We narrowed to 9,757 results for: PEP
-
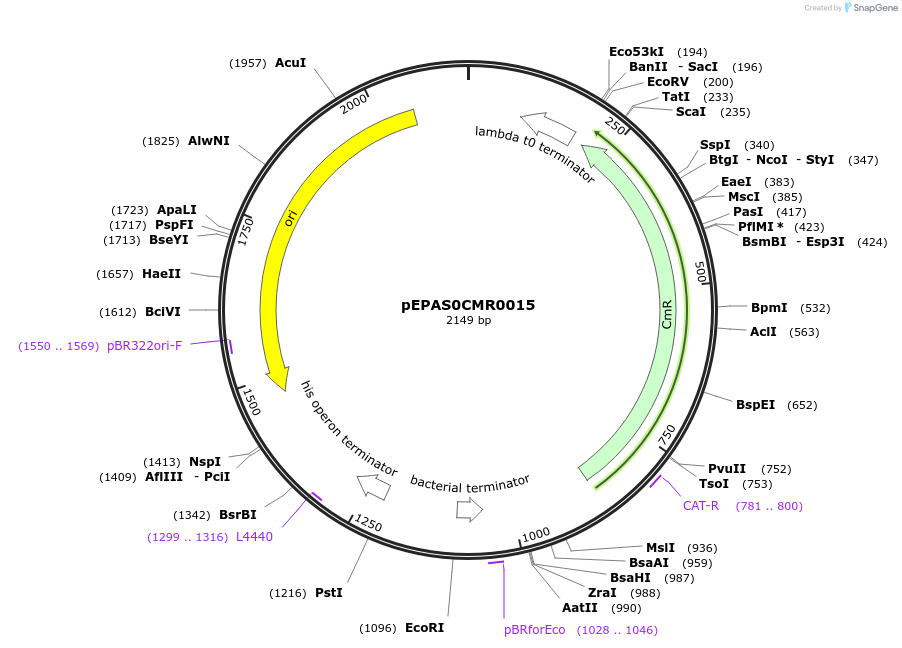
Plasmid#154560PurposePhytobricks Level 0 partDepositorInsertMinSyn_064
UseSynthetic BiologyAvailable SinceOct. 16, 2020AvailabilityAcademic Institutions and Nonprofits only -
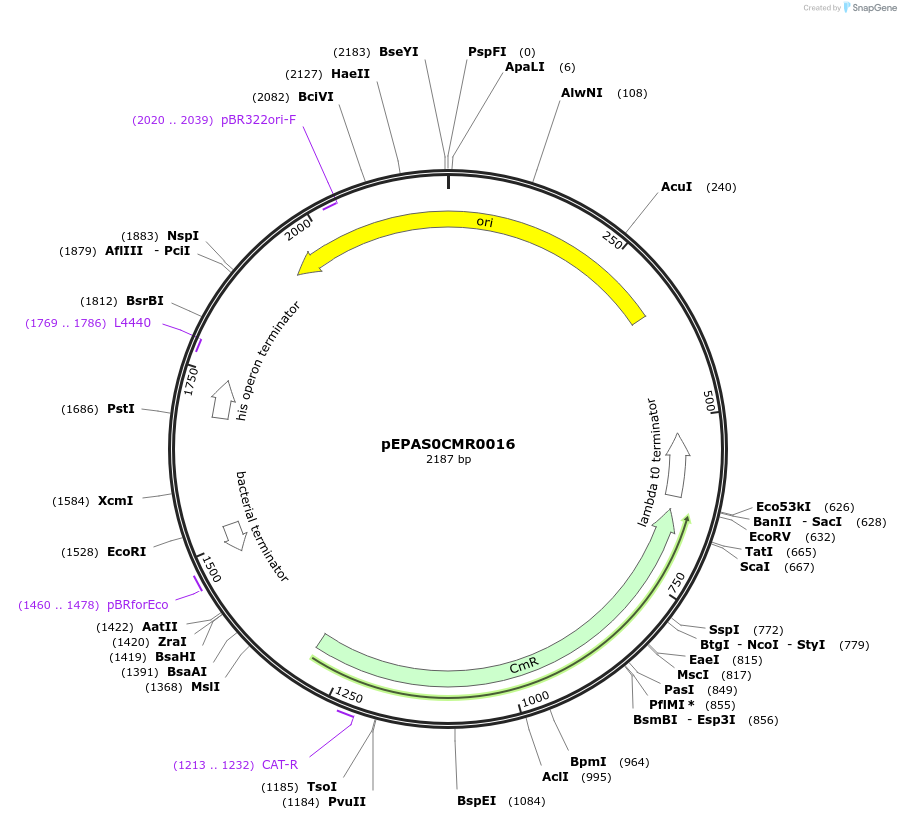
pEPAS0CMR0016
Plasmid#154561PurposePhytobricks Level 0 partDepositorInsertMinSyn_065
UseSynthetic BiologyAvailable SinceOct. 16, 2020AvailabilityAcademic Institutions and Nonprofits only -
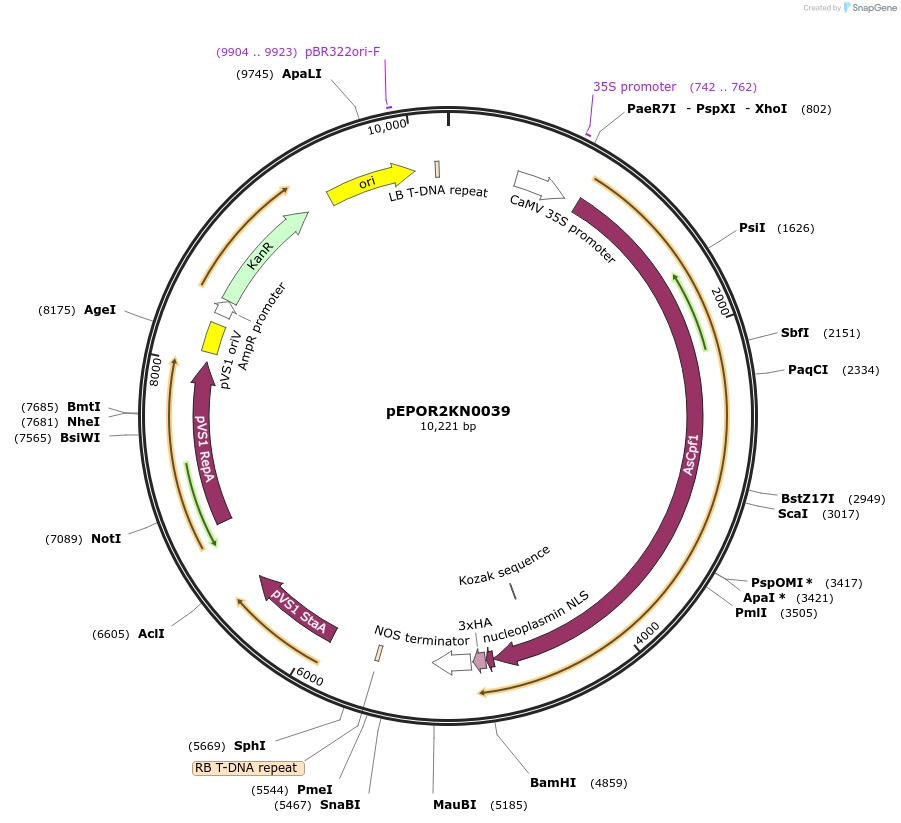
pEPOR2KN0039
Plasmid#117606PurposeLevel2 MoClo construct, containing level1 plant expression cassettes for AsCas12a(pEPOR1CB0014) and sgRNA_AtADH1 {AT1G77120}(pEPOR1CB0028)DepositorInsert[35S:AsCas12a(pEPOR1CB0014) ] +[AtU6-26:sgRNA]
ExpressionPlantAvailable SinceNov. 16, 2018AvailabilityAcademic Institutions and Nonprofits only -
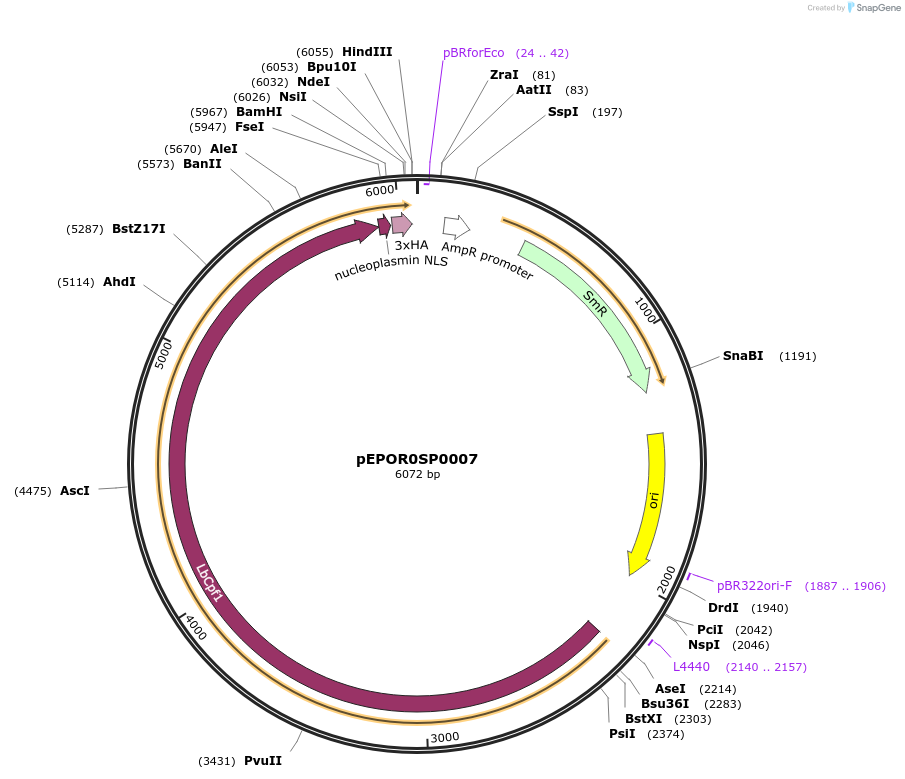
pEPOR0SP0007
Plasmid#117534PurposeLevel0 Golden Gate part: CDS, LbCas12aDepositorInsertLbCas12a (with stop codon)
UseCRISPRAvailable SinceNov. 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
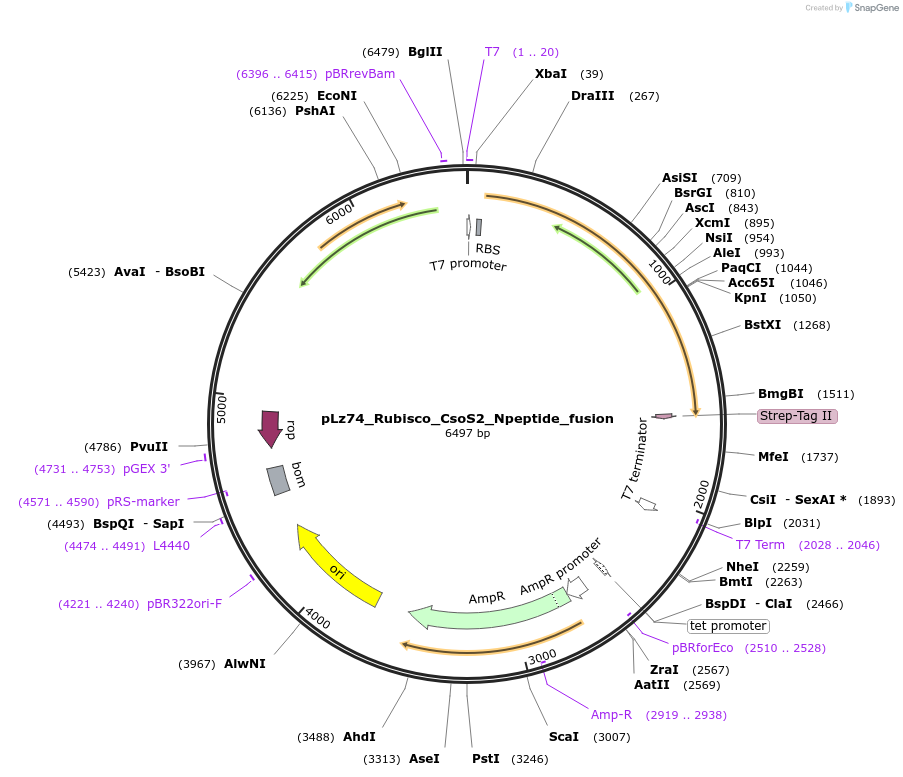
pLz74_Rubisco_CsoS2_Npeptide_fusion
Plasmid#140840PurposeSame as pBz15 but with N*-peptide fusion on CbbL, construct used for protein crystallizationDepositorInsertForm IA Rubisco with CsoS2 N-peptide fusion; CbbL-Npeptide, CbbS
TagsCsoS2 N-peptide consensus to CbbL C-terminus and …ExpressionBacterialPromoterT7Available SinceMay 8, 2020AvailabilityAcademic Institutions and Nonprofits only -
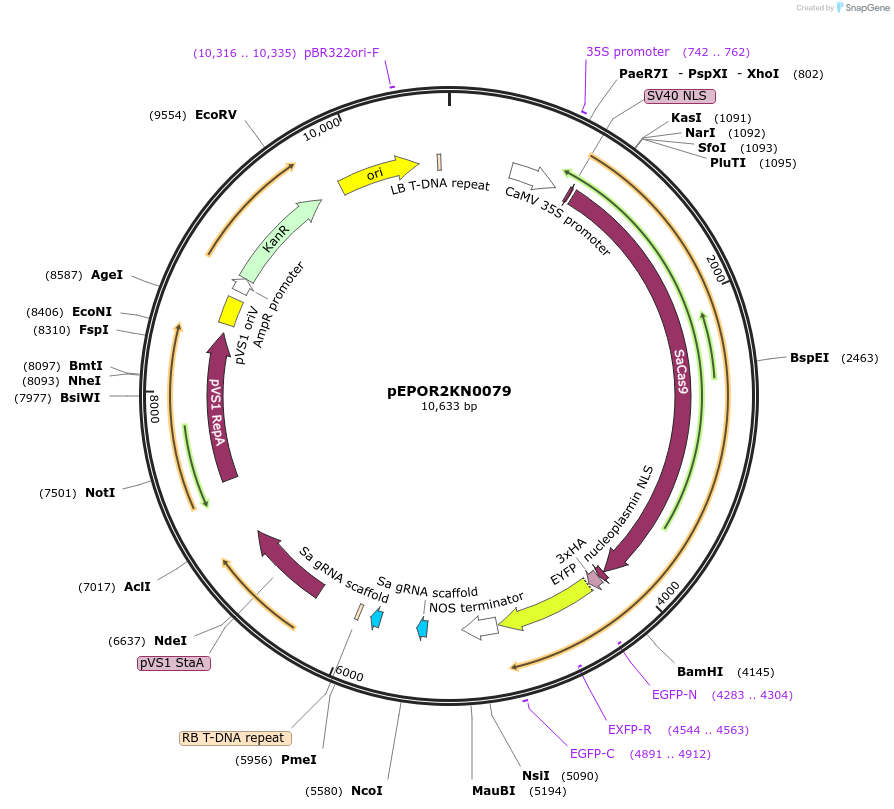
pEPOR2KN0079
Plasmid#117646PurposeLevel2 MoClo construct, containing level1 plant expression cassettes for SaCas9(pEPOR1CB0015) and sgRNA_ENTH/ANT {AT1G14910}(pEPOR1CB0100)and sgRNA_ENTH/ANT {AT2G01600}(pEPOR1CB0111)DepositorInsert[35S:SaCas9(pEPOR1CB0015) ] +[AtU6-26:sgRNA]
ExpressionPlantAvailable SinceNov. 16, 2018AvailabilityAcademic Institutions and Nonprofits only -
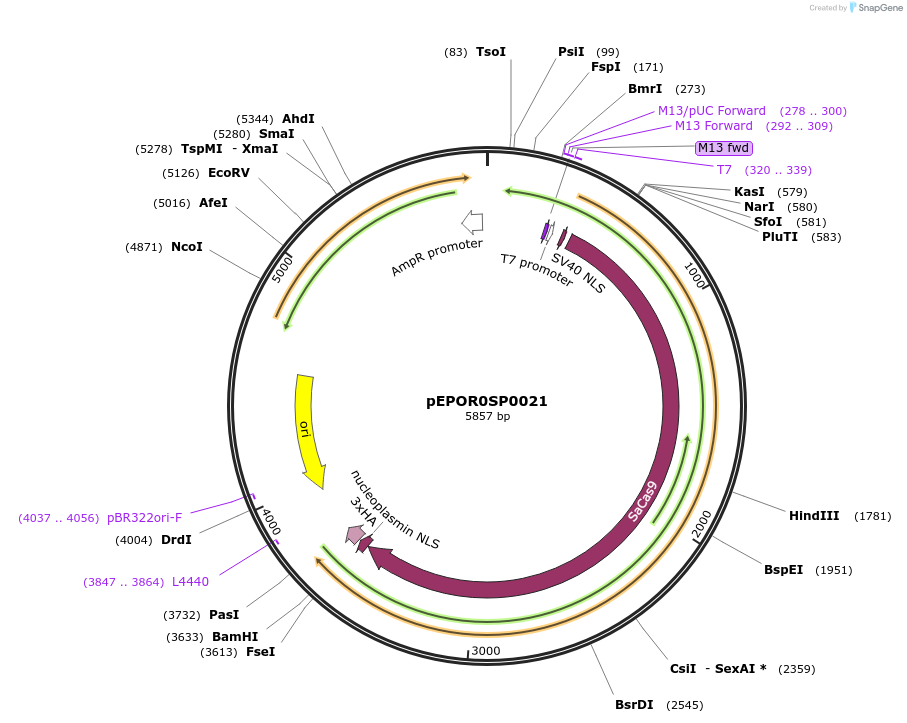
pEPOR0SP0021
Plasmid#117532PurposeLevel0 Golden Gate part: CDS, SaCas9 (no stop codon)DepositorInsertSaCas9 (no stop codon)
UseCRISPRAvailable SinceApril 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
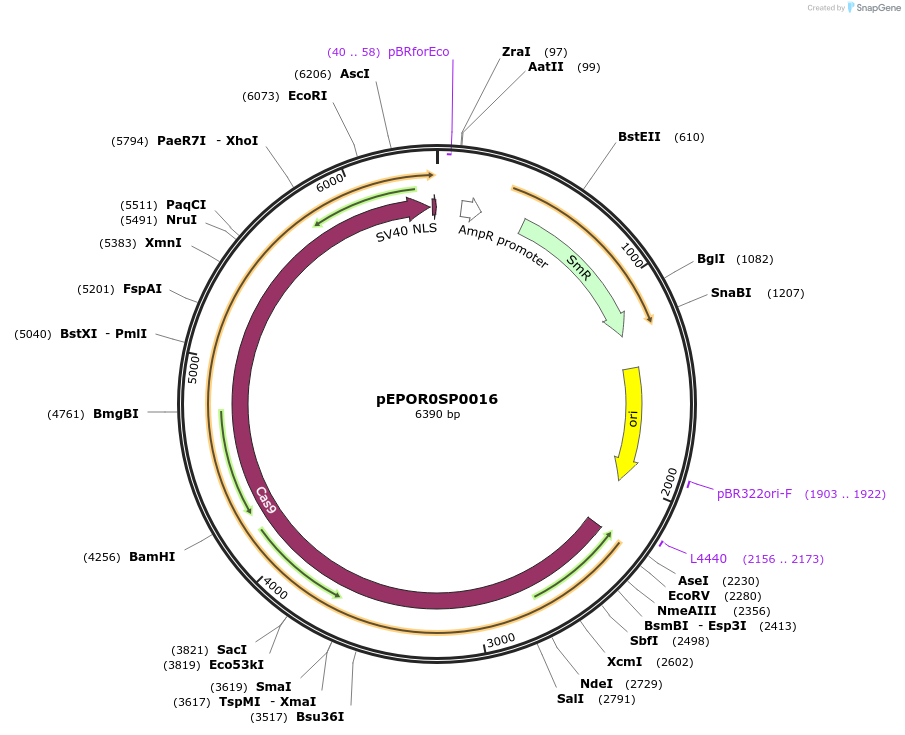
pEPOR0SP0016
Plasmid#117526PurposeLevel0 Golden Gate part: CDS, Sp-eCas9 1.0DepositorInsertSp-eCas9 1.0 (with stop codon)
UseCRISPRMutationK810A, K1003A, R1060AAvailable SinceNov. 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
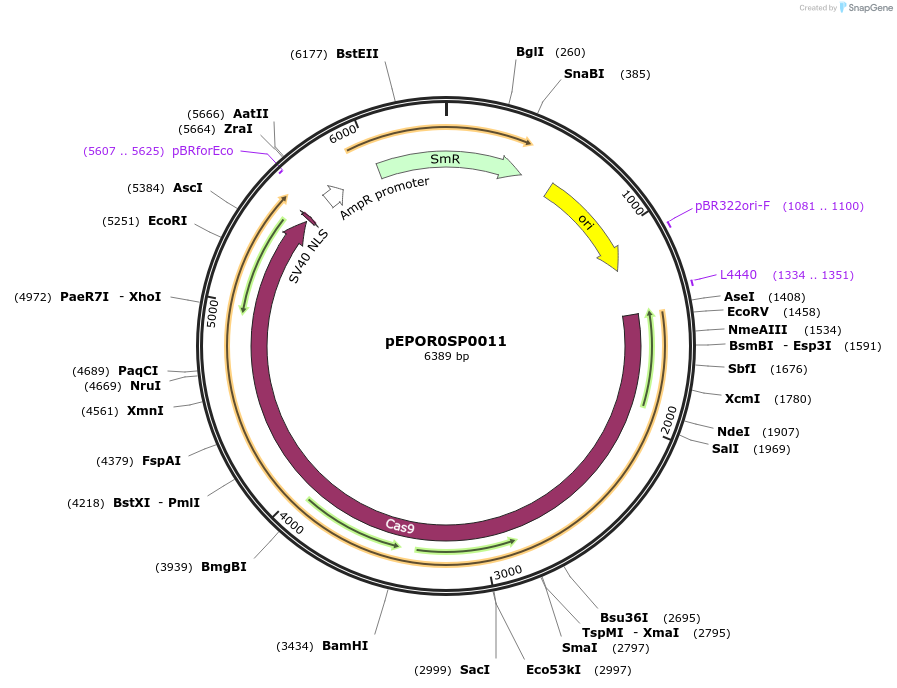
pEPOR0SP0011
Plasmid#117530PurposeLevel0 Golden Gate part: CDS, Sp-xCas9 3.7 (no stop codon)DepositorInsertSp-xCas9 3.7 (no stop codon)
UseCRISPRMutationA262T, R324L, S409I, E480K, E543D, M694I, E1219VAvailable SinceNov. 7, 2018AvailabilityAcademic Institutions and Nonprofits only -
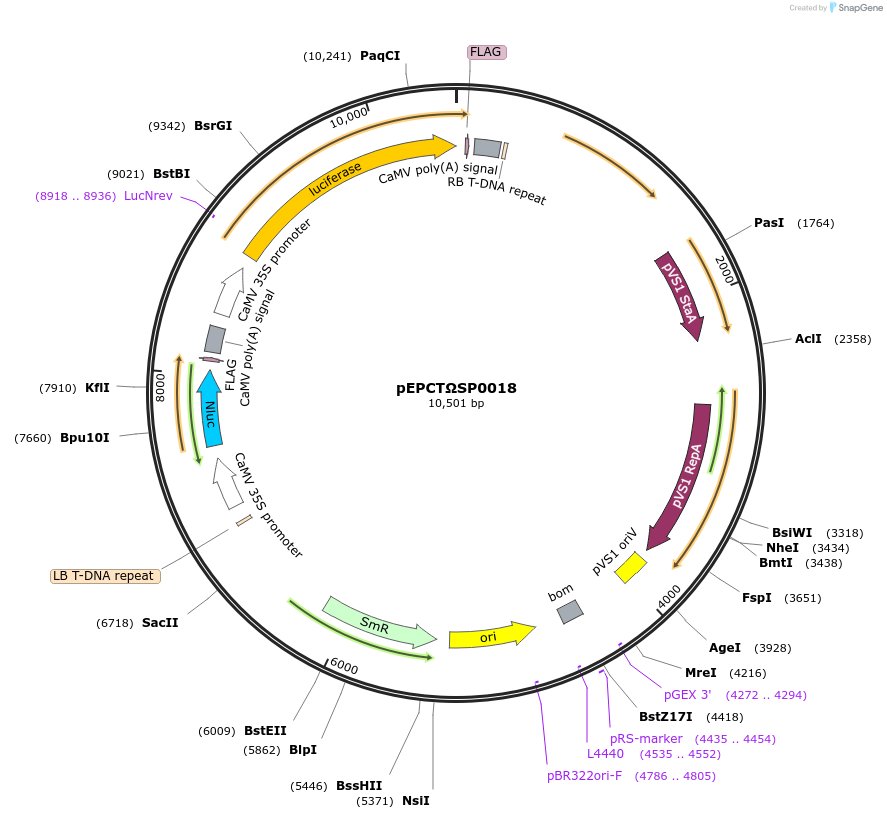
pEPCTΩSP0018
Plasmid#187566PurposeModule for constitutive expression of nanoluc luciferase and firefly luciferase driven by 35s promotersDepositorInsertFirefly Luciferase and NanoLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
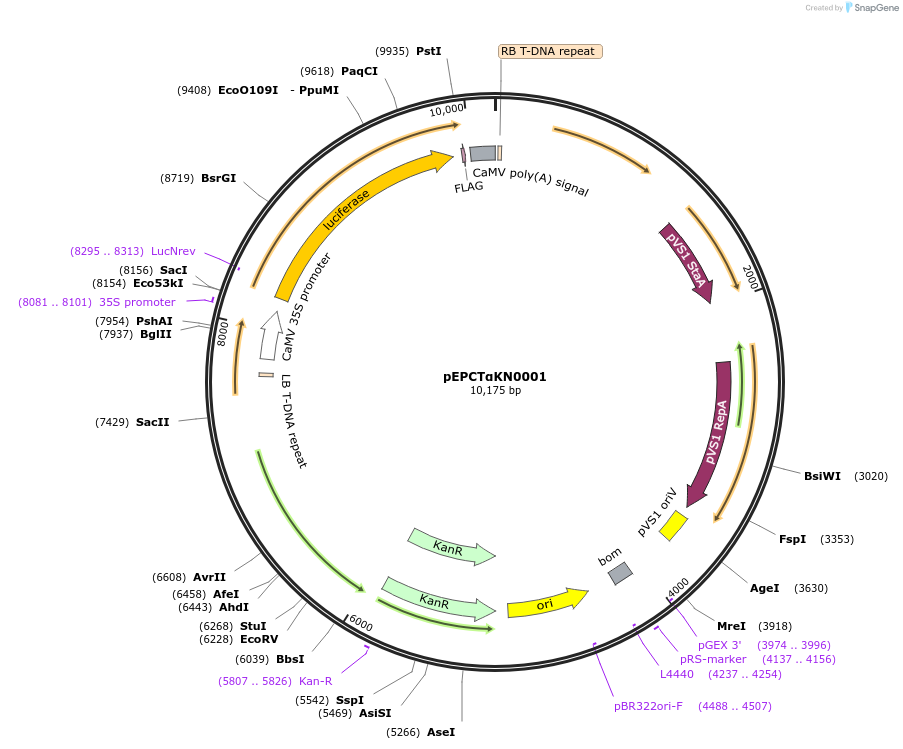
pEPCTαKN0001
Plasmid#187568PurposeTranscriptional unit for constitutive expression of firefly luciferase driven by the 35s promoterDepositorInsertLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
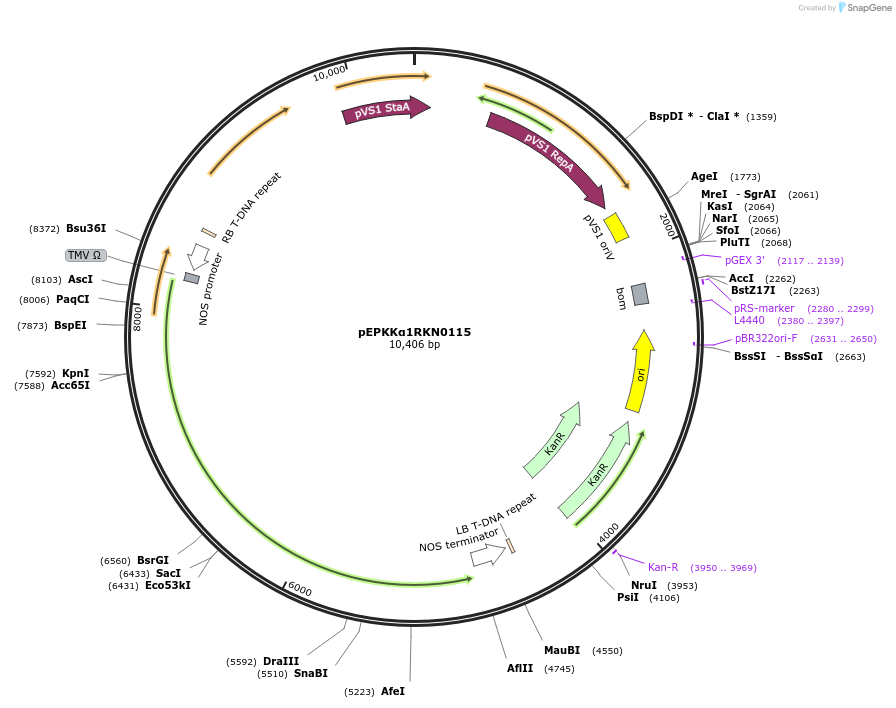
pEPKKα1RKN0115
Plasmid#187571PurposeTranscriptional unit for constitutive expression of a TALE driven by the nos promoterDepositorInsertTALE
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
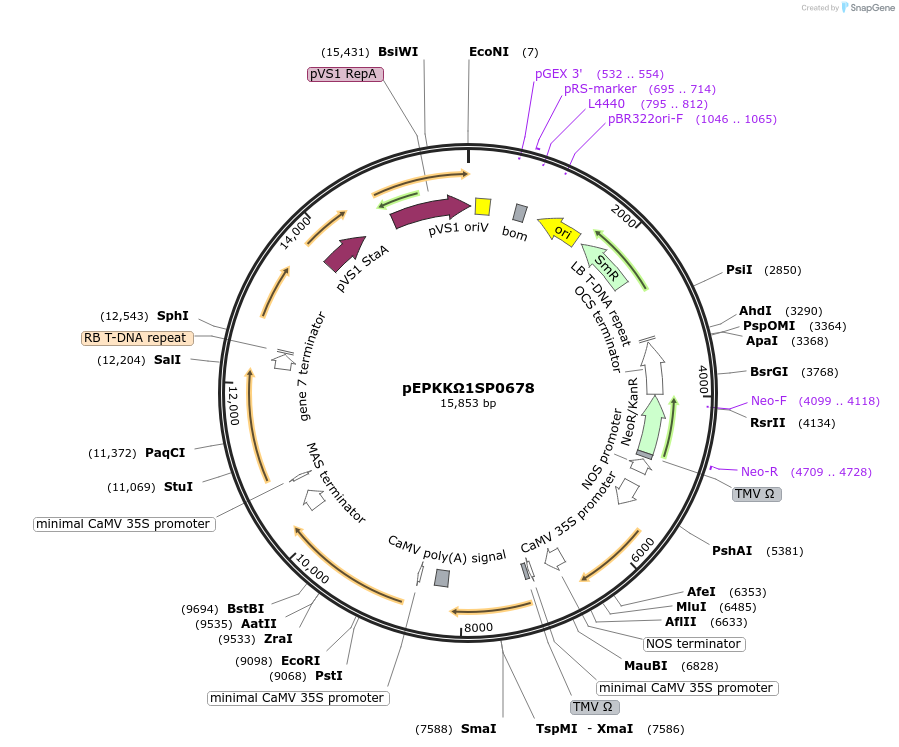
pEPKKΩ1SP0678
Plasmid#187605PurposeModule for copper inducible expression of AtrD11, HarFAR and EaDActDepositorInsert35s:TMVΩ:CUP2:GAl4:nos
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
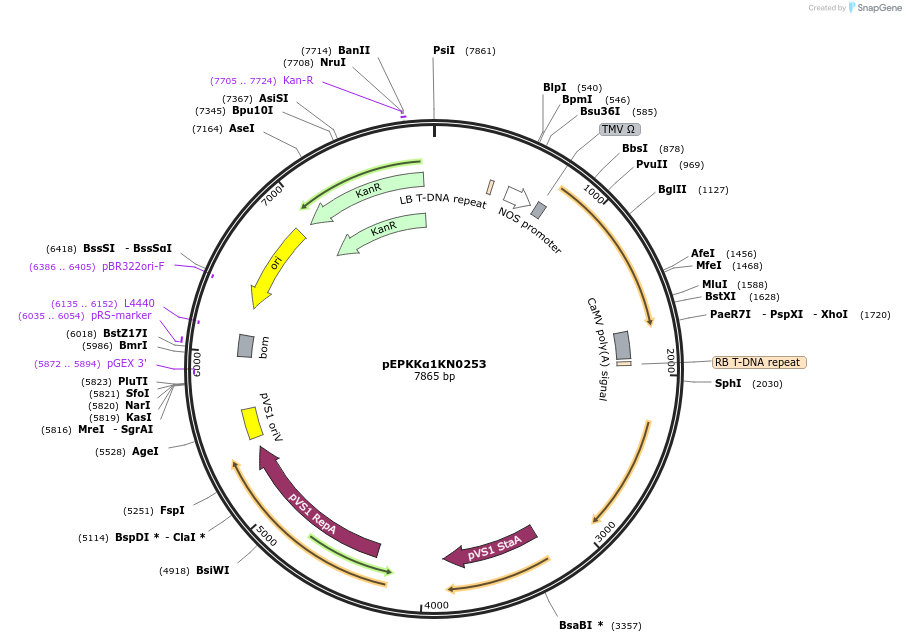
pEPKKα1KN0253
Plasmid#187560PurposeTranscriptional unit for constitutive expression of CUP2:GAL4 driven by the nos promoterDepositorInsertCUP2:GAL4
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
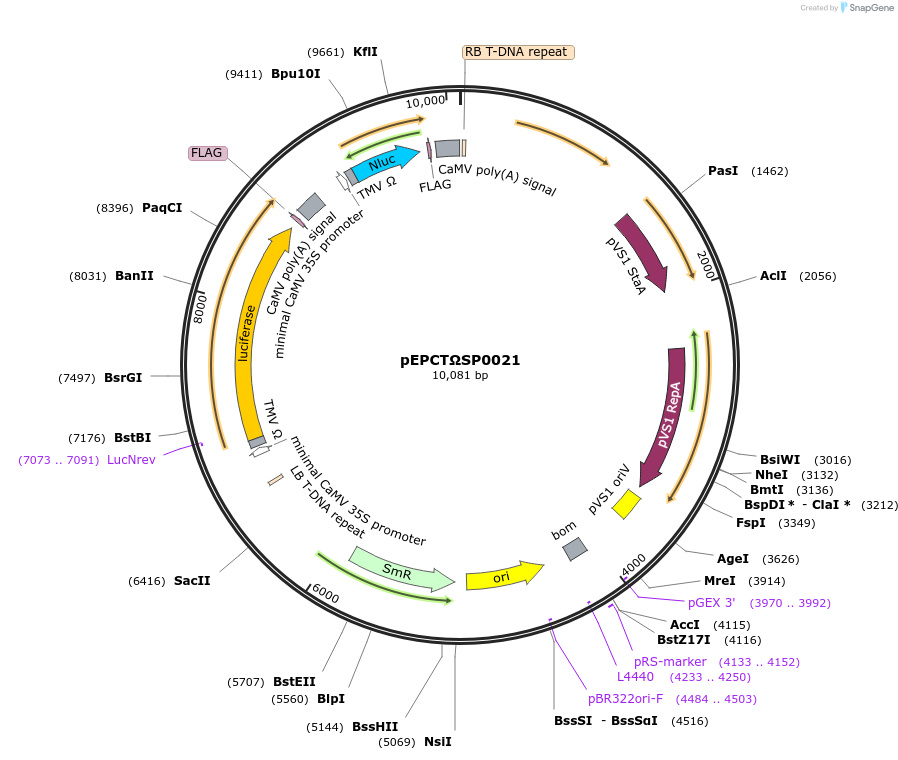
pEPCTΩSP0021
Plasmid#187561PurposeModule for copper-inducible expression of firefly luciferase and nanoluc luciferase driven by minimal synthetic promoters with binding sites for CUP2DepositorInsertFirefly Luciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
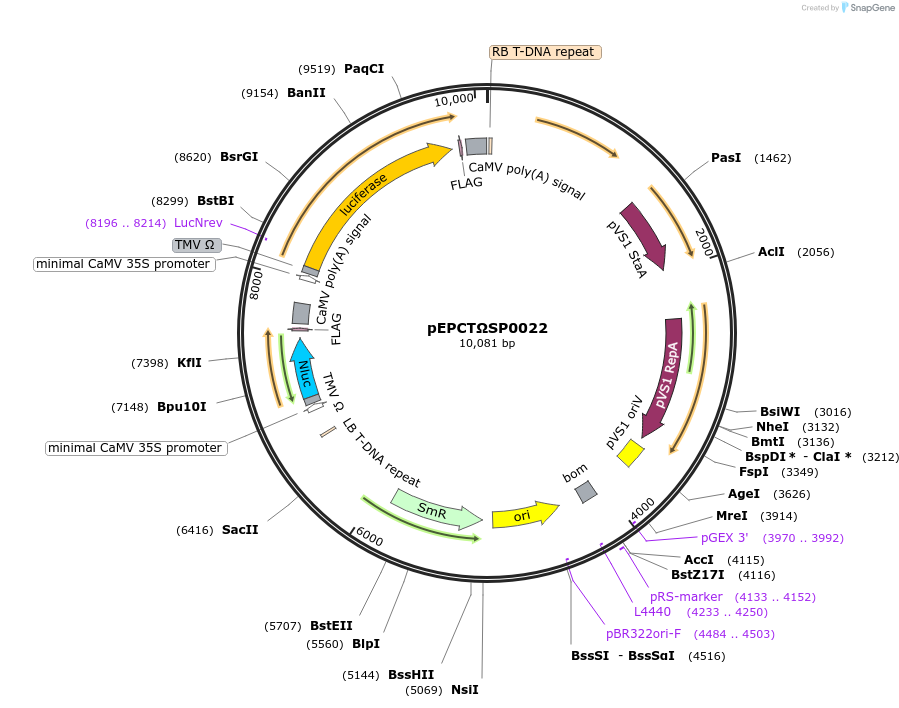
pEPCTΩSP0022
Plasmid#187562PurposeModule for copper-inducible expression of nanoluc luciferase and firefly luciferase driven by minimal synthetic promoters with binding sites for CUP2DepositorInsertFirefly Luciferase and NanoLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
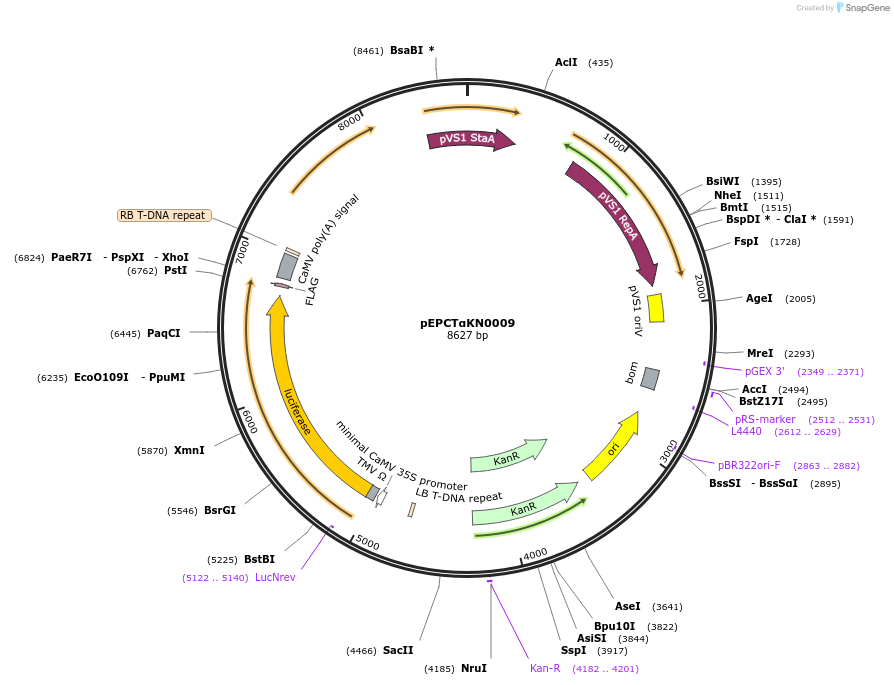
pEPCTαKN0009
Plasmid#187563PurposeTranscriptional unit for copper-inducible expression of firefly luciferase driven by a minimal synthetic promoter with binding sites for CUP2DepositorInsertLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
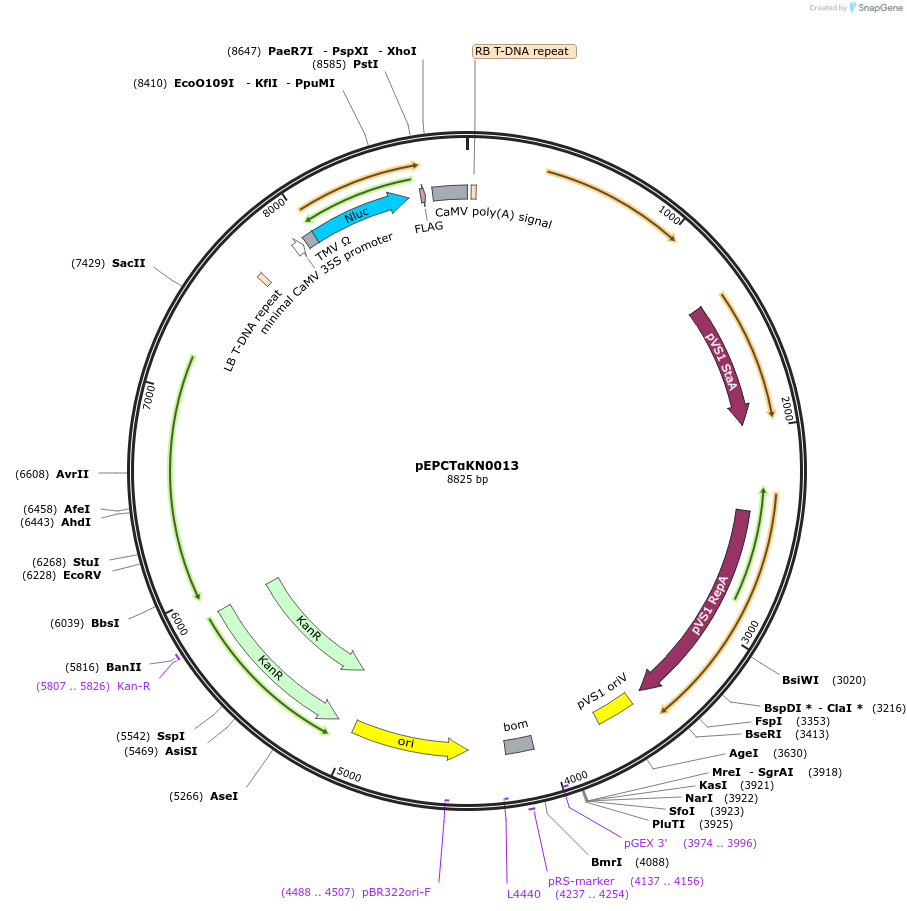
pEPCTαKN0013
Plasmid#187564PurposeTranscriptional unit for copper-inducible expression of nanoluc luciferase driven by a minimal synthetic promoter with binding sites for CUP2DepositorInsertLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
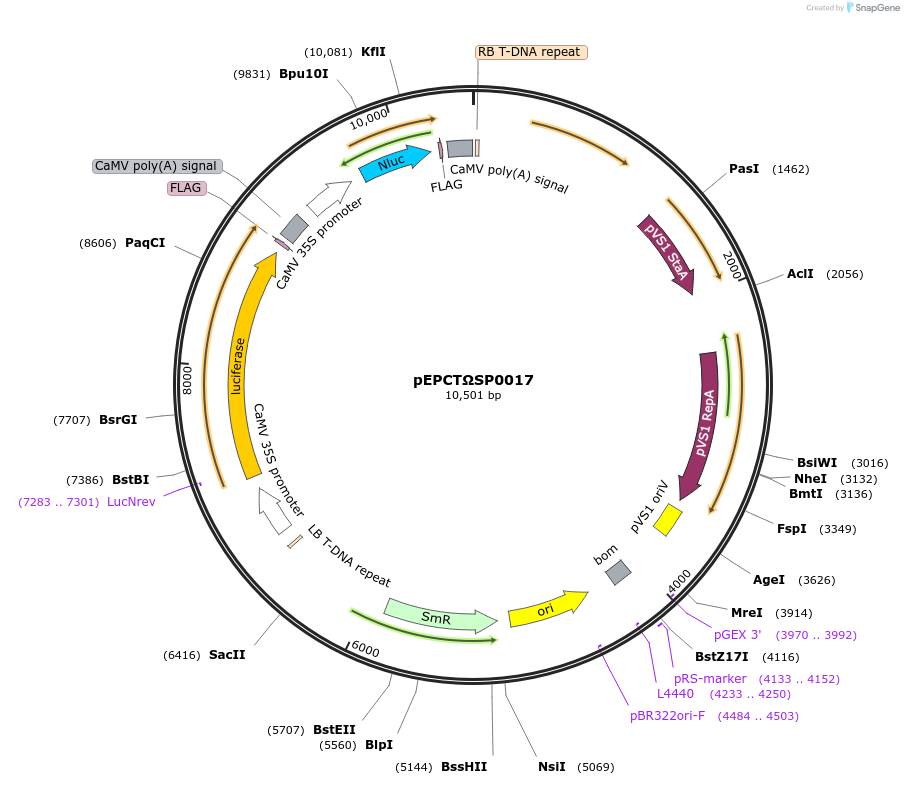
pEPCTΩSP0017
Plasmid#187565PurposeModule for constitutive expression of firefly luciferase and nanoluc luciferase driven by 35s promotersDepositorInsertFirefly Luciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
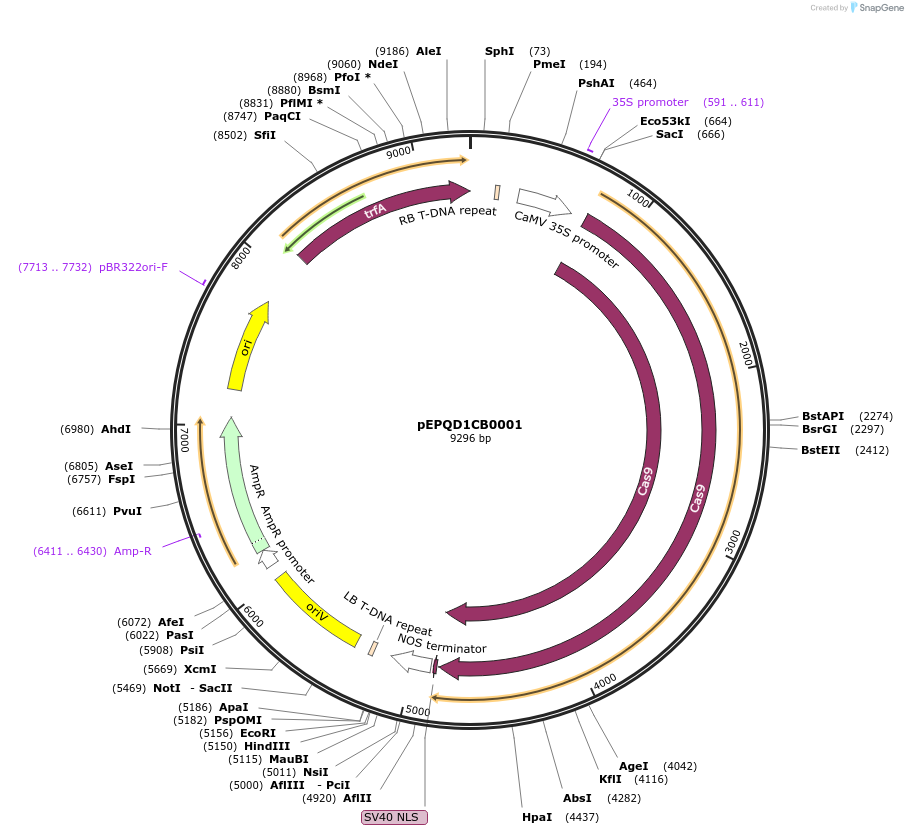
pEPQD1CB0001
Plasmid#185625PurposeMoClo Level 1, position 2 reverse, transcriptional unit for expression of plant codon optimized Cas9 from Streptococcus pyogenes driven by 35S promoterDepositorInsertSpCas9
UseCRISPR and Synthetic BiologyAvailable SinceAug. 4, 2022AvailabilityAcademic Institutions and Nonprofits only