We narrowed to 15,312 results for: nts
-
Plasmid#30297DepositorInsertSC1918/H1 HA designed binding protein HB81
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
pETCON-HB82/design_21
Plasmid#30298DepositorInsertSC1918/H1 HA designed binding protein HB82
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
pETCON-HB7/design_52
Plasmid#30299DepositorInsertSC1918/H1 HA designed binding protein HB7
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
pETCON-HB9/design_41
Plasmid#30300DepositorInsertSC1918/H1 HA designed binding protein HB9
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
pETCON-IgG3/design_30
Plasmid#30302DepositorInsertIgG Fc Designed Binder IgG3
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
pETCON-IgG9/design_59
Plasmid#30303DepositorInsertIgG Fc Designed Binder IgG9
ExpressionYeastAvailable SinceJune 8, 2011AvailabilityAcademic Institutions and Nonprofits only -
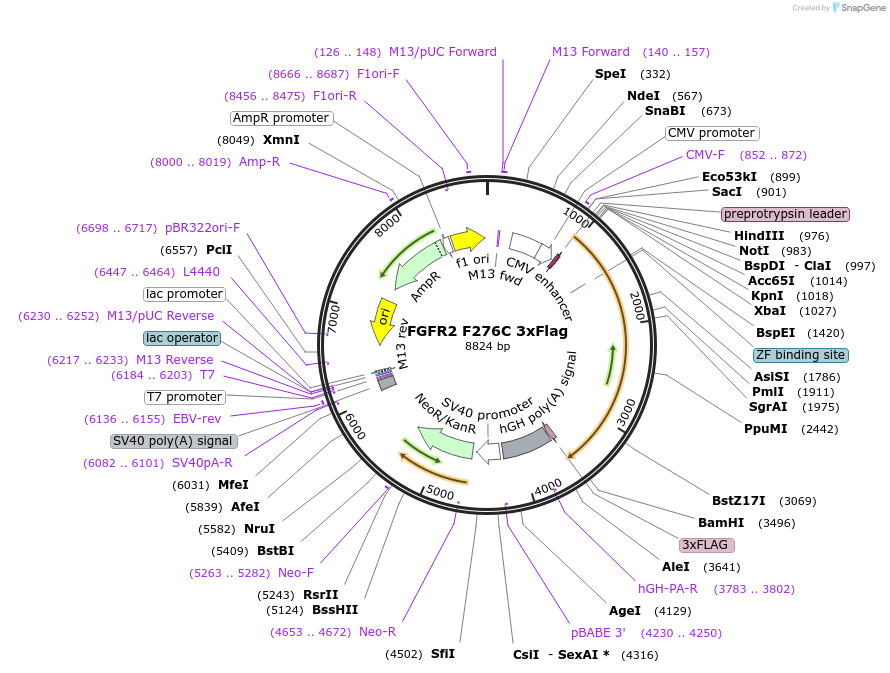
FGFR2 F276C 3xFlag
Plasmid#110064PurposeFGFR2 F276C expression in mammalian cells (JCOPO paper)DepositorInsertfibroblast growth factor receptor 2 (FGFR2 Human)
TagsFlagExpressionMammalianMutationmutated Phenylalanine 276 to Cysteine (F276C)PromoterCMVAvailable SinceMay 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
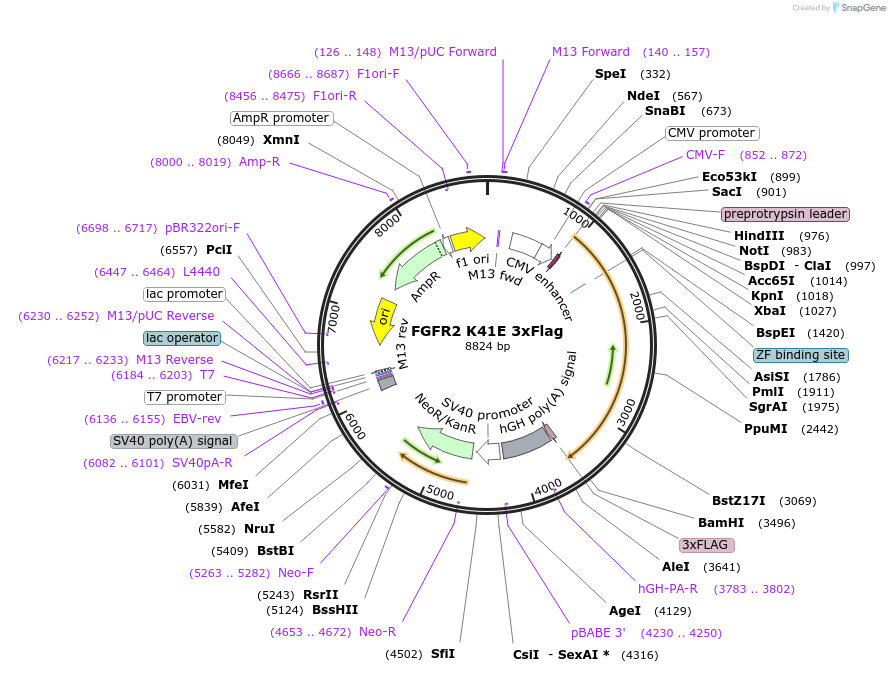
FGFR2 K41E 3xFlag
Plasmid#110105Purposeexpresses FGFR2 with a single amino acid change in mammalian cellsDepositorInsertfibroblast growth factor receptor 2 (FGFR2 Human)
TagsFlagExpressionMammalianMutationmutated Lysine 41 to Glutamic acid (K41E)PromoterCMVAvailable SinceMay 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
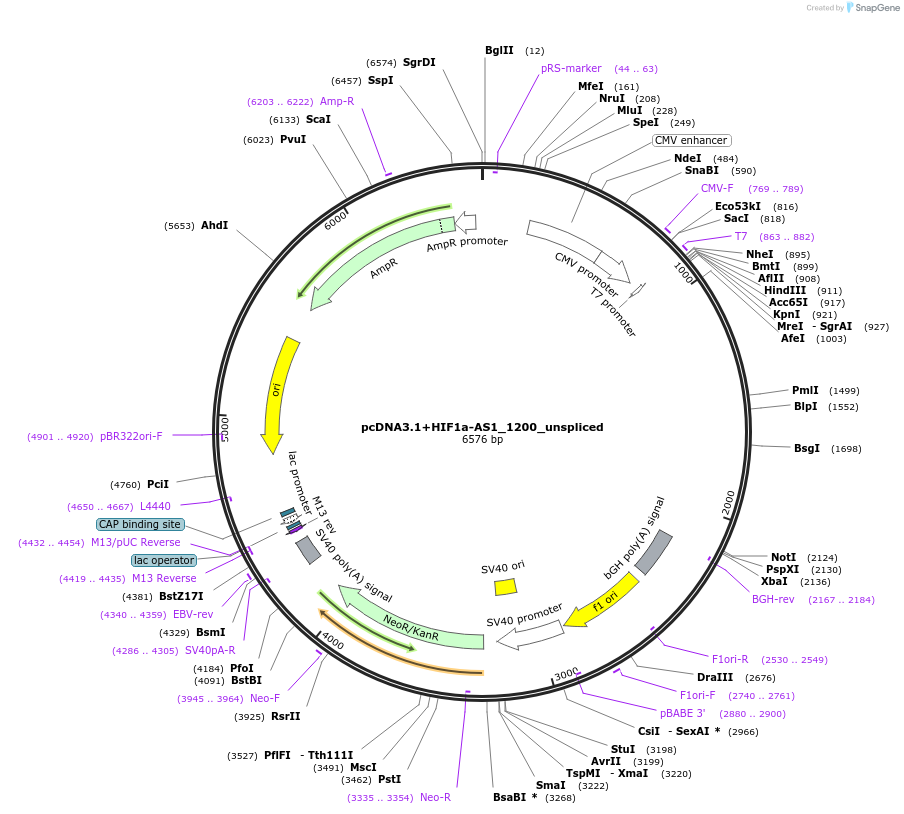
pcDNA3.1+HIF1a-AS1_1200_unspliced
Plasmid#194176PurposeExpresses first 1200 nt of lncRNA HIF1a-AS1 (unspliced)DepositorAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
E. coli BL21 (DE3) ΔEntD
Bacterial Strain#192874PurposeDeletion of native E. coli EntD, a 4'-PPTase required for native enterobactin synthesis. Allows expression of NRPS carrier proteins that do not harbor a phosphopantetheinyl group.DepositorBacterial ResistanceNoneSpeciesEscherichia coliAvailable SinceSept. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
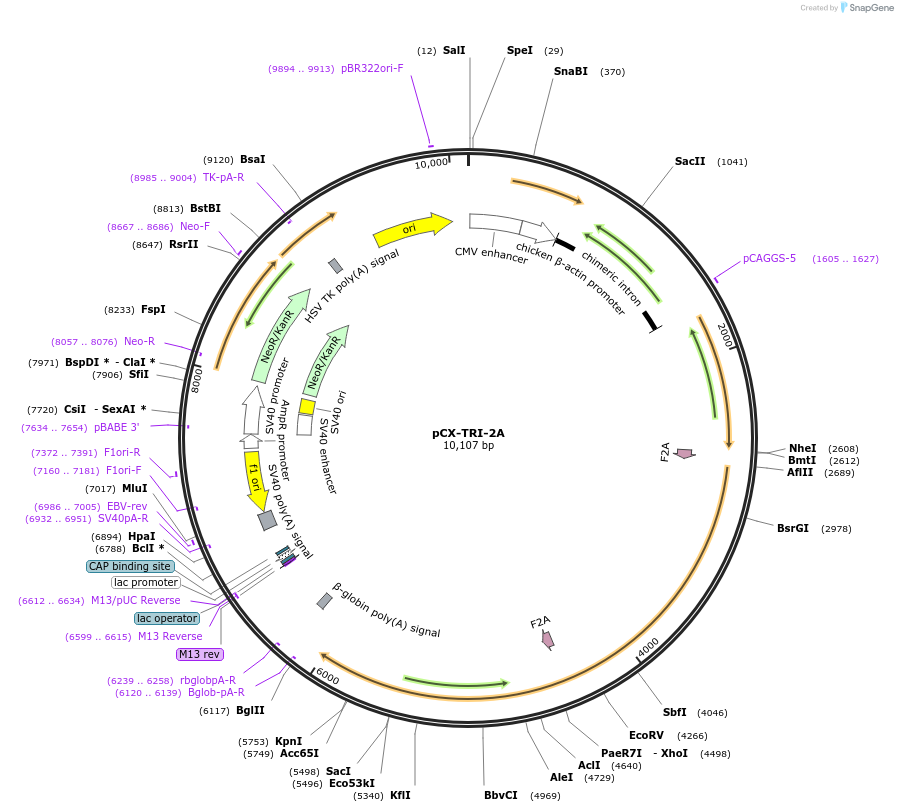
pCX-TRI-2A
Plasmid#74674Purposemammalian expression of TRI-2A (hHO1/F2A/hCD73/F2A/hCD39)DepositorTagsF2AExpressionMammalianPromoterpCAGGSAvailable SinceFeb. 14, 2017AvailabilityAcademic Institutions and Nonprofits only -
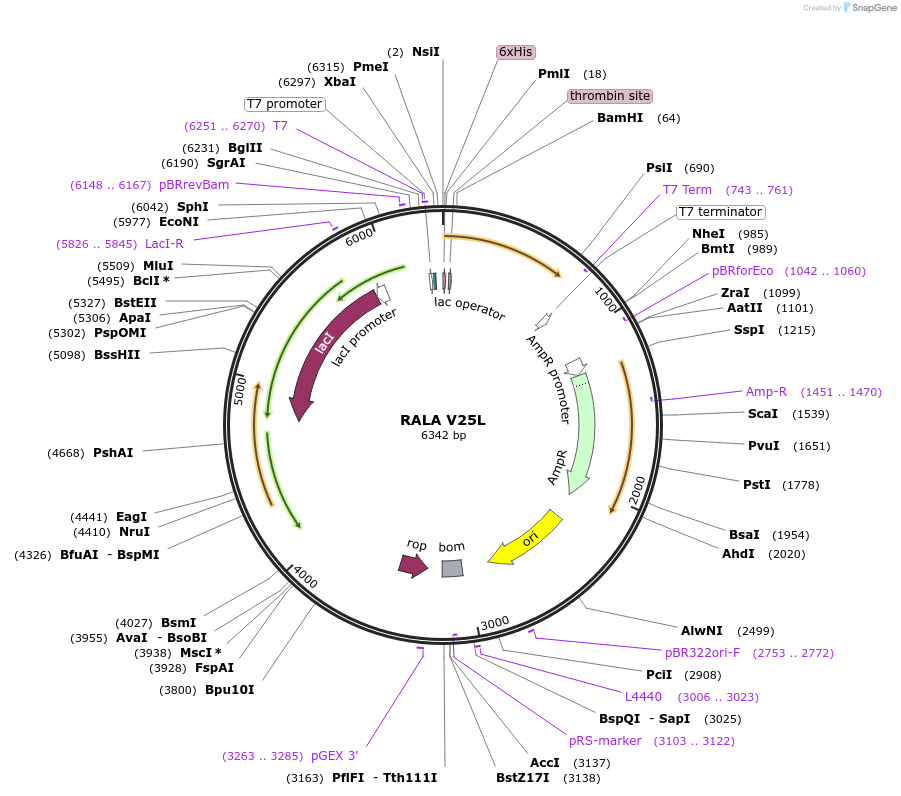
RALA V25L
Plasmid#122927PurposeBacterial expression of RALA V25LDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
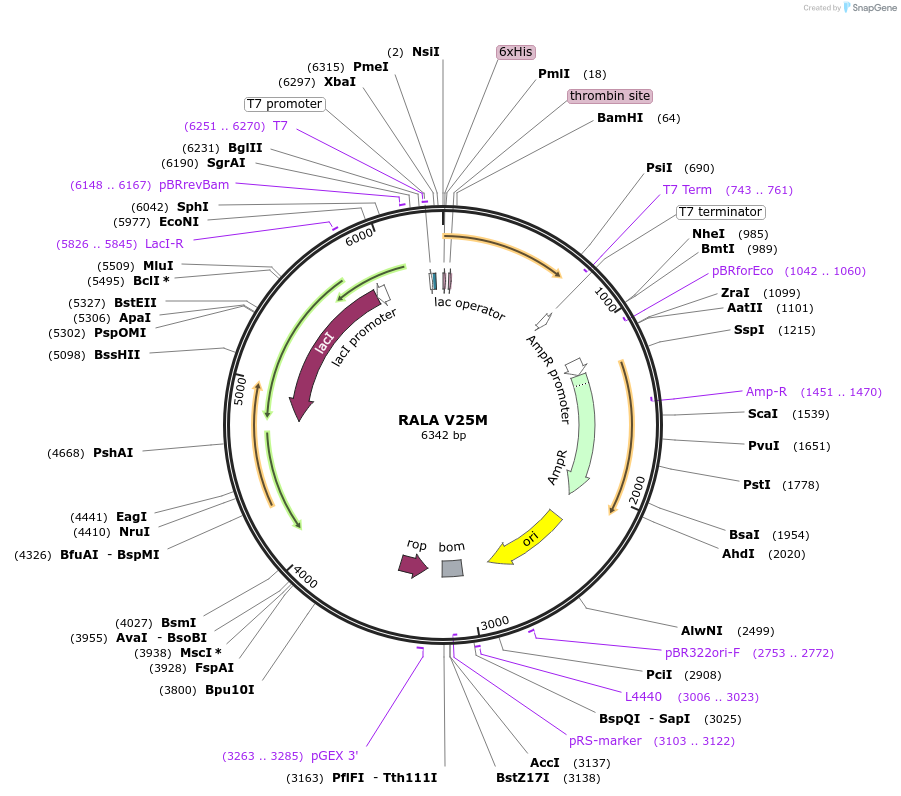
RALA V25M
Plasmid#122928PurposeBacterial expression of RALA V25MDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
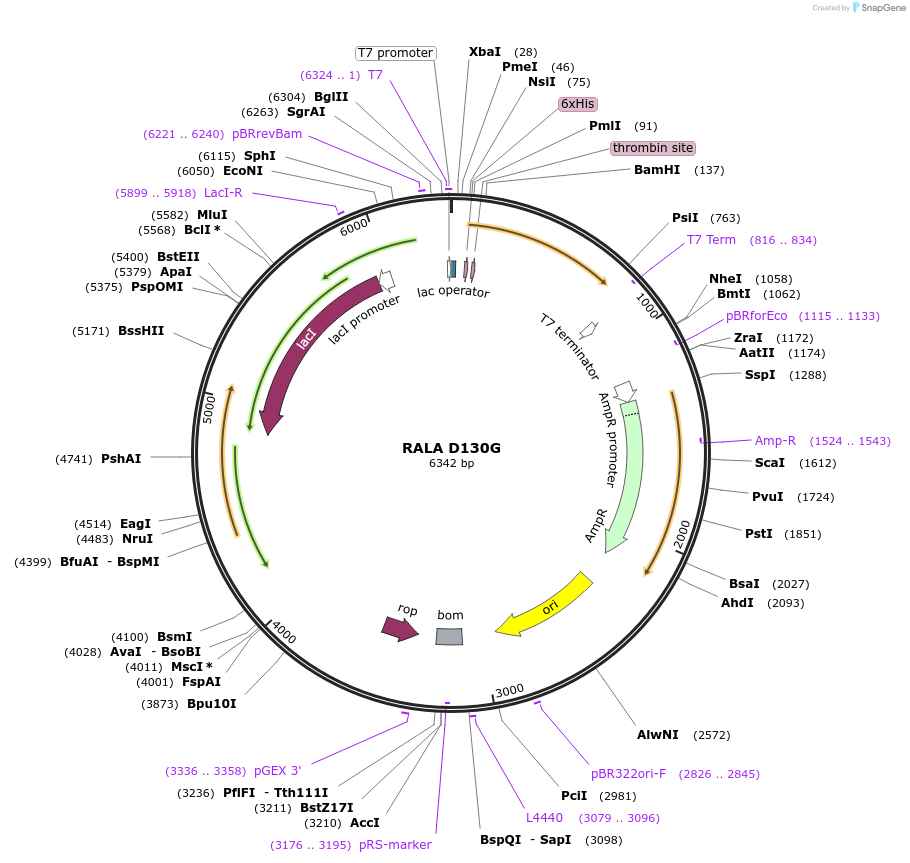
RALA D130G
Plasmid#122929PurposeBacterial expression of RALA D130GDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
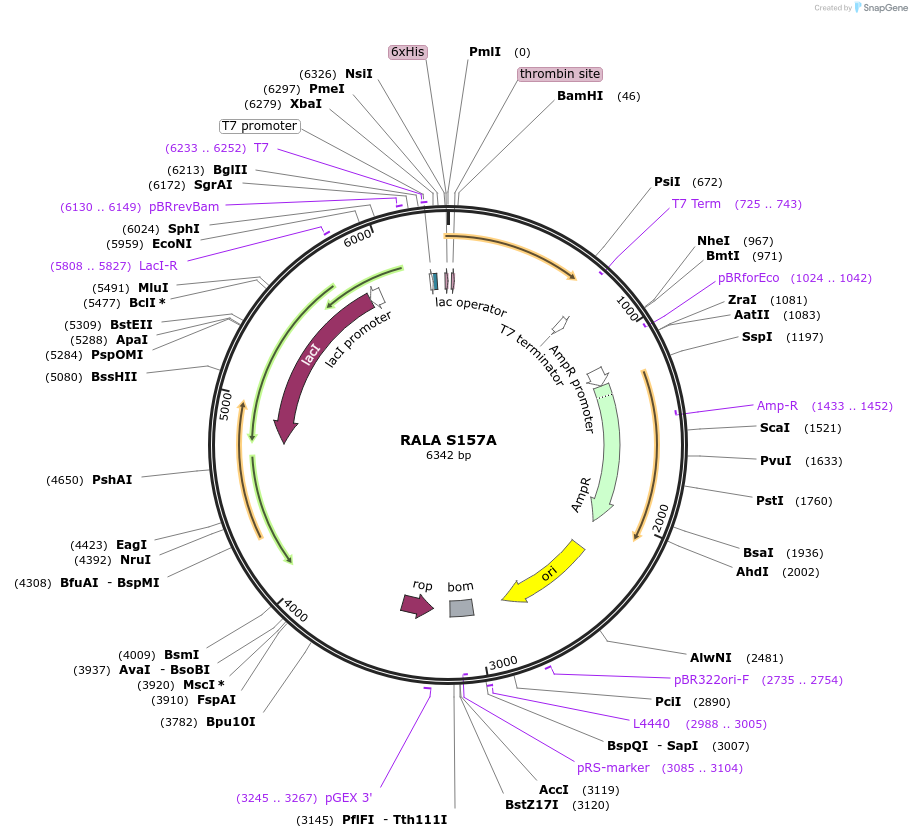
RALA S157A
Plasmid#122930PurposeBacterial expression of RALA S157ADepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
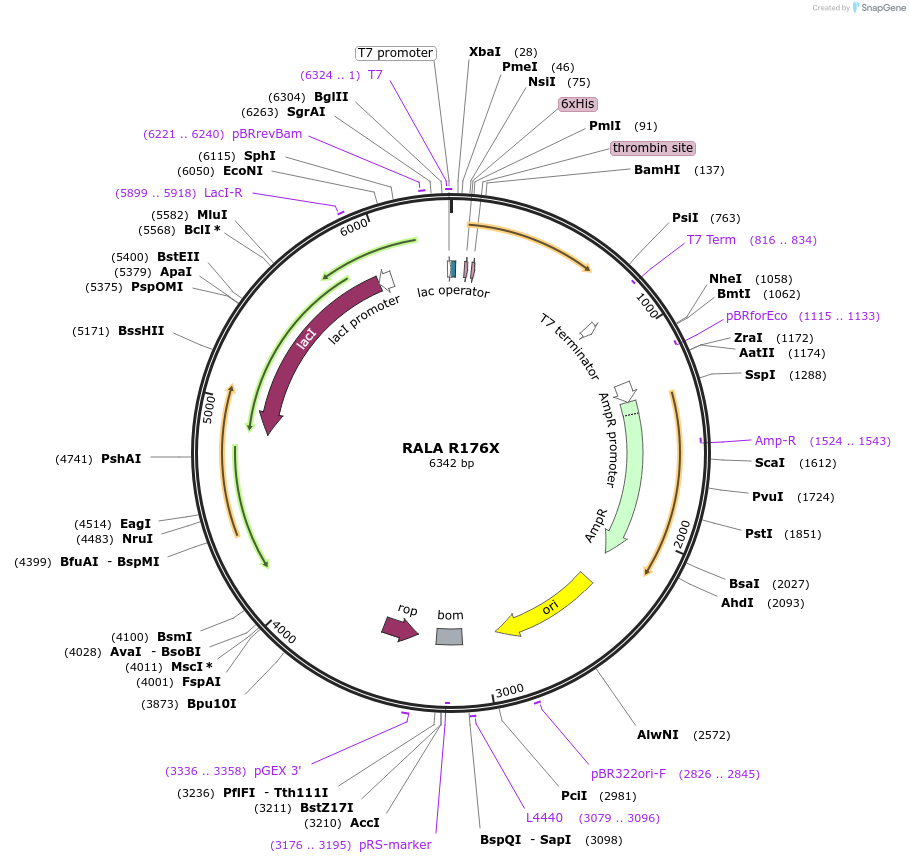
RALA R176X
Plasmid#122931PurposeBacterial expression of RALA R176XDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
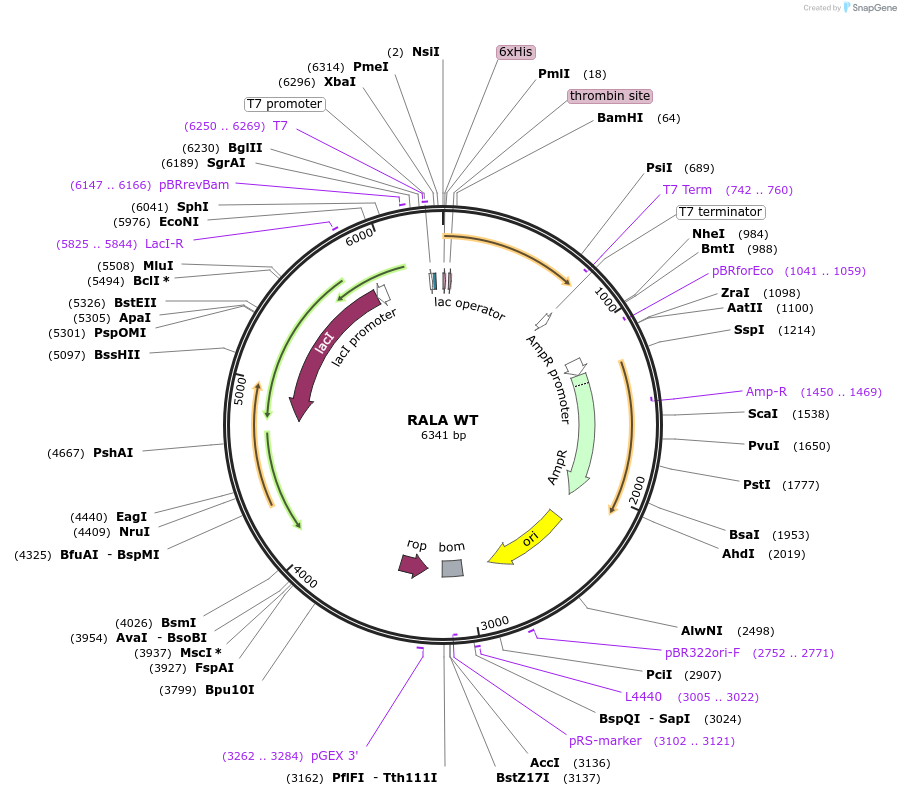
RALA WT
Plasmid#122925PurposeBacterial expression of RALA WTDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
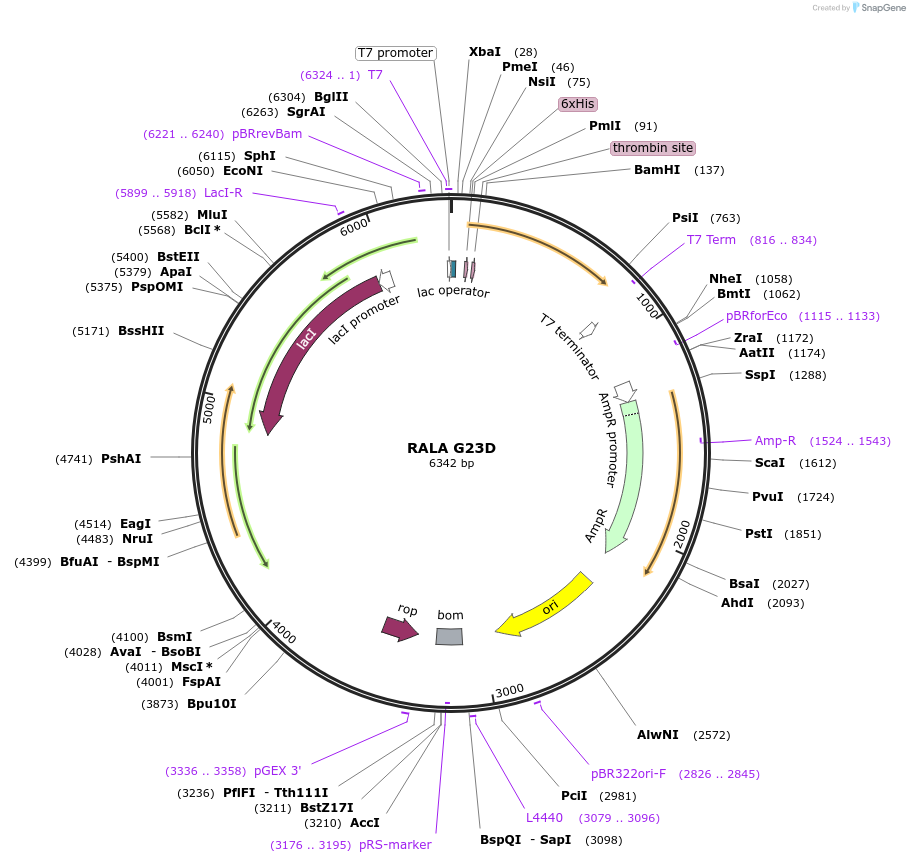
RALA G23D
Plasmid#122926PurposeBacterial expression of RALA G23DDepositorAvailable SinceApril 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
p11-5’_10xN_PAM-site3 (RTW572)
Pooled Library#160134PurposePlasmid library harboring a randomized 10nt PAM 5’ of target site 3. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 5’ PAMs (e.g. Cas12a)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
p11-3’_8xN_PAM-site1 (RTW554)
Pooled Library#160132PurposePlasmid library harboring a randomized 8nt PAM 3’ of target site 1. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 3’ PAMs (e.g. Cas9)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only