of noncanonical Wnt/Fz signaling, regulates gastrulation movements. Veeman MT, Slusarski DC, Kaykas A, Louie SH, Moon RT. Curr Biol. 2003 Apr 15. 13(8):680-5. 10.1016/S0960-9822(03)00240-9 PubMed 12699626