-
Depositing Labs
-
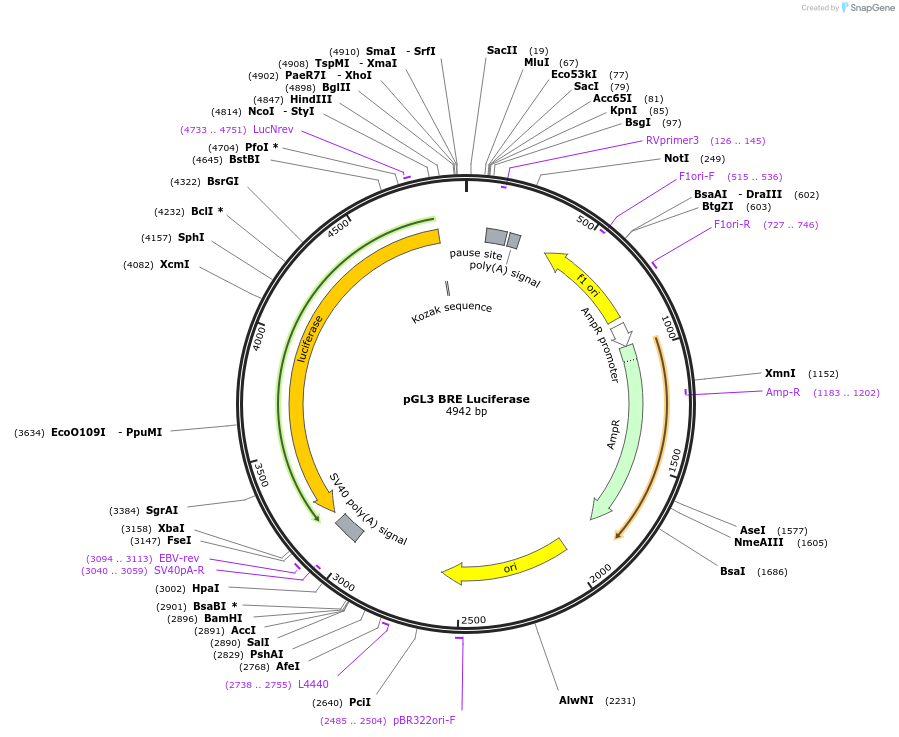
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 45126 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepGL3-Basic
-
Backbone manufacturerPromega
- Backbone size w/o insert (bp) 4818
-
Vector typeMammalian Expression, Luciferase ; BMP/Smad Transcriptional Reporter
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameId1-BRE
-
Alt nameBRE-Luc reporter
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site NheI (not destroyed)
- 3′ cloning site NheI (not destroyed)
- 5′ sequencing primer RVprimer3
- 3′ sequencing primer LucNRev
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Synthetic oligonucleotides containing two CAGC sites, GGCGCC palindrome and MluI 5′-overhang (on the sense strand) and two SBE sites, one GGCG site and NheI 5′-overhang (on the antisense strand) were ligated and inserted into NheI sites in pGL3-MLP-luc minimal promoter vector. Thus the construct was created with a 5′ site for NheI site followed by 2 SBEs, 1 GGCG site, 2 CAGC′ sites, 2 GGCGCC palindromes, 2 CAGC′ sites, 1 GGCG site, 2 SBEs, and an NheI 3′ site.
Based on the above description and the associated publication, the deduced sequence of the full hairpin is:
GCTAGCTCAGACCGTTAGACGCCAGGACGGGCTGTCAGGCTGGCGCCG CGGCGCCAGCCTGACAGCCCGTCCTGGCGTCTAACGGTCTGAGCTAGC
The presence of the hairpin can make sequencing this region difficult by traditional Sanger Sequencing. Protocols for "difficult templates" or "GC rich" may help.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pGL3 BRE Luciferase was a gift from Martine Roussel & Peter ten Dijke (Addgene plasmid # 45126 ; http://n2t.net/addgene:45126 ; RRID:Addgene_45126) -
For your References section:
Identification and functional characterization of distinct critically important bone morphogenetic protein-specific response elements in the Id1 promoter. Korchynskyi O, ten Dijke P. J Biol Chem. 2002 Feb 15;277(7):4883-91. Epub 2001 Nov 29. 10.1074/jbc.M111023200 PubMed 11729207