We narrowed to 20,670 results for: ARIA
-
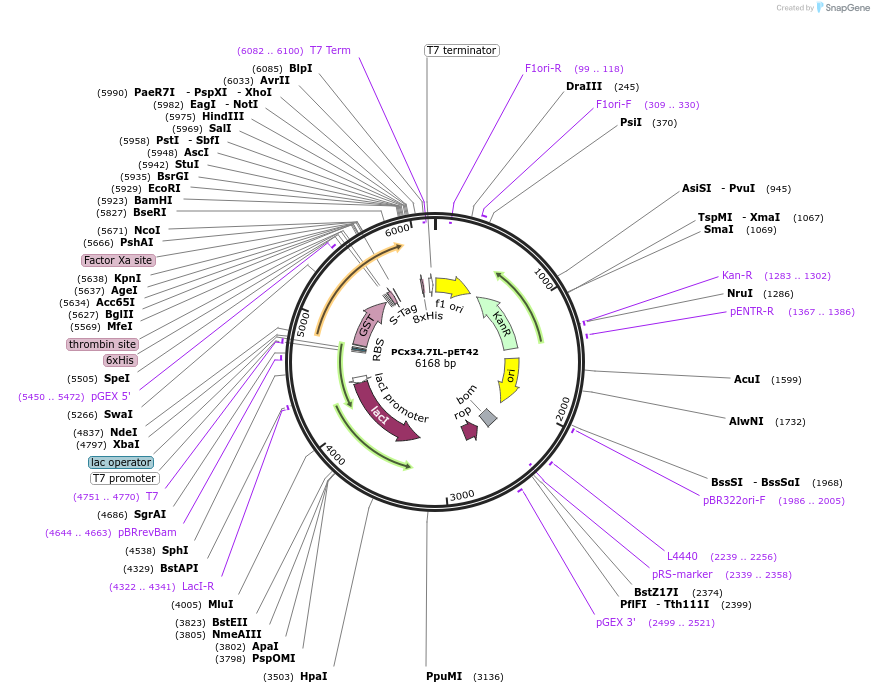
Plasmid#42915DepositorInsertConnexin 34.7 cytoplasmic loop
Tags6x His, GST, and S-tagExpressionBacterialMutationcytoplasmic loop is aa 102-179PromoterT7Available SinceMarch 28, 2013AvailabilityAcademic Institutions and Nonprofits only -
PL06010B1E06 (smedwi3)
Plasmid#35697DepositorInsertSmedwi3
PromoterT7 sense, T3 antisenseAvailable SinceApril 2, 2012AvailabilityAcademic Institutions and Nonprofits only -
pASUS17R
Plasmid#1205DepositorAvailable SinceAug. 19, 2005AvailabilityAcademic Institutions and Nonprofits only -
SARS-CoV-2 Spike (S) Receptor Binding Domain (RBD) library N501Y Tile 2 (positions 437-527)
Pooled Library#174296PurposeThis library contains mutations to SARS-CoV-2 Spike RBD N501Y variant in which all surface exposed residues on S RBD (Tile 2, positions 437-527) were mutated to every other amino acid.DepositorExpressionYeastAvailable SinceSept. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
SARS-CoV-2 Spike (S) Receptor Binding Domain (RBD) library N501Y Tile 1 (positions 333-436)
Pooled Library#174295PurposeThis library contains mutations to SARS-CoV-2 Spike RBD N501Y variant in which all surface exposed residues on S RBD (Tile 1, positions 333-436) were mutated to every other amino acid.DepositorExpressionYeastAvailable SinceSept. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
SARS-CoV-2 Spike (S) Receptor Binding Domain (RBD) library E484K Tile 2 (positions 437-527)
Pooled Library#174294PurposeThis library contains mutations to SARS-CoV-2 Spike RBD E484K variant in which all surface exposed residues on S RBD (Tile 2, positions 437-527) were mutated to every other amino acid.DepositorExpressionYeastAvailable SinceSept. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
SARS-CoV-2 Spike (S) Receptor Binding Domain (RBD) library E484K Tile 1 (positions 333-436)
Pooled Library#174293PurposeThis library contains mutations to SARS-CoV-2 Spike RBD E484K variant in which all surface exposed residues on S RBD (Tile 1, positions 333-436) were mutated to every other amino acid.DepositorExpressionYeastAvailable SinceSept. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
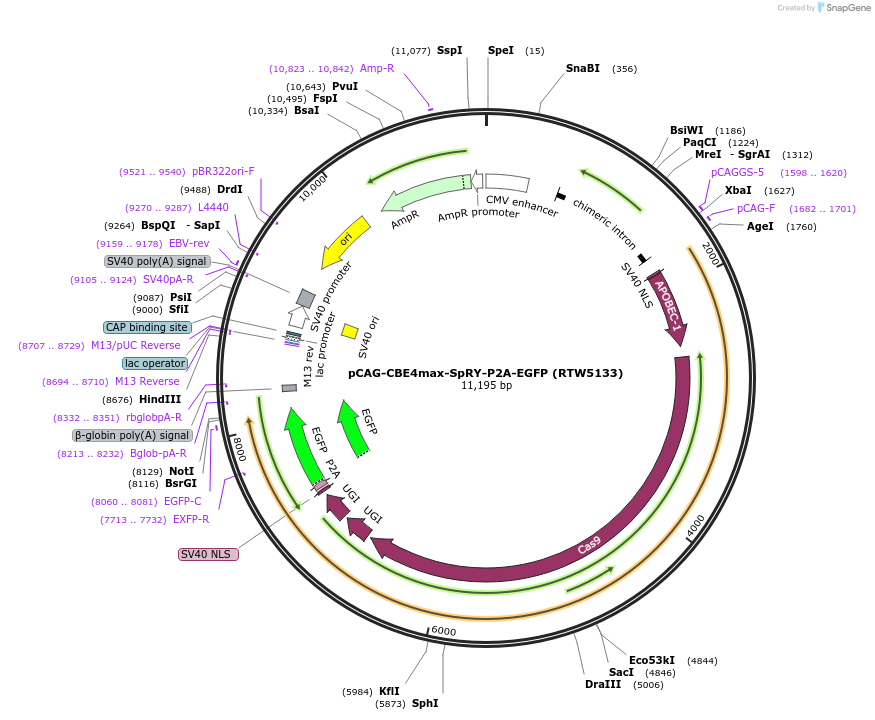
pCAG-CBE4max-SpRY-P2A-EGFP (RTW5133)
Plasmid#139999PurposeCAG promoter expression plasmid for human codon optimized BE4max C-to-T base editor with SpRY(D10A/A61R/L1111R/D1135L/S1136W/G1218K/E1219Q/N1317R/A1322R/R1333P/R1335Q/T1337R) and P2A-EGFPDepositorInserthuman codon optimized CBE4max SpCas9 variant named SpRY with P2A-EGFP
UseCRISPRTagsP2A-EGFPExpressionMammalianMutationnSpCas9=D10A; SpRY=A61R/L1111R/D1135L/S1136W/G121…PromoterCAGAvailable SinceMarch 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
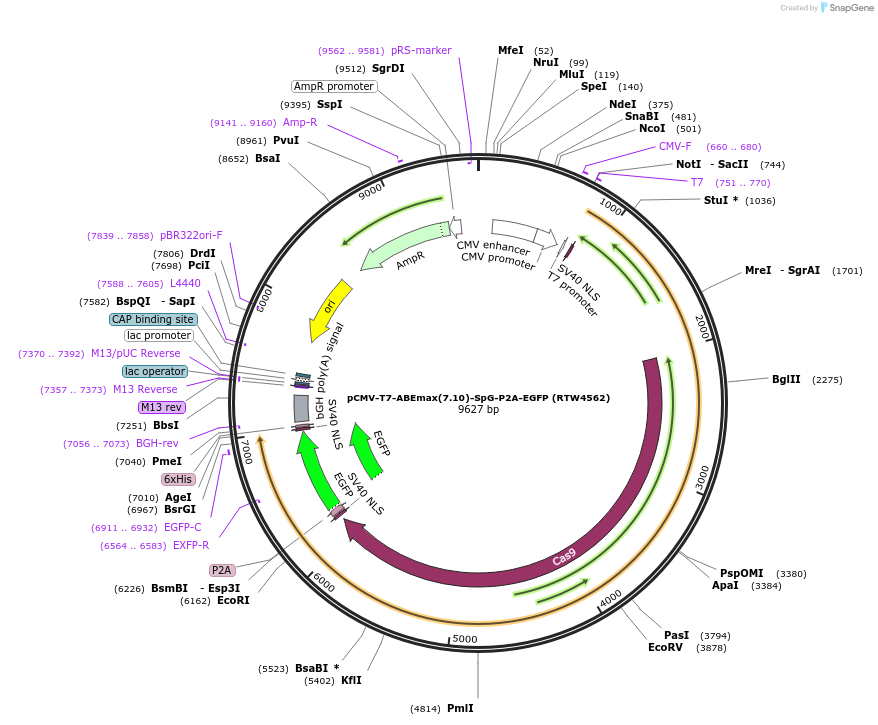
pCMV-T7-ABEmax(7.10)-SpG-P2A-EGFP (RTW4562)
Plasmid#140002PurposeCMV and T7 promoter expression plasmid for human codon optimized ABEmax(7.10) A-to-G base editor with SpG(D10A/D1135L/S1136W/G1218K/E1219Q/R1335Q/T1337R) and P2A-EGFPDepositorInserthuman codon optimized ABEmax(7.10) SpCas9 variant named SpG with P2A-EGFP
UseCRISPR; In vitro transcription; t7 promoterTagsP2A-EGFPExpressionMammalianMutationnSpCas9=D10A; SpG=D1135L/S1136W/G1218K/E1219Q/R13…PromoterCMV and T7Available SinceMarch 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
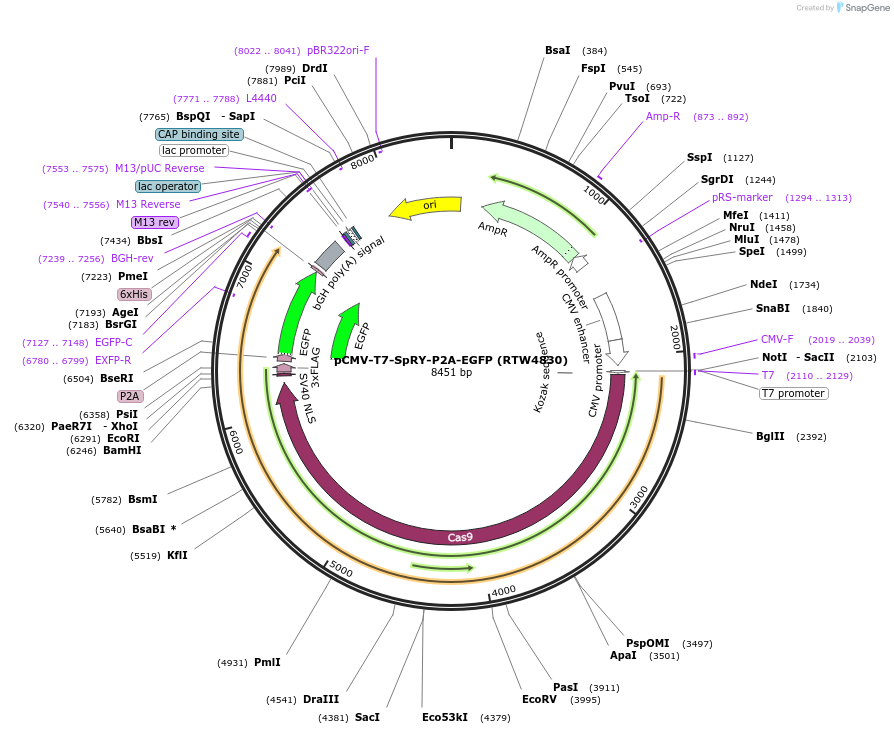
pCMV-T7-SpRY-P2A-EGFP (RTW4830)
Plasmid#139989PurposeCMV and T7 promoter expression plasmid for human codon optimized SpRY(A61R/L1111R/D1135L/S1136W/G1218K/E1219Q/N1317R/A1322R/R1333P/R1335Q/T1337R) with a c-terminal bi-partite NLS, 3x flag tag, and P2A-EGFPDepositorInserthuman codon optimized SpCas9 variant named SpRY with BPNLS-3xFLAG-P2A-EGFP
UseCRISPR; In vitro transcription; t7 promoterTagsBPNLS-3xFLAG-P2A-EGFPExpressionMammalianMutationSpRY=A61R/L1111R/D1135L/S1136W/G1218K/E1219Q/N131…PromoterCMV and T7Available SinceMarch 26, 2020AvailabilityAcademic Institutions and Nonprofits only -
-
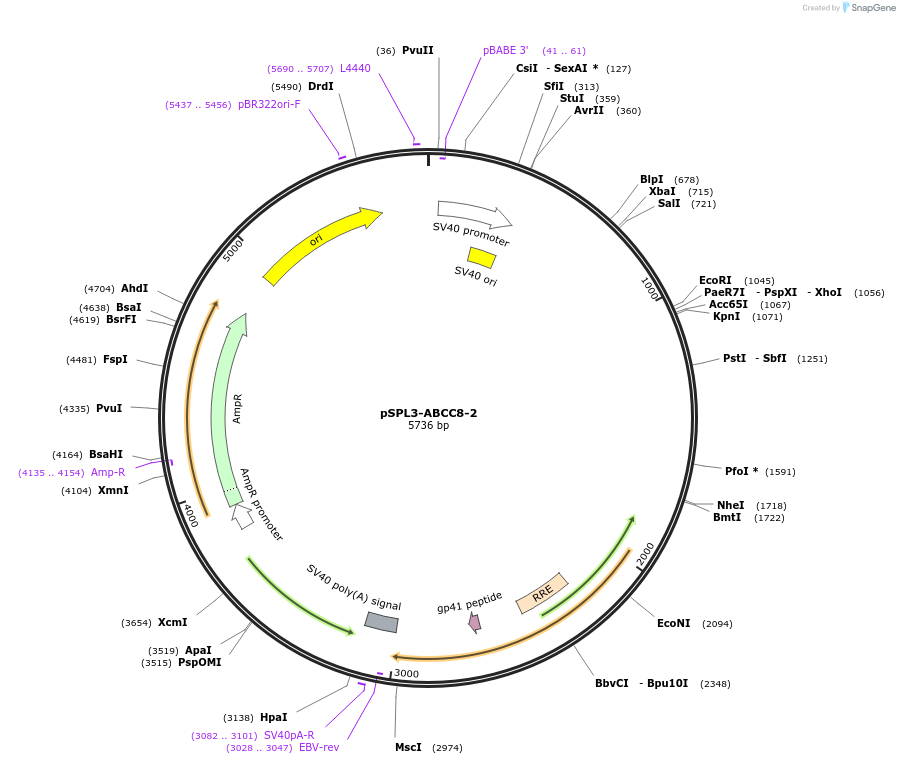
pSPL3-ABCC8-2
Plasmid#135916PurposeEncodes ABCC8 exon 2 wild type for the analysis of splicing variantsDepositorInsertATP-binding Cassette subfamily C member 8 (ABCC8 Human)
UseExon trappingExpressionMammalianPromoterSV40Available SinceMarch 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
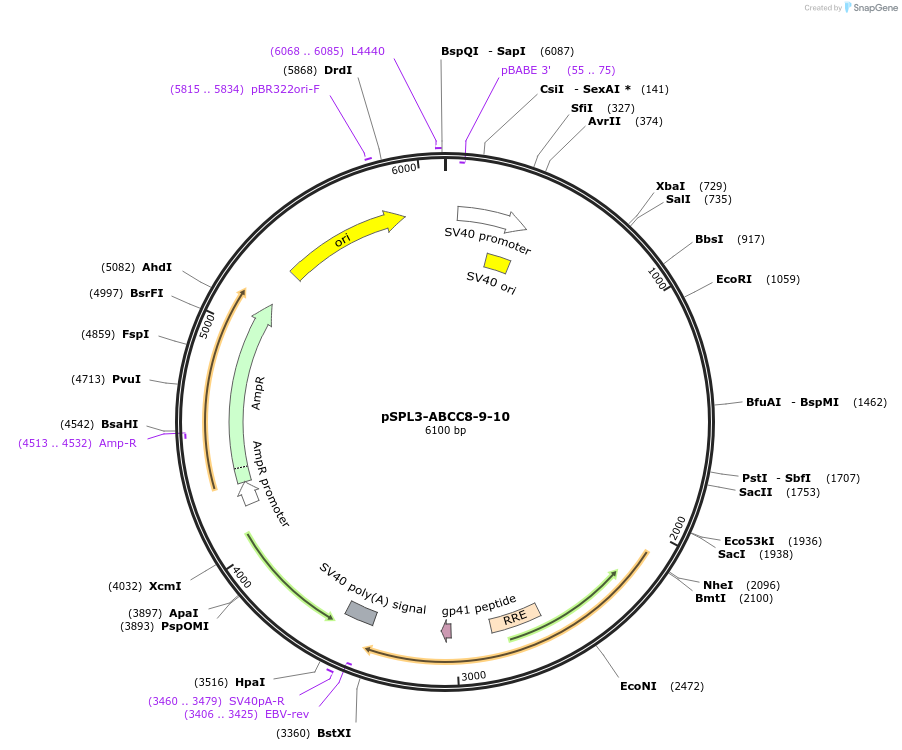
pSPL3-ABCC8-9-10
Plasmid#135918PurposeEncodes ABCC8 exons 9 and 10 wild type for the analysis of splicing variantsDepositorInsertATP-binding Cassette subfamily C member 8 (ABCC8 Human)
UseExon trappingExpressionMammalianPromoterSV40Available SinceMarch 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
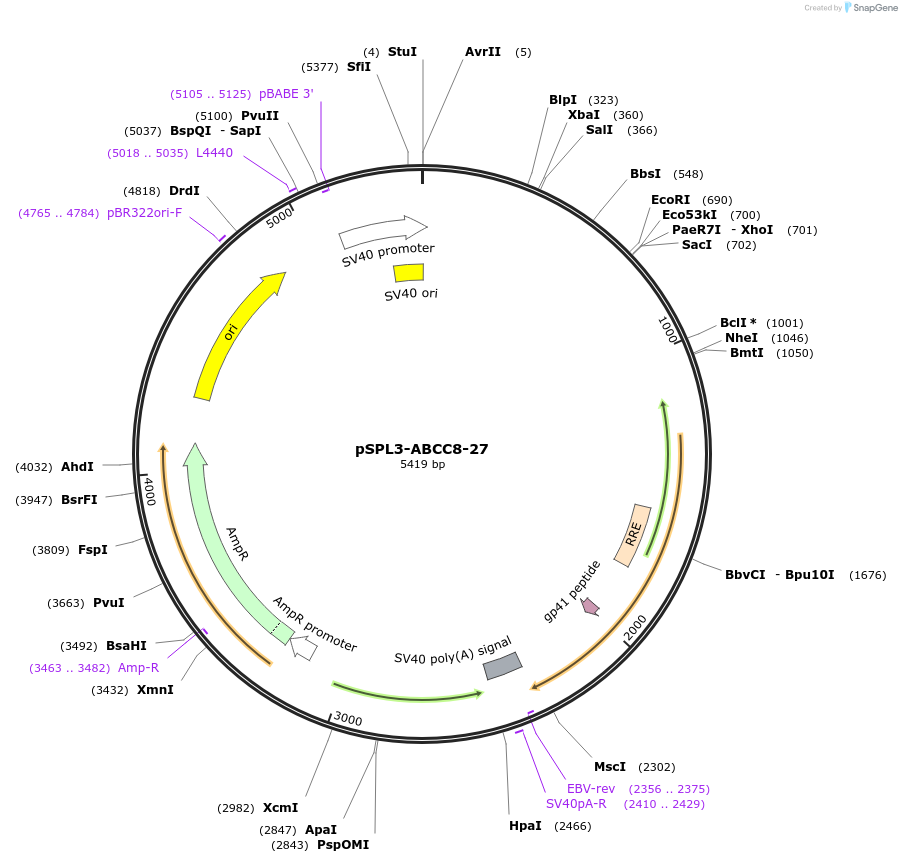
pSPL3-ABCC8-27
Plasmid#135919PurposeEncodes ABCC8 exon 27 wild type for the analysis of splicing variantsDepositorInsertATP-binding Cassette subfamily C member 8 (ABCC8 Human)
UseExon trappingExpressionMammalianPromoterSV40Available SinceMarch 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
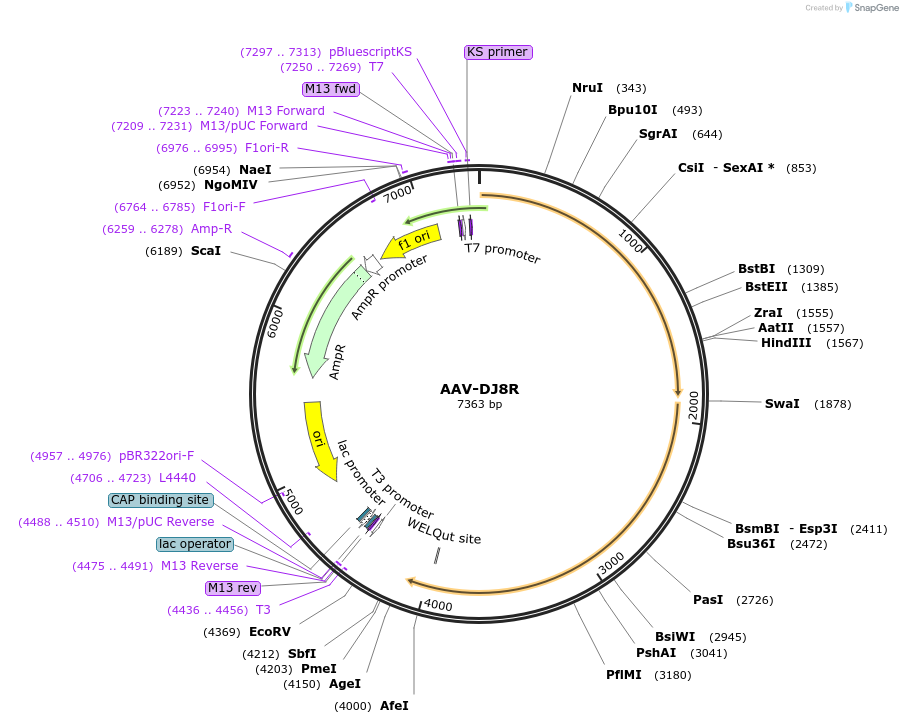
AAV-DJ8R
Plasmid#213974PurposeAAV packaging vector containing AAV2-rep gene and CapDJR modified AAV-DJ8 capsid coding sequenceDepositorTypeEmpty backboneUseAAVAvailable SinceMarch 7, 2024AvailabilityAcademic Institutions and Nonprofits only -
-
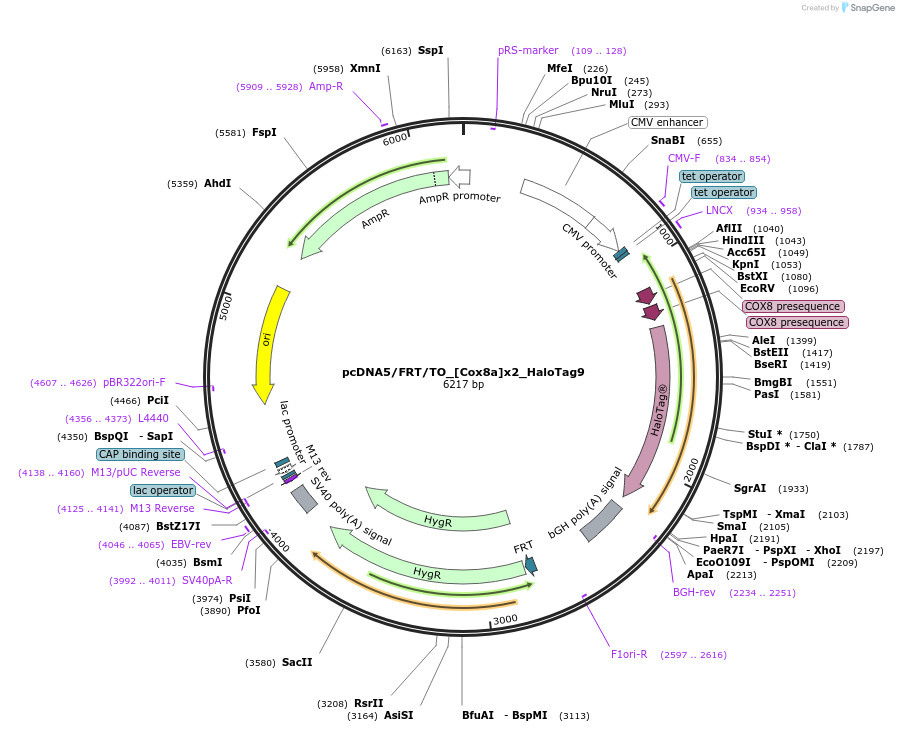
pcDNA5/FRT/TO_[Cox8a]x2_HaloTag9
Plasmid#175539PurposeExpression of HaloTag9 at the inner membrane of mitochondria in mammalian cellsDepositorInsertCox8a-HaloTag9
ExpressionMammalianMutationHaloTag9 = HaloTag7-Q165H-P174RAvailable SinceDec. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
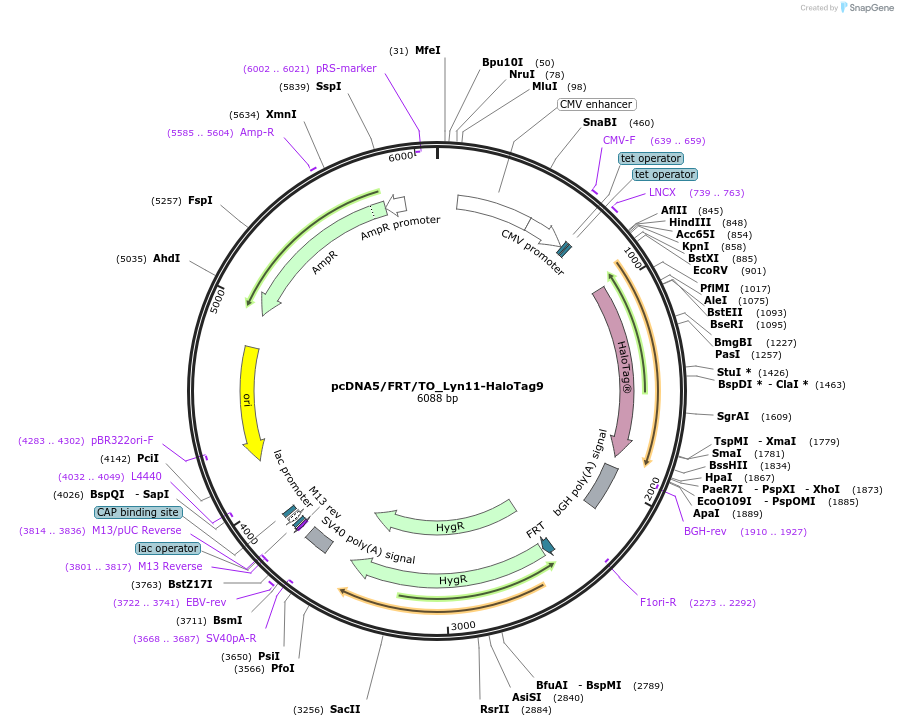
pcDNA5/FRT/TO_Lyn11-HaloTag9
Plasmid#175543PurposeExpression of HaloTag9 at the inner leaflet of the plasma membrane of mammalian cellsDepositorInsertLyn11-HaloTag9
ExpressionMammalianMutationHaloTag9 = HaloTag7-Q165H-P174RAvailable SinceDec. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
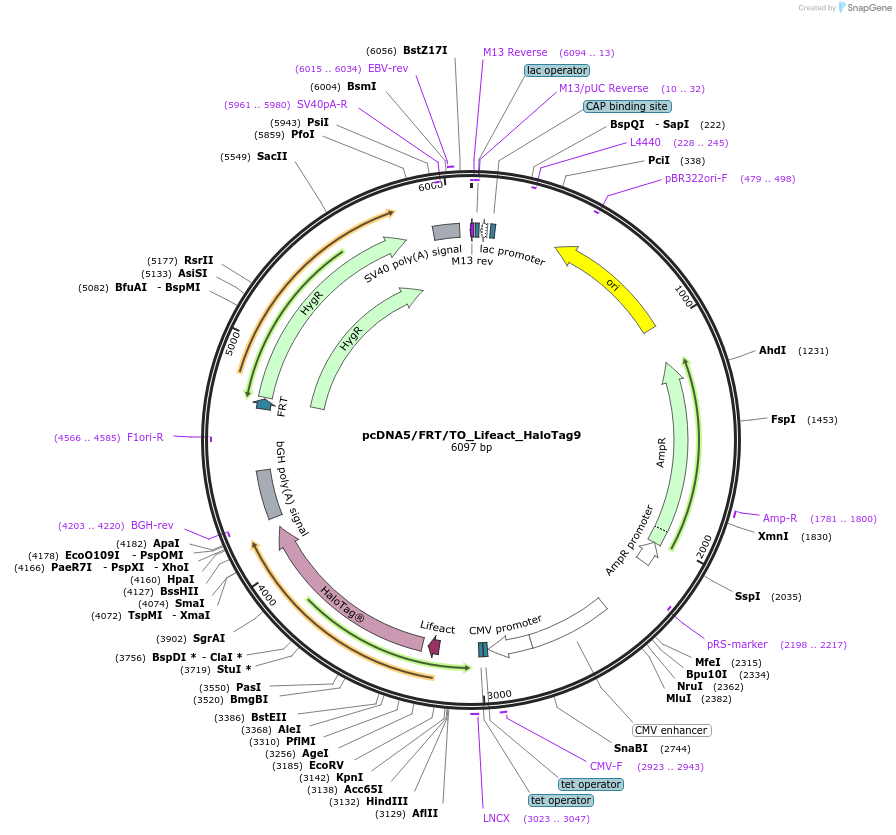
pcDNA5/FRT/TO_Lifeact_HaloTag9
Plasmid#175545PurposeExpression of HaloTag9 at the actin filaments of mammalian cellsDepositorInsertLifeact-HaloTag9
ExpressionMammalianMutationHaloTag9 = HaloTag7-Q165H-P174RAvailable SinceDec. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
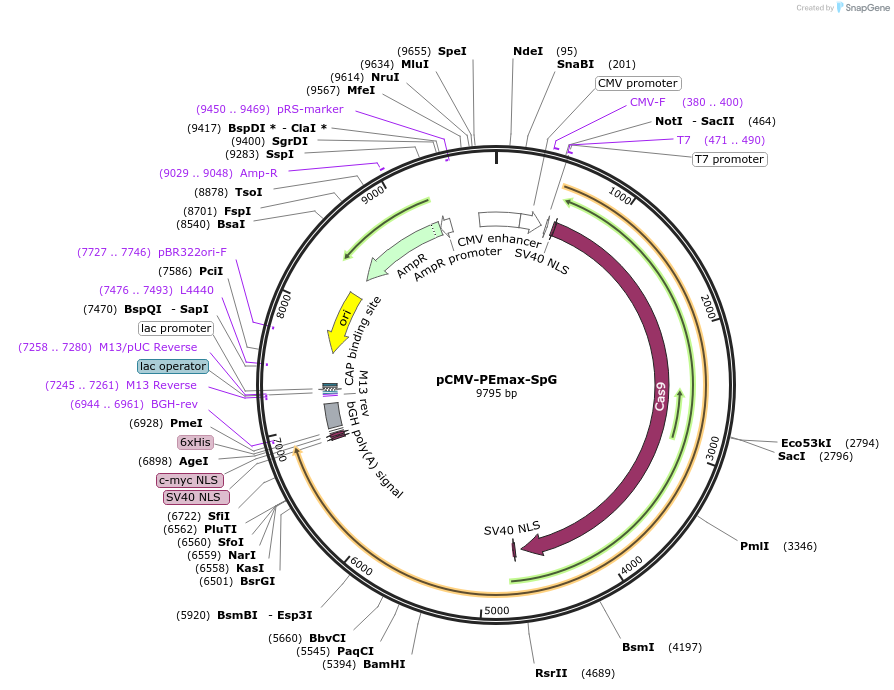
pCMV-PEmax-SpG
Plasmid#211549PurposePrime editing in mammalian cellsDepositorInsertPEmax-SpG
UseCRISPRExpressionMammalianPromoterCMVAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only