-
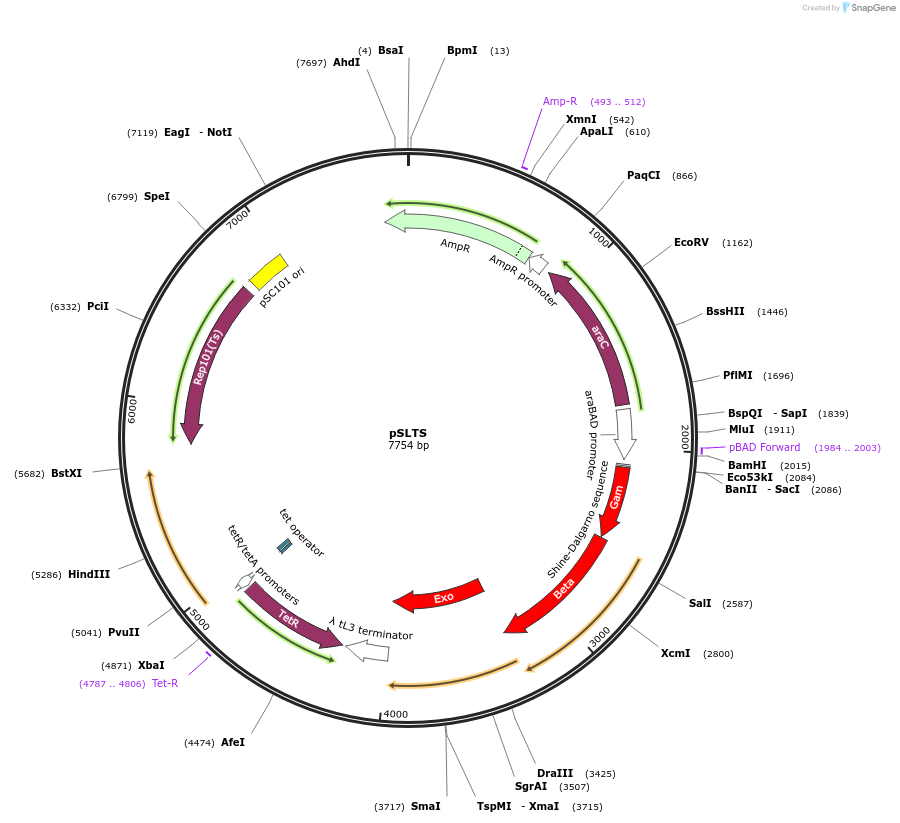
Purpose(Empty Backbone) Expresses the lambda Red recombinase under control of the arabinose-inducible P(araB) promoter and I-SceI under control of the anhydrotetracycline-inducible P(tetA) promoter.
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 59386 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepKDTS
-
Backbone manufacturerHerring and Blattner
- Backbone size (bp) 7742
-
Modifications to backboneWe constructed pSLTS by correcting three mutations found in pKDTS. One of these mutations resulted in replacement of 5 amino acid residues at the N-terminus of I-SceI with a single amino acid (MHMKNIK to MHQ), and the other two resulted in missense changes of ASN11 to Ser and Lys33 to Arg in TetR (AAC to AGC and AAG to AGG, respectively). Please see GenBank: KP897155.1 for more details.
-
Vector typeBacterial Expression ; Genome Engineering
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature30°C
-
Growth Strain(s)Mach1
-
Growth instructionspSLTS has a temperature sensitive origin of replication and will be lost when cells are grown at temperatures higher than 30 ℃.
-
Copy numberLow Copy
Cloning Information
- Cloning method Gibson Cloning
- 5′ sequencing primer TAGGCGCAATCACTTTCGTCTACTC
- 3′ sequencing primer TTGAGTGACATGCAAAGTAAGTATGATCTC
- (Common Sequencing Primers)
Resource Information
-
Addgene Notes
-
A portion of this plasmid was derived from a plasmid made bypSLTS was constructed from pKDTS, which was originally constructed by C. D. Herring and F. R. Blattner (J. Bacteriology 186: 2673(2004)).
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
We developed GetX (https://sourceforge.net/projects/getx/) a stand-alone python script that allows the user to design mutation cassettes for scarless genome editing in bacteria using our previously described two-step recombination method. Please see the document linked under the Resource Information heading above for additional information on installing and using the script.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pSLTS was a gift from Shelley Copley (Addgene plasmid # 59386 ; http://n2t.net/addgene:59386 ; RRID:Addgene_59386) -
For your References section:
A versatile and highly efficient method for scarless genome editing in Escherichia coli and Salmonella enterica. Kim J, Webb AM, Kershner JP, Blaskowski S, Copley SD. BMC Biotechnol. 2014 Sep 25;14(1):84. 10.1186/1472-6750-14-84 PubMed 25255806