-
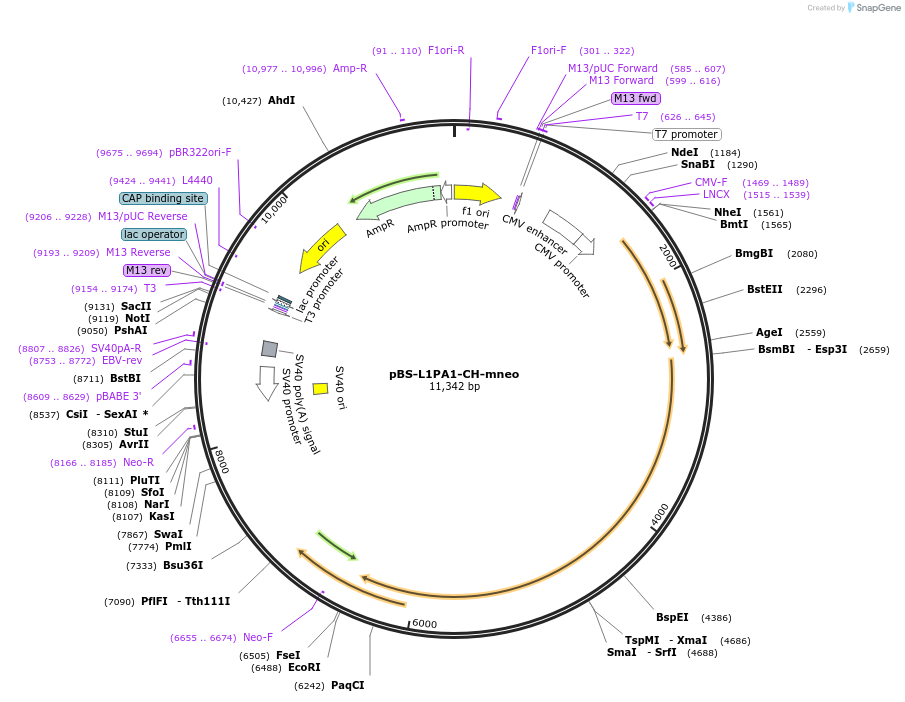
Purposecontains the fully codon optimized ORF1 and ORF2 of the human L1RP
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 51288 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepBluescript II
-
Backbone manufacturerStratagene
-
Vector typeMammalian Expression
-
Selectable markersNeomycin (select with G418)
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameL1
-
Alt nameL1RE1
-
Alt nameLine1
-
SpeciesH. sapiens (human)
-
Entrez GeneL1RE1 (a.k.a. L1.2, LRE1)
- Promoter CMV
-
Tag
/ Fusion Protein
- mneoI indicator cassette containing an inverted neomycin resistance gene (C terminal on insert)
Cloning Information
- Cloning method Unknown
- 5′ sequencing primer T7
- 3′ sequencing primer SP6
- (Common Sequencing Primers)
Resource Information
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
L1 is tagged with neoI indicator cassette containing an inverted neomycin resistance gene. The SV40 promoter (SV40p) drives transcription of the neomycin gene which is disrupted by an intron in the opposite orientation that renders it non-functional. The intron is only spliced out from tagged L1 transcripts generated by the CMV promoter. When retrotransposition of the spliced L1 RNA occurs, the new insert will contain a functional neo gene that is expressed from the SV40 promoter.
The L1 open reading frames were codon optimized using Primo Optimum 3.4 (http://www.changbioscience.com/primo/primoo.html) which makes synonymous changes to optimal codons within the nucleotide sequence. A 63 bp inter-ORF with 92% sequence identity to L1RP was utilized for all constructs. Sequence changes to this region were generated to introduce restriction enzyme sites used for plasmid construction. The optimized nucleotide sequence is 74.9% identical to the sequence of L1RP.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pBS-L1PA1-CH-mneo was a gift from Astrid Roy-Engel (Addgene plasmid # 51288 ; http://n2t.net/addgene:51288 ; RRID:Addgene_51288) -
For your References section:
Evolutionary conservation of the functional modularity of primate and murine LINE-1 elements. Wagstaff BJ, Barnerssoi M, Roy-Engel AM. PLoS One. 2011 May 10;6(5):e19672. doi: 10.1371/journal.pone.0019672. 10.1371/journal.pone.0019672 PubMed 21572950