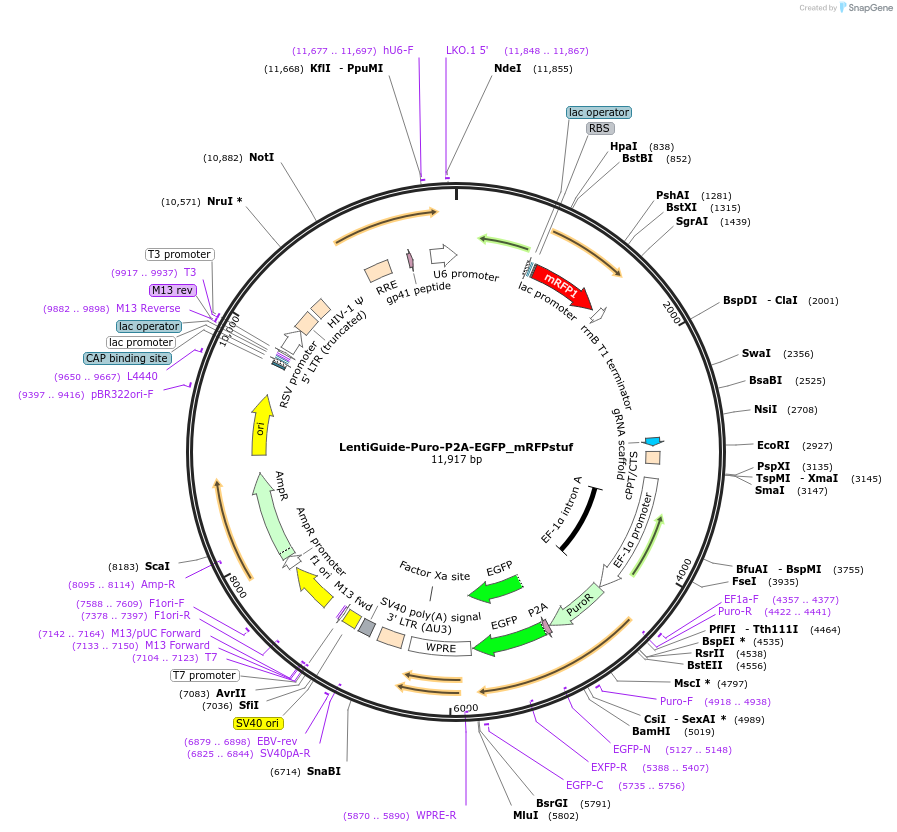
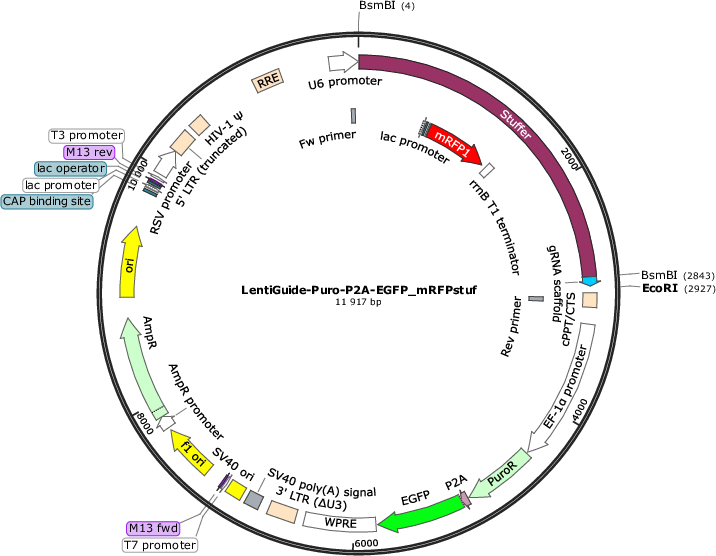
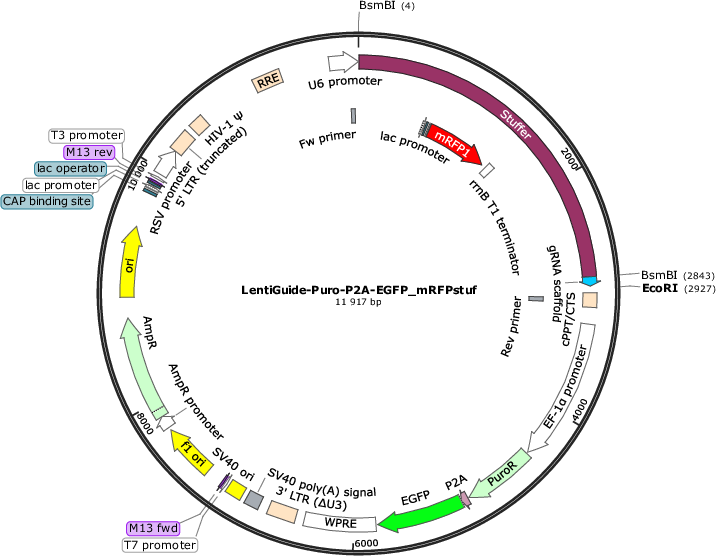
LentiGuide-Puro-P2A-EGFP_mRFPstuf
(Plasmid
#137730)
-
Purpose(Empty Backbone) Expresses customizable S. Pyogenes sgRNA from U6 promoter and PuroR-P2A-EGFP from EF-1a promoter. Stuffer contains mRFP from a bacterial promoter, enabling simple visual quality control step of sgRNA
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 137730 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonelentiGuide-Puro-P2A-EGFP
-
Backbone manufacturerFeng Zhang Lab
- Backbone size (bp) 10963
-
Modifications to backboneA monomeric red fluorescent protein (mRFP) sequence driven by the lac promoter was cloned into the stuffer region of the lentiGuide-Puro-P2A-EGFP vector (#137729). Furthermore, a unique EcoRI site was introduced following the sgRNA scaffold.
-
Vector typeMammalian Expression, Bacterial Expression, Mouse Targeting, Lentiviral, CRISPR, Synthetic Biology
-
Selectable markersPuromycin
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Growth instructionsBest to use 32°C for sgRNA library cloning into the linearized vector
-
Copy numberUnknown
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer AATGGACTATCATATGCTTACCGTAACTTGAAAGTATTTCG
- 3′ sequencing primer CTTTAGTTTGTATGTCTGTTGCTATTATGTCTACTATTCTTTCC
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe original lentiGuide-Puro plasmid (#52963) was obtained from Addgene.
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
LentiGuide-Puro-P2A-GFP_mRFPstuf was generated from the lentiGuide-Puro-P2A-GFP (#137729), which was generated from lentiGuide-Puro (#52963).
The plasmid has mRFP driven by the bacterial lac promotor in the stuffer (the mRFP cassette was a kind gift from Dr. Erik Holmgren). The stuffer, 2839 bp, is excised by BsmBI digestion and replaced by a CRISPR library (or individual gRNA spacers). If the BsmBI digestion and the subsequent cloning is inadequate, the resulting CRISPR library can be contaminated by plasmids containing the stuffer instead of gRNAs. The bacterial mRFP cassette in the stuffer enables rapid identification of bacterial colonies carrying plasmids with retained stuffer (bacteria are red) and colonies without the stuffer (bacteria are white). This can be used as a simple visual quality control step for the gRNA cloning efficiency, as well as guide the selection of bacterial colonies where the stuffer has been excised.
A unique EcoRI site was introduced directly following the sgRNA scaffold, facilitating cloning in this region (introducing UMIs, introducing mutations in the sgRNA scaffold, etc).
The plasmid contains both PuroR and EGFP, enabling the selection of successfully infected cells based on both these selection markers. We typically add a pre-determined dose of puromycin (~5-15ug/ml) 24h after infection, using non-infected cells as controls for the efficiency of the selection. After 24-48h selection all non-infected control cells are typically dead, and the infected cells are ~100% EGFP positive. The EGFP signal can also be used for the simple determination of MOI of the viral preparation.
The provided sequencing primers are the primers traditionally used for NGS sequencing of gRNA representation in lentiGuide-Puro (#52963). This primer pair can also be used for Sanger sequencing the insert, amplifying the DNA sample (60oC annealing temp), and sequencing the ~300bp amplicon using the reversed primer. The forward primer is too close to the gRNA sequence for Sanger sequence. 5’-TGTAAACACAAAGATATTAGTACAAAATACGTGAC-3´ can be used as an alternative forward primer, resulting in a larger amplicon, better suited for Sanger sequencing.
LentiGuide-Puro-P2A-GFP_mRFPstuf vector was generated from the lentiGuide-Puro-P2A-GFP vector (#137729). This construct has RFP driven by a bacterial promotor in the stuffer (a kind gift from Dr. Erik Holmgren).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
LentiGuide-Puro-P2A-EGFP_mRFPstuf was a gift from Fredrik Wermeling (Addgene plasmid # 137730 ; http://n2t.net/addgene:137730 ; RRID:Addgene_137730) -
For your References section:
IL-4 controls activated neutrophil FcgammaR2b expression and migration into inflamed joints. Panda SK, Wigerblad G, Jiang L, Jimenez-Andrade Y, Iyer VS, Shen Y, Boddul SV, Guerreiro-Cacais AO, Raposo B, Kasza Z, Wermeling F. Proc Natl Acad Sci U S A. 2020 Jan 24. pii: 1914186117. doi: 10.1073/pnas.1914186117. 10.1073/pnas.1914186117 PubMed 31980518