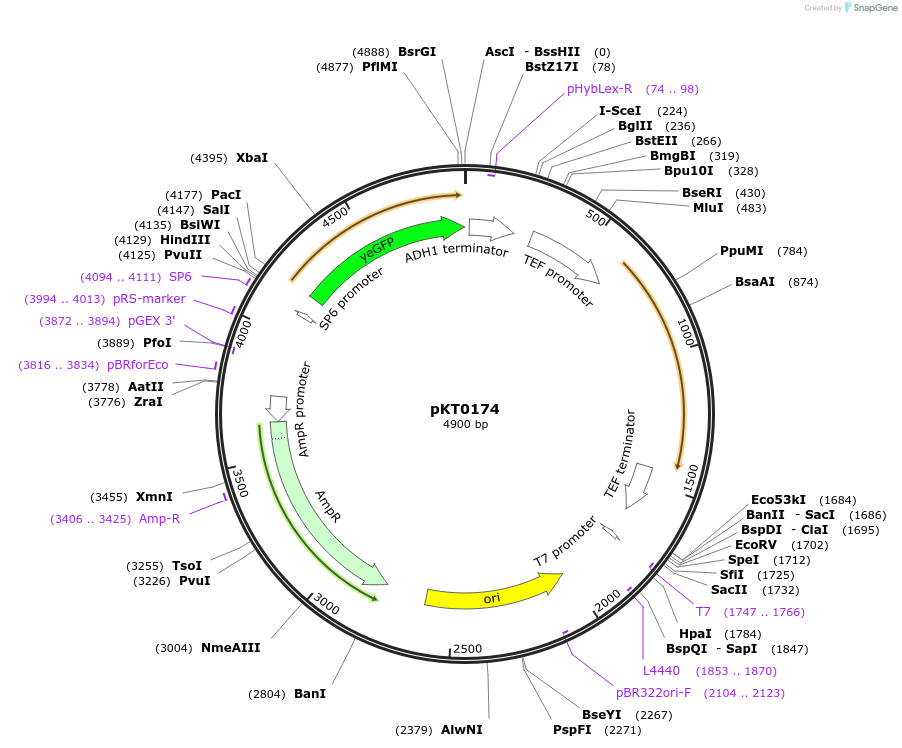
pKT0174
(Plasmid
#8720)
-
Depositing Lab
-
Publication
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 8720 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepFA6a-link
- Backbone size w/o insert (bp) 4900
-
Vector typeYeast Expression
-
Selectable markersURA3 ; Candida albicans
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberUnknown
Gene/Insert
-
Gene/Insert nameyECFP
-
Alt nameCodon-optimized GFP variant
-
Alt namepKT174 (pFA6a–link–yECFP–CaURA3)
-
MutationyEGFP with F64L, S65T, Y66W, N144I, M153T, V163A
-
Tag
/ Fusion Protein
- Codon optimized linker (N terminal on backbone)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site PacI (not destroyed)
- 3′ cloning site AscI (not destroyed)
- 5′ sequencing primer SP6
- (Common Sequencing Primers)
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Full name of this plasmid is pFA6a-link-yECFP-CaURA3. yECFP is created by mutagenesis from a codon-optimized green fluorescent protein - yEGFP1 (Cormack et al. 1997). This yEGFP variant is cloned into pDH5, replacing YFP. An codon-optimized linker was added into the yEGFP variant to improve expression level. C. albicans URA3 (CaURA3) was introduced by subcloning the BglII/SacI fragment of pAG60 (Goldstein et al., 1999) into the plasmid, in place of SpHIS5.
Note that yECFP in this plasmid contains N144I mutation relative to yEGFP. The associated publication mentions a N146I mutation, but N146 is present in the yECFP in this plasmid. The yeast GFP used to create this plasmid contained a point mutation M233I that is present in all GFP variants derived from this original yEGFP. The mutation had no effect on fluorescence (see associated publication for more details).
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pKT0174 was a gift from Kurt Thorn (Addgene plasmid # 8720 ; http://n2t.net/addgene:8720 ; RRID:Addgene_8720) -
For your References section:
Optimized cassettes for fluorescent protein tagging in Saccharomyces cerevisiae. Sheff MA, Thorn KS. Yeast 2004 Jun;21(8):661-70. 10.1002/yea.1130 PubMed 15197731