We narrowed to 20,365 results for: REN
-
Plasmid#203948PurposeLevel 0 part. Cas12a coding sequence from Francisella novicida in CDS1 positionDepositorInsertFnCas12a
UseSynthetic BiologyAvailable SinceMay 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
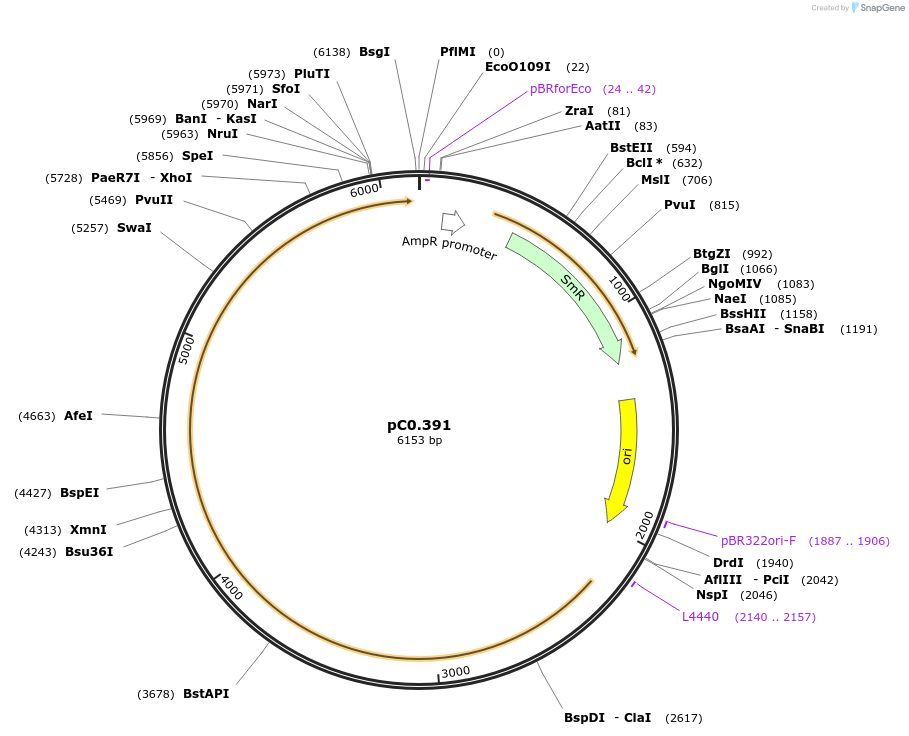
pC0.385
Plasmid#203936PurposeUp Flank. Neutral site. Up flanking sequence of ω3 acyl-lipid desaturase (desB, FEK30_04840) in Synechococcus sp. PCC 11901.DepositorInsertdesB UP
UseSynthetic BiologyAvailable SinceMay 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
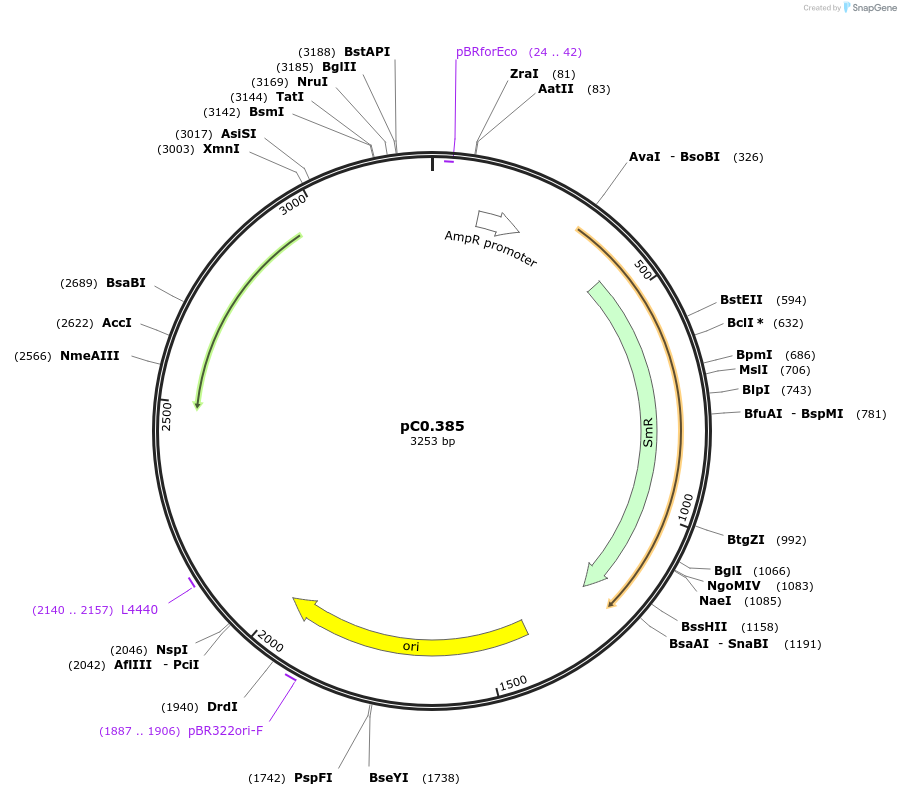
pC0.386
Plasmid#203938PurposeUp Flank. Neutral site. Up flanking sequence of glycogen synthase A1 (glgA1, FEK30_14880) in Synechococcus sp. PCC 11901.DepositorInsertglgA1 UP
UseSynthetic BiologyAvailable SinceMay 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
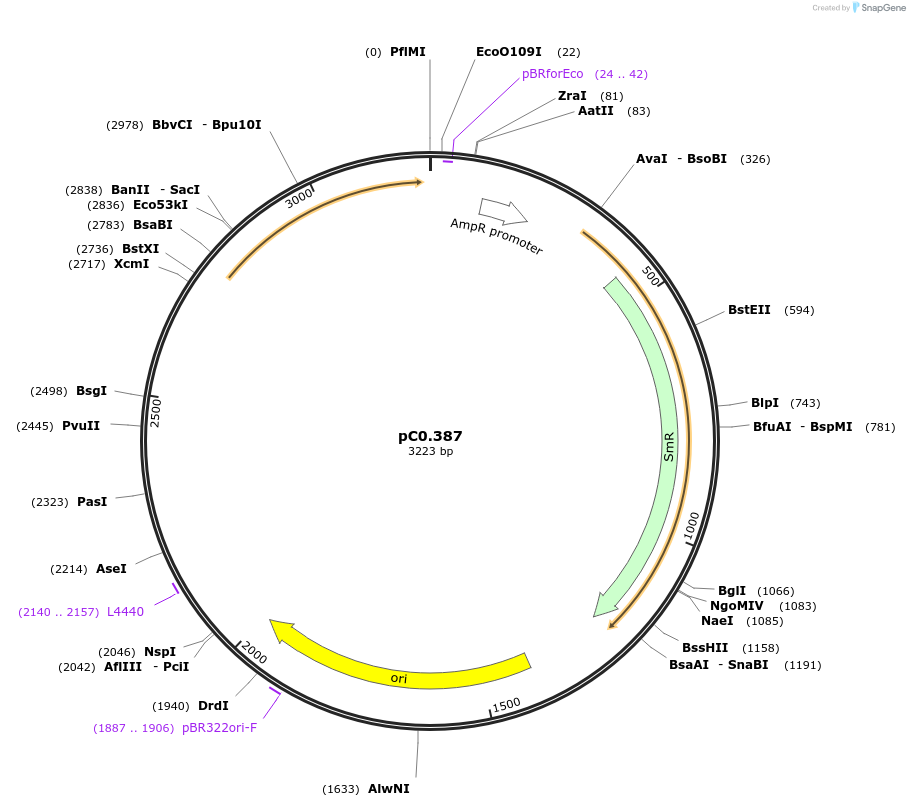
pC0.387
Plasmid#203939PurposeDown Flank. Neutral site. Down flanking sequence of glycogen synthase A1 (glgA1, FEK30_14880) in Synechococcus sp. PCC 11901.DepositorInsertglgA1 DOWN
UseSynthetic BiologyAvailable SinceMay 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
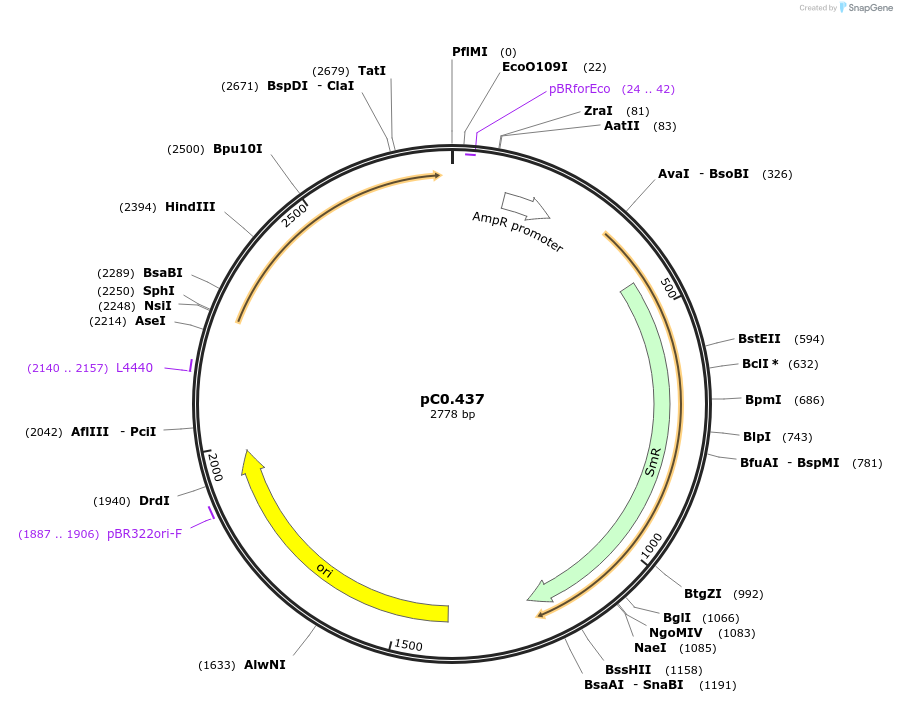
pC0.437
Plasmid#203942PurposeUp Flank. Neutral site. Up flanking sequence of intergenic region (NS1) between FEK30_11550 and FEK30_11555 in Synechococcus sp. PCC 11901.DepositorInsertNS1 UP
UseSynthetic BiologyAvailable SinceMay 10, 2024AvailabilityAcademic Institutions and Nonprofits only -
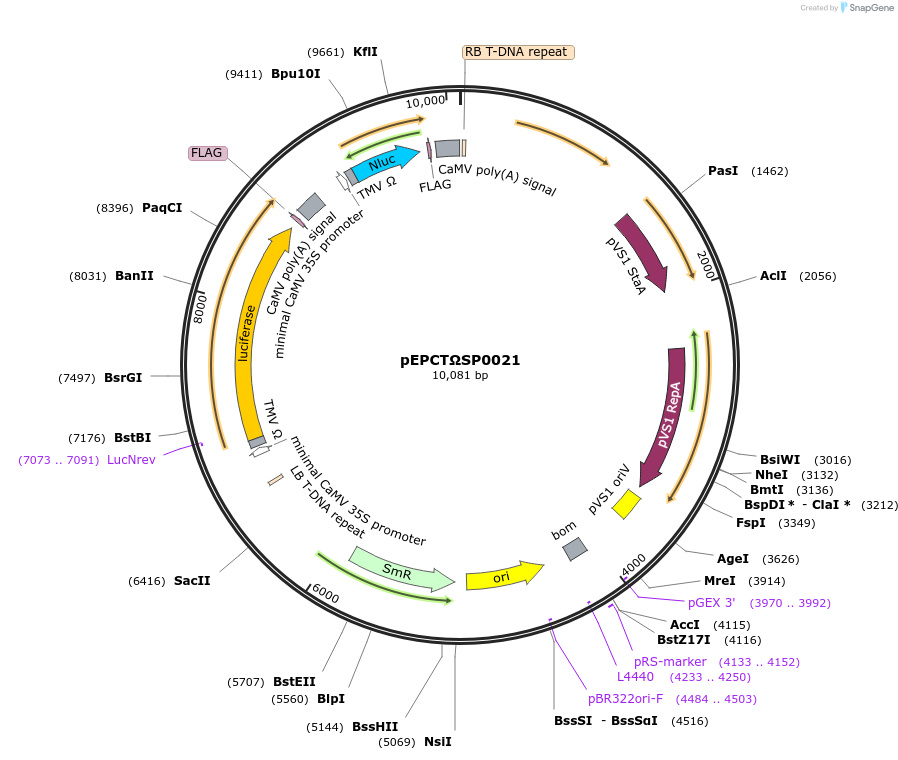
pEPCTΩSP0021
Plasmid#187561PurposeModule for copper-inducible expression of firefly luciferase and nanoluc luciferase driven by minimal synthetic promoters with binding sites for CUP2DepositorInsertFirefly Luciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
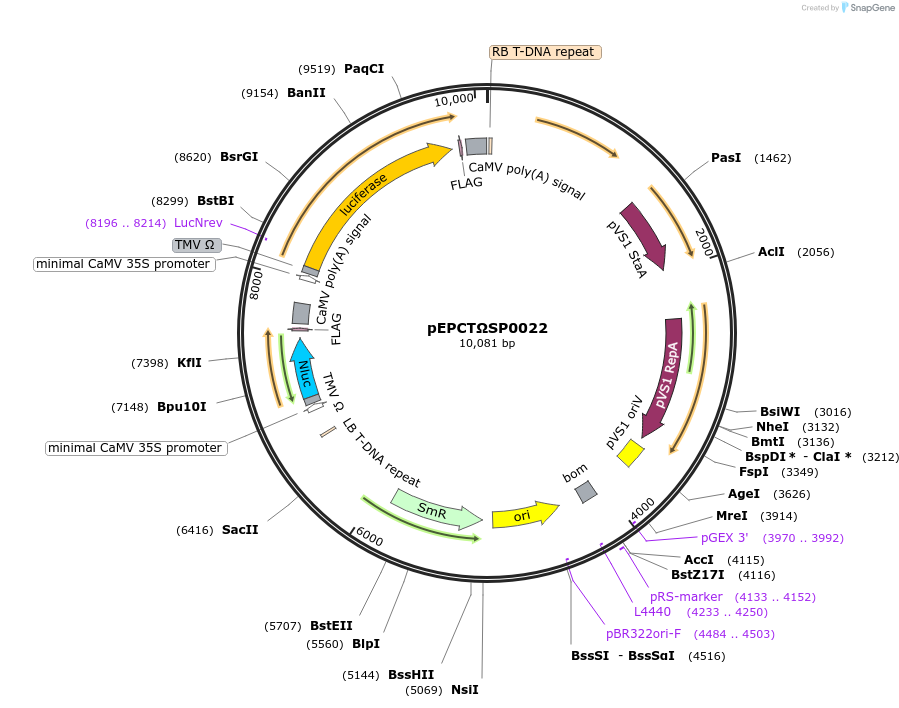
pEPCTΩSP0022
Plasmid#187562PurposeModule for copper-inducible expression of nanoluc luciferase and firefly luciferase driven by minimal synthetic promoters with binding sites for CUP2DepositorInsertFirefly Luciferase and NanoLuciferase
ExpressionPlantAvailable SinceFeb. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
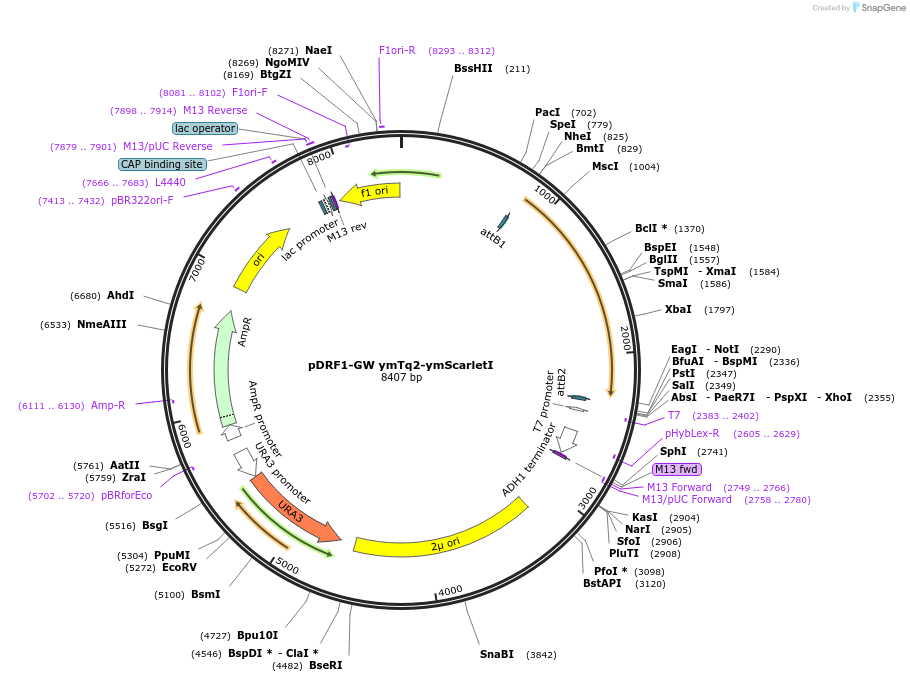
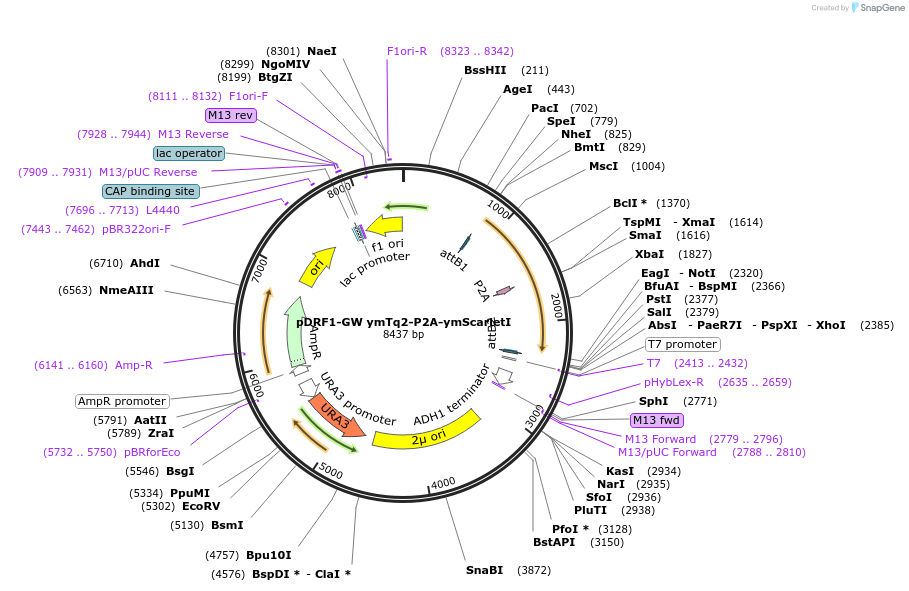
pDRF1-GW ymTq2-ymScarletI
Plasmid#205512PurposeExpression of ymTq2-ymScarletl in yeastDepositorInsertymTq2-ymScarletl
ExpressionYeastAvailable SinceSept. 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
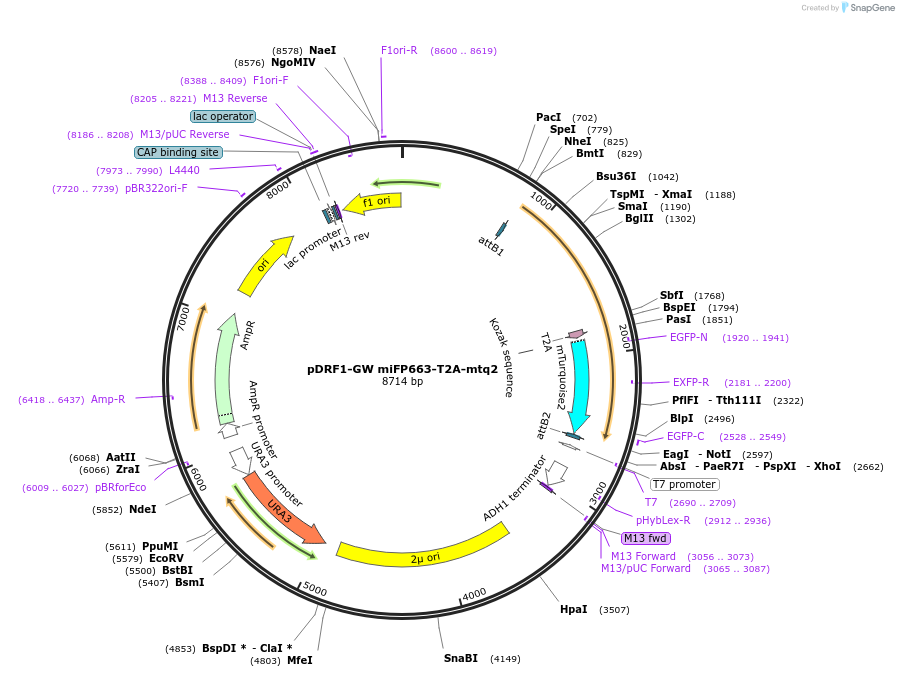
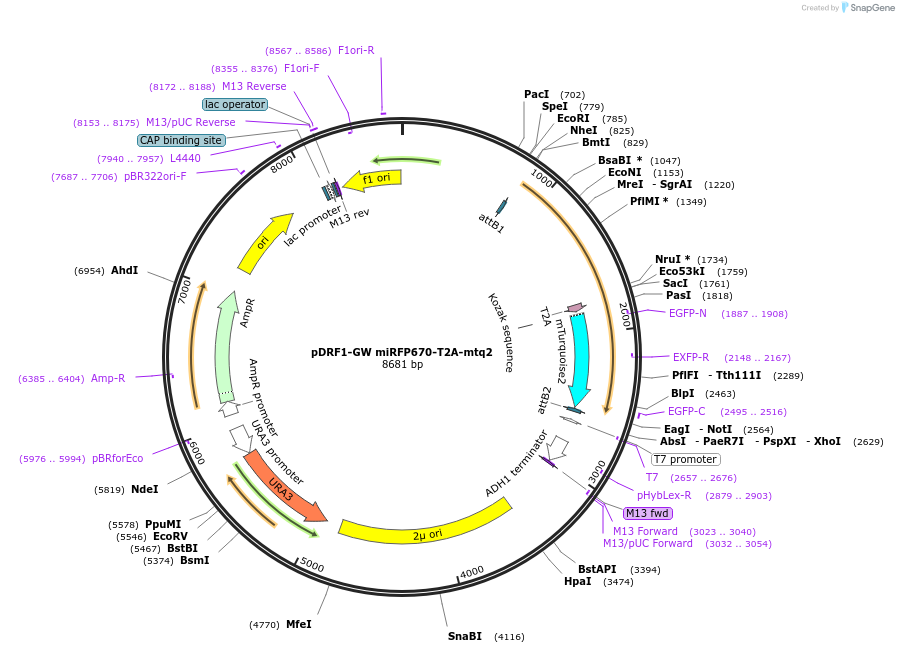
pDRF1-GW miFP663-T2A-mtq2
Plasmid#205504PurposeExpression of miFP663-T2A-mtq2 in yeastDepositorInsertmiFP663-T2A-mtq2
ExpressionYeastAvailable SinceSept. 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
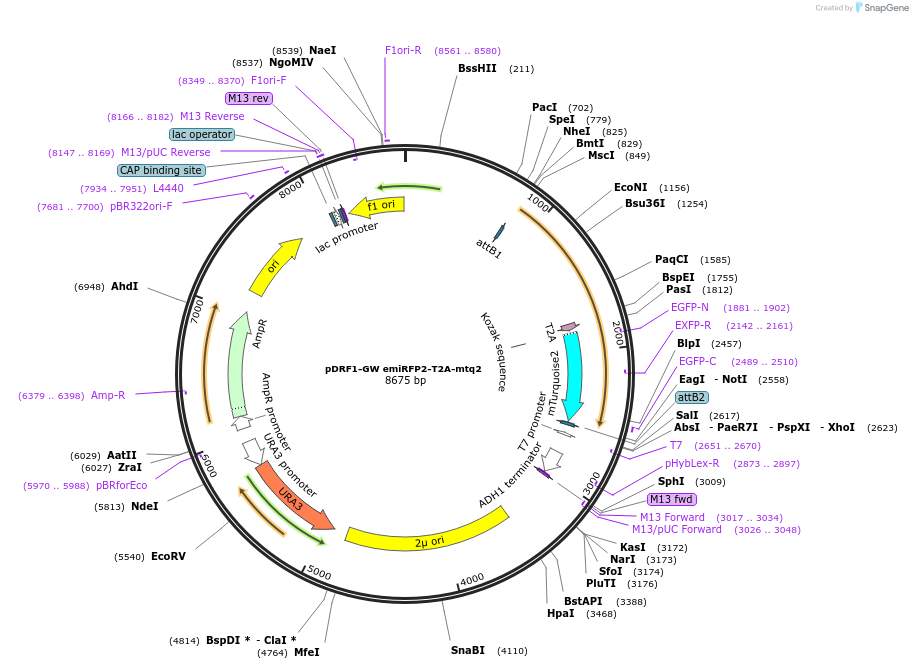
pDRF1-GW emiRFP2-T2A-mtq2
Plasmid#205497PurposeExpression of emiRFP2-T2A-mtq2 in yeastDepositorInsertemiRFP2-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pDRF1-GW ymTq2-P2A-ymScarletI
Plasmid#205500PurposeExpression of ymTq2-P2A-ymScarletI in yeastDepositorInsertymTq2-P2A-ymScarletI
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
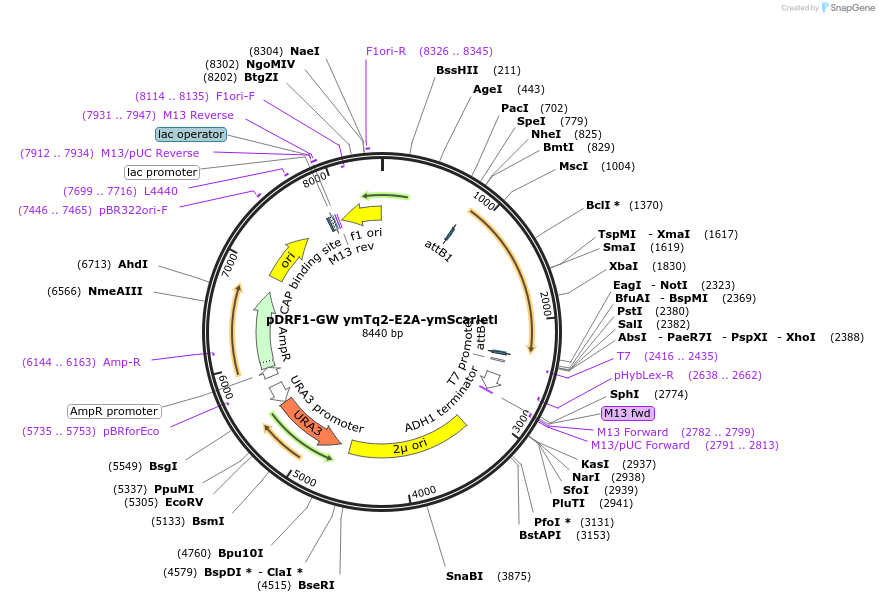
pDRF1-GW ymTq2-E2A-ymScarletl
Plasmid#205510PurposeExpression of ymTq2-E2A-ymScarletl in yeastDepositorInsertymTq2-E2A-ymScarletl
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
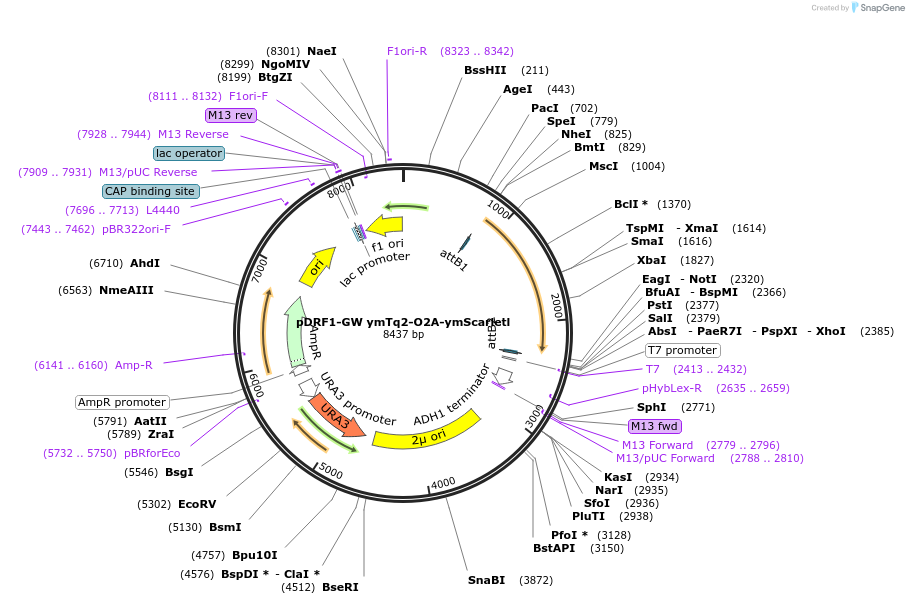
pDRF1-GW ymTq2-O2A-ymScarletl
Plasmid#205509PurposeExpression of ymTq2-O2A-ymScarletl in yeastDepositorInsertymTq2-O2A-ymScarletl
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
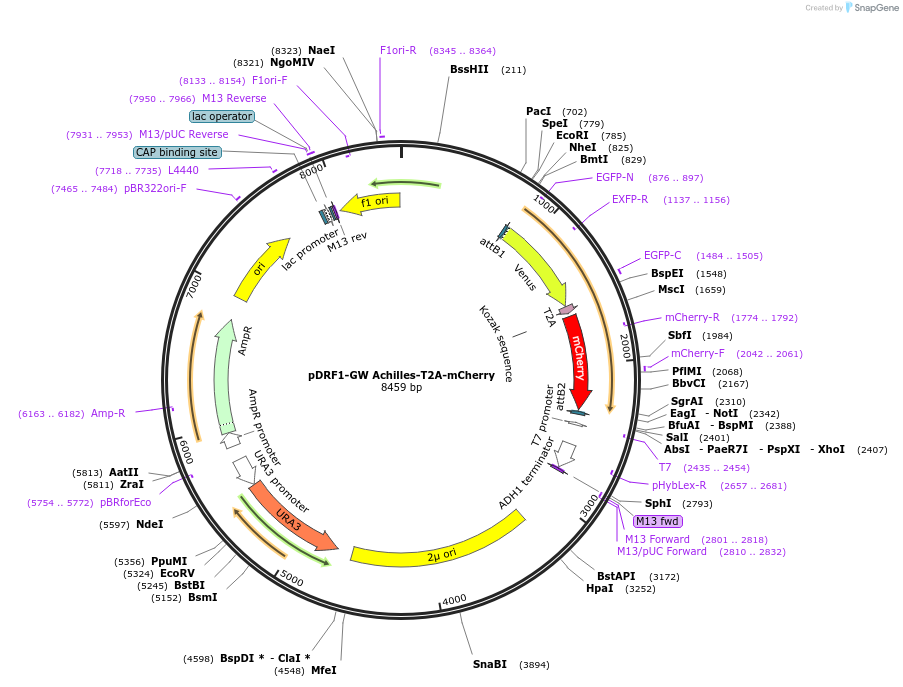
pDRF1-GW Achilles-T2A-mCherry
Plasmid#205499PurposeExpression of Achilles-T2A-mCherry in yeastDepositorInsertAchilles-T2A-mCherry
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
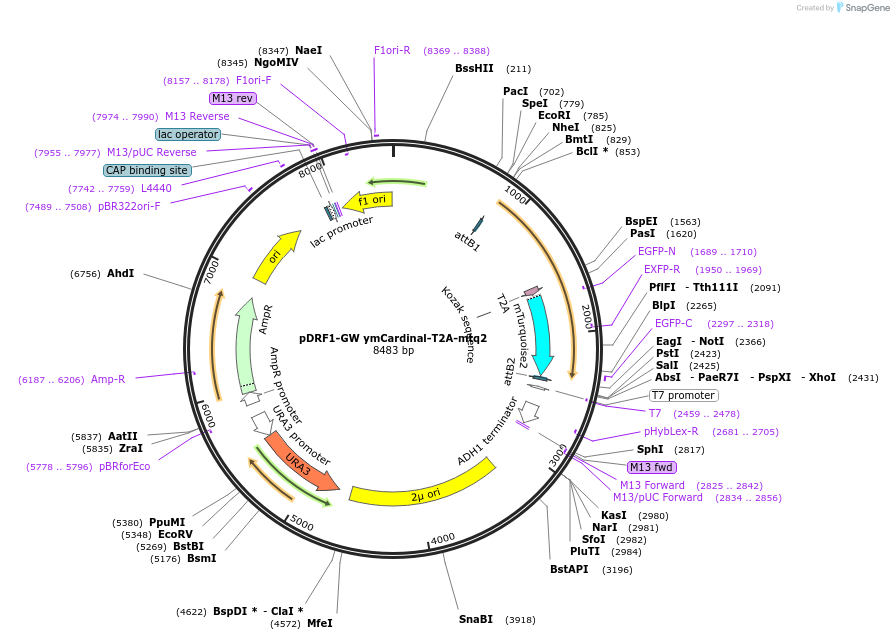
pDRF1-GW ymCardinal-T2A-mtq2
Plasmid#205508PurposeExpression of ymCardinal-T2A-mtq2 in yeastDepositorInsertymCardinal-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
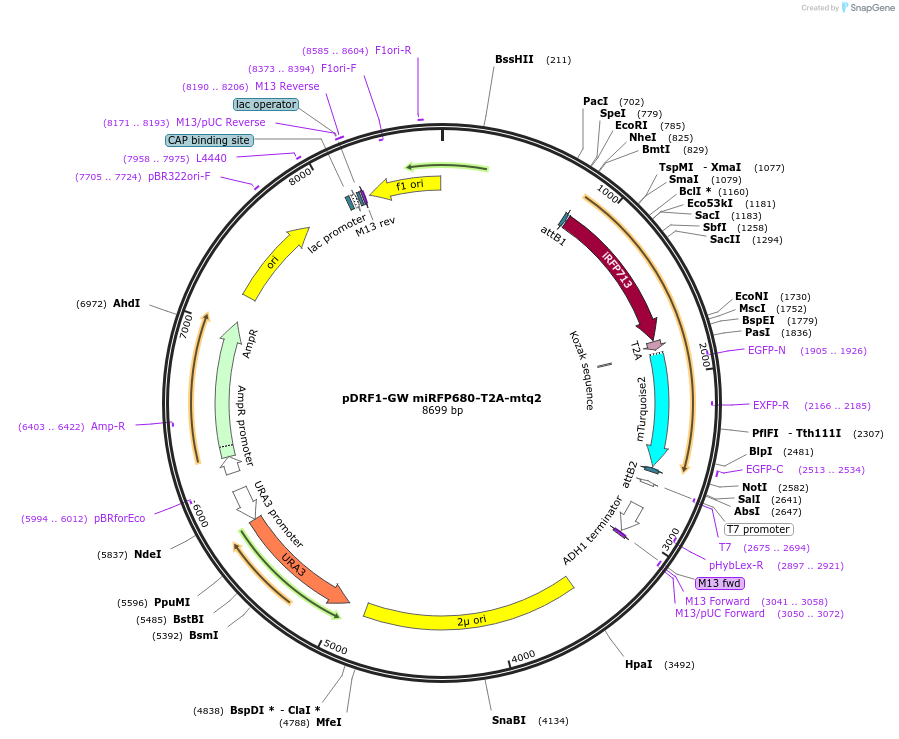
pDRF1-GW miRFP680-T2A-mtq2
Plasmid#205502PurposeExpression of miRFP680-T2A-mtq2 in yeastDepositorInsertmiRFP680-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
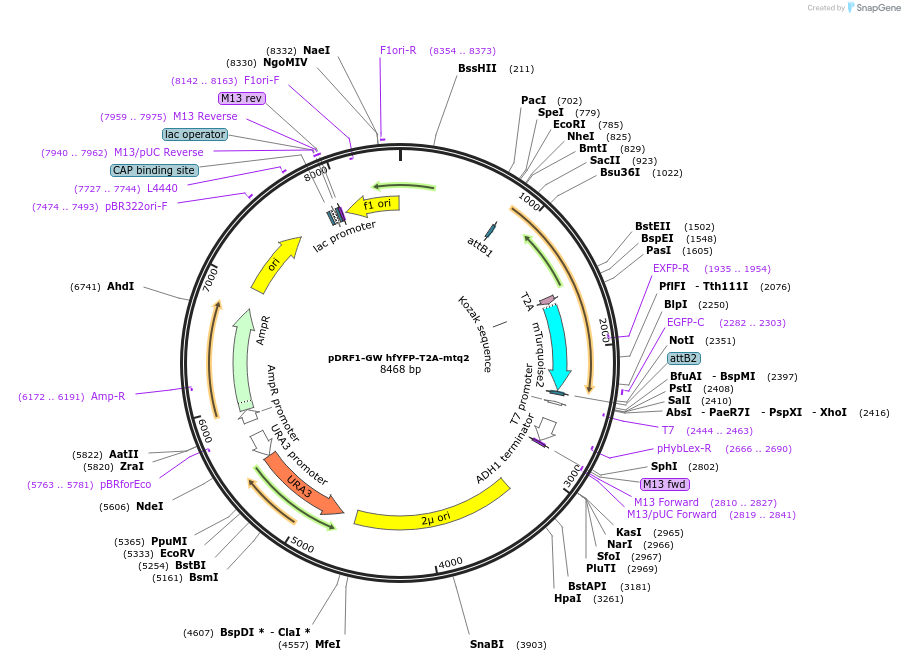
pDRF1-GW hfYFP-T2A-mtq2
Plasmid#205496PurposeExpression of hfYFP-T2A-mtq2 in yeastDepositorInserthfYFP-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
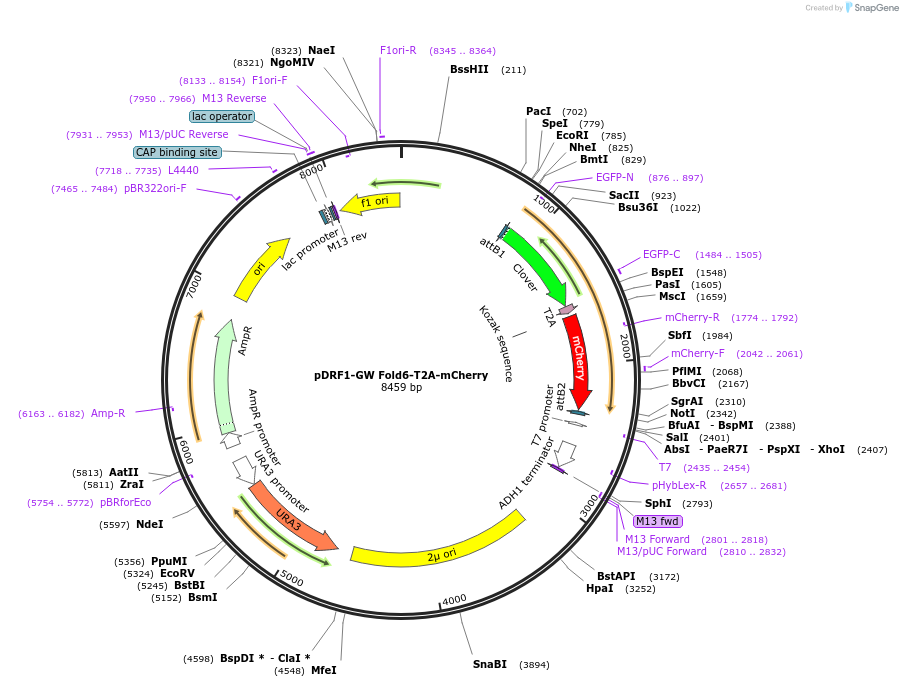
pDRF1-GW Fold6-T2A-mCherry
Plasmid#205492PurposeExpression of Fold6-T2A-mCherry in yeastDepositorInsertFold6-T2A-mCherry
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
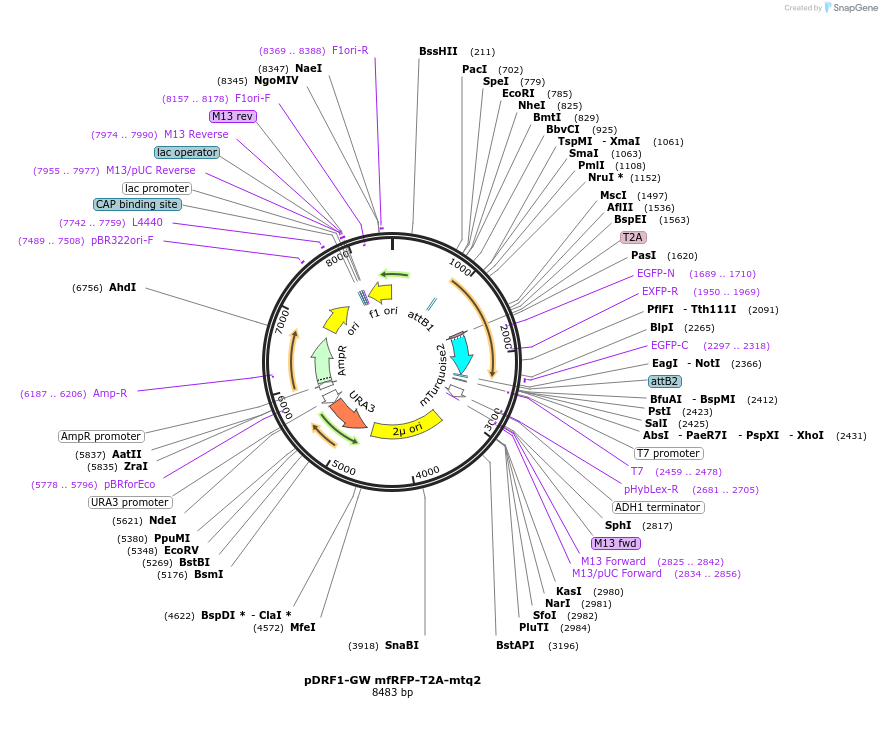
pDRF1-GW mfRFP-T2A-mtq2
Plasmid#205494PurposeExpression of mfRFP-T2A-mtq2 in yeastDepositorInsertmfRFP-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
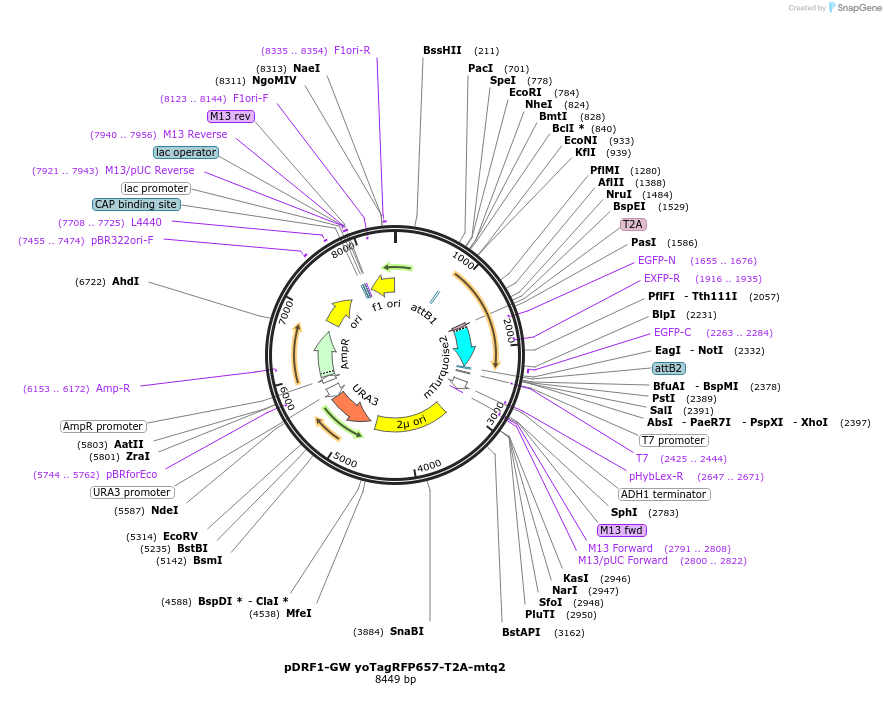
pDRF1-GW yoTagRFP657-T2A-mtq2
Plasmid#205505PurposeExpression of yoTagRFP657-T2A-mtq2 in yeastDepositorInsertyoTagRFP657-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pDRF1-GW miRFP2-T2A-mtq2
Plasmid#205503PurposeExpression of miRFP2-T2A-mtq2 in yeastDepositorInsertmiRFP2-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
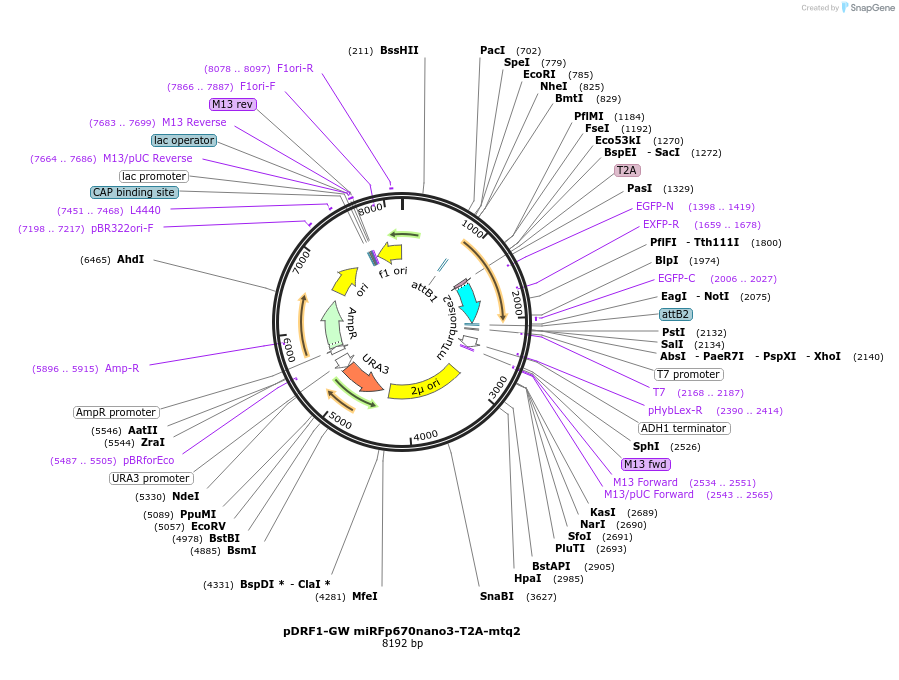
pDRF1-GW miRFp670nano3-T2A-mtq2
Plasmid#205501PurposeExpression of miRFp670nano3-T2A-mtq2 in yeastDepositorInsertmiRFp670nano3-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
pDRF1-GW miRFP670-T2A-mtq2
Plasmid#205491PurposeExpression of miRFP670-T2A-mtq2 in yeastDepositorInsertmiRFP670-T2A-mtq2
ExpressionYeastAvailable SinceSept. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
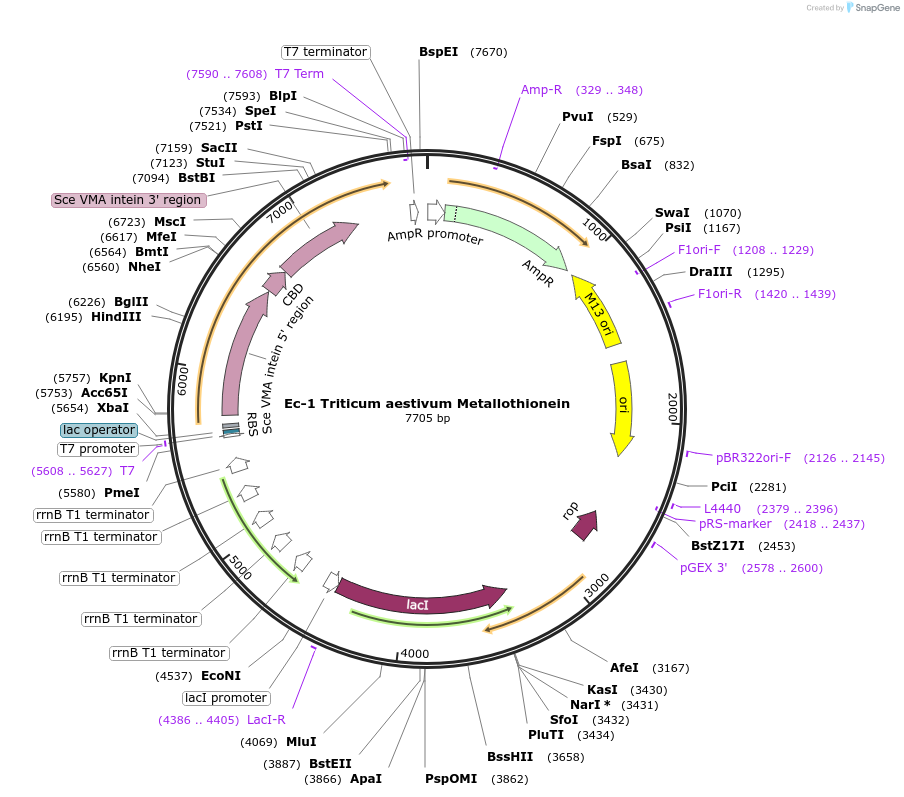
Ec-1 Triticum aestivum Metallothionein
Plasmid#200296PurposeBacterial expression plasmids for production of recombinant Triticum aestivum MetallothioneinDepositorInsertGene of metallothionein with Chiting Binding Domanin-Intein
TagsChiting Binding Domanin-InteinExpressionBacterialAvailable SinceJuly 17, 2023AvailabilityAcademic Institutions and Nonprofits only -
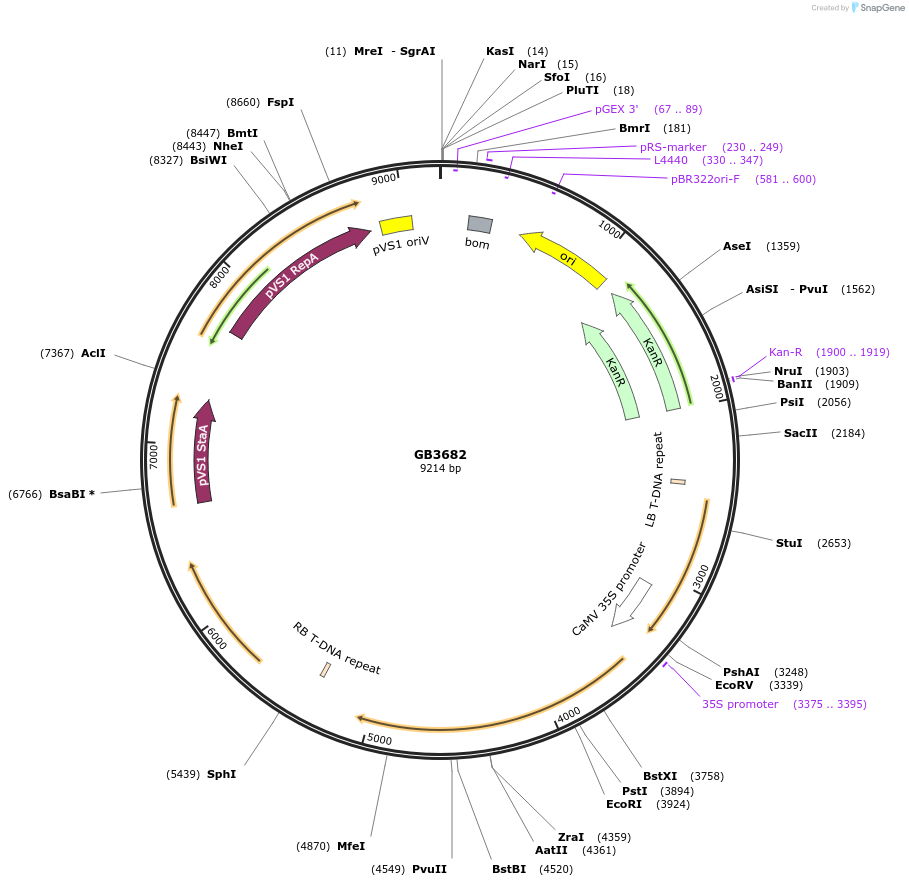
GB3682
Plasmid#187807PurposeTranscriptional unit for expression of alcohol O-acetyltransferase from Saccharomyces pastorianus strain CBS 1483 chromosome SeVIII-SeXV, codon optimized for Nicotiana.DepositorInsertSpATF1-2
ExpressionPlantAvailable SinceJune 1, 2023AvailabilityAcademic Institutions and Nonprofits only -
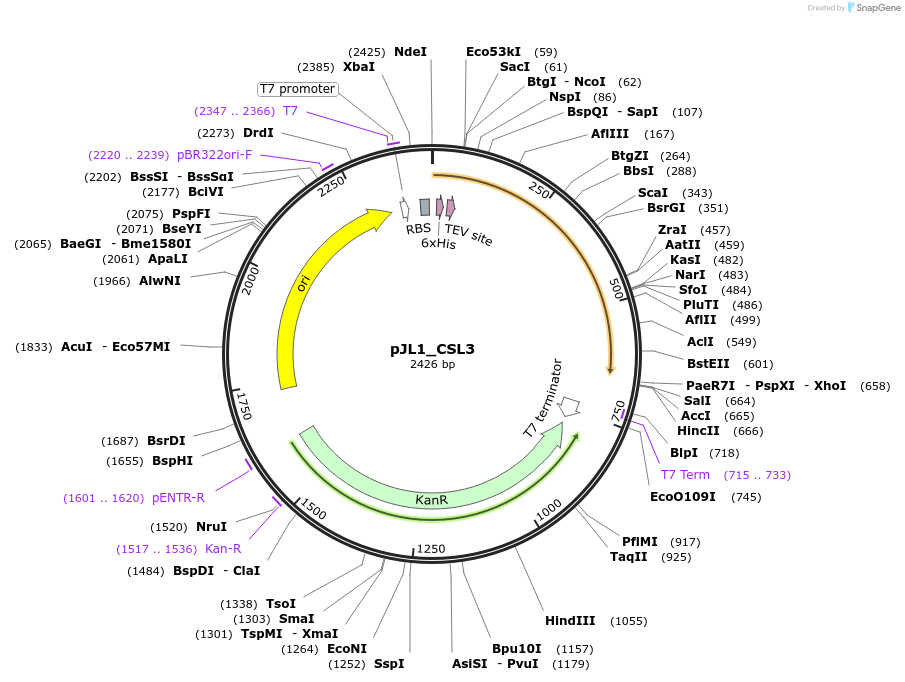
pJL1_CSL3
Plasmid#200173PurposeAmino acids 1-195 of CSL3 with an N-terminal 6xHis-tag followed by a TEV site in the pJL1 vectorDepositorInsertCSL3
Tags6x His tag and TEV cleavage siteExpressionBacterialPromoterT7Available SinceMay 23, 2023AvailabilityAcademic Institutions and Nonprofits only -
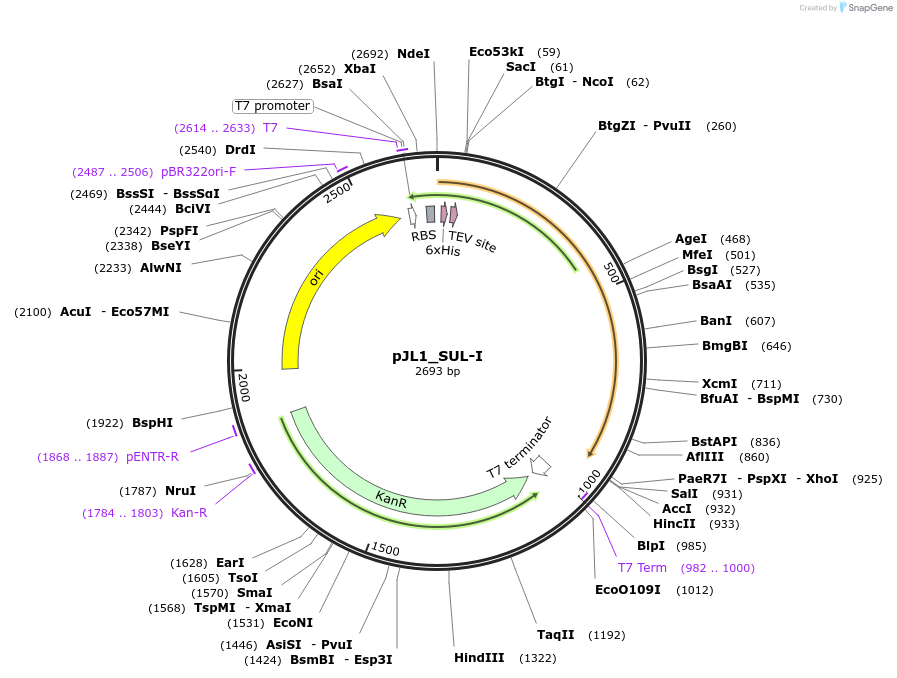
pJL1_SUL-I
Plasmid#200172PurposeAmino acids 25-308 of SUL-I with an N-terminal 6xHis-tag followed by a TEV site in the pJL1 vectorDepositorInsertSUL-I
Tags6x His tag and TEV cleavage siteExpressionBacterialPromoterT7Available SinceMay 23, 2023AvailabilityAcademic Institutions and Nonprofits only -
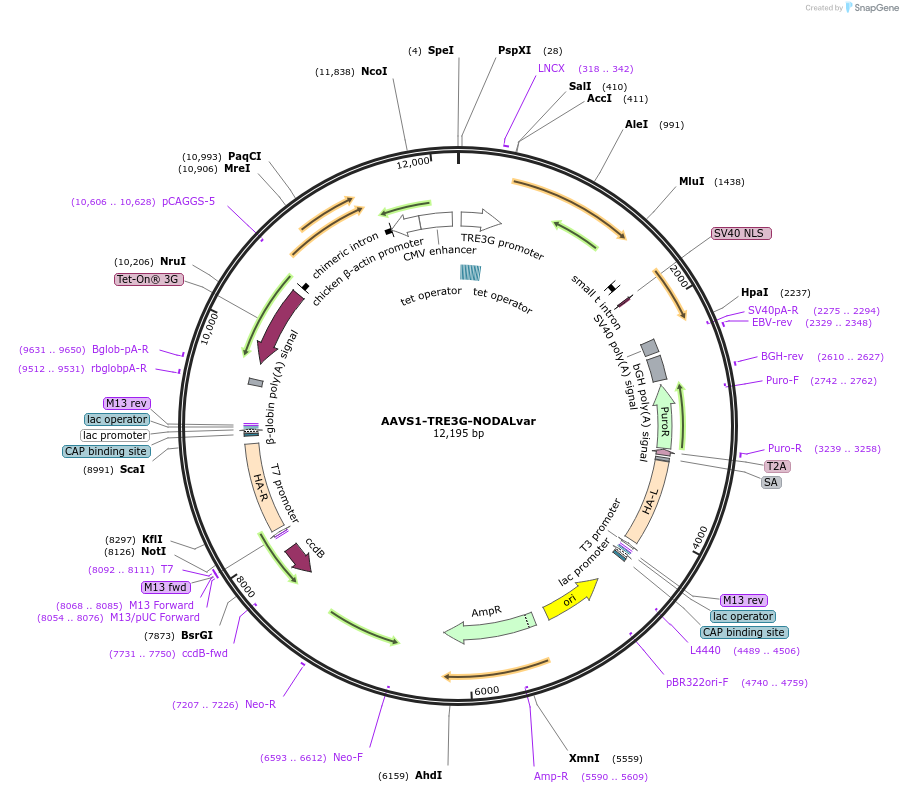
AAVS1-TRE3G-NODALvar
Plasmid#115640PurposeFor targeted integration and inducible expression of a human NODAL splice variant using doxycyclineDepositorAvailable SinceSept. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
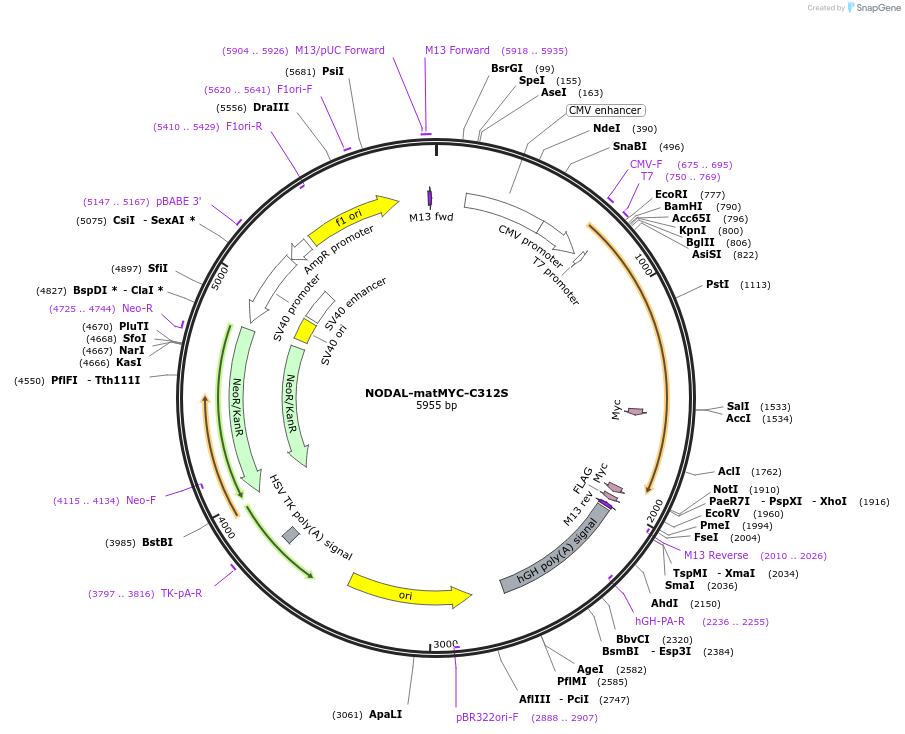
NODAL-matMYC-C312S
Plasmid#115256PurposeFor mammalian expression of the human NODAL open reading frame (with a mutated Cysteine residue) with an internal MYC tagDepositorAvailable SinceSept. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
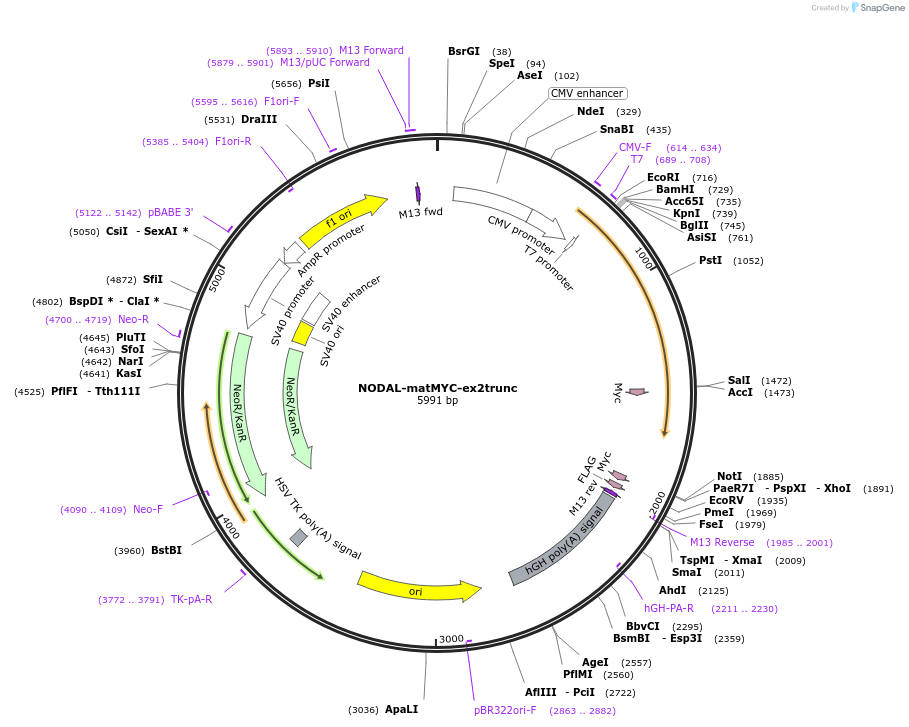
NODAL-matMYC-ex2trunc
Plasmid#115257PurposeFor mammalian expression of a C-terminal truncated human NODAL open reading frame with an internal MYC tagDepositorAvailable SinceSept. 27, 2022AvailabilityAcademic Institutions and Nonprofits only